Clear Sky Science · en
Whole-genome resequencing and genetic diversity of five indigenous cattle breeds from China
Why Local Cows Matter to All of Us
Across China’s Sichuan Province, small farmers rely on rugged yellow cattle that can handle steep hills, hot humid summers, and scarce feed. These animals quietly support food security and rural livelihoods, yet their unique traits are at risk of being lost as modern breeds and crossbreeding spread. This study reads nearly every letter of DNA in several Sichuan cattle breeds, creating a detailed genetic map that can help protect these animals, improve beef production, and prepare livestock for a changing climate.
Cattle Built for Tough Landscapes
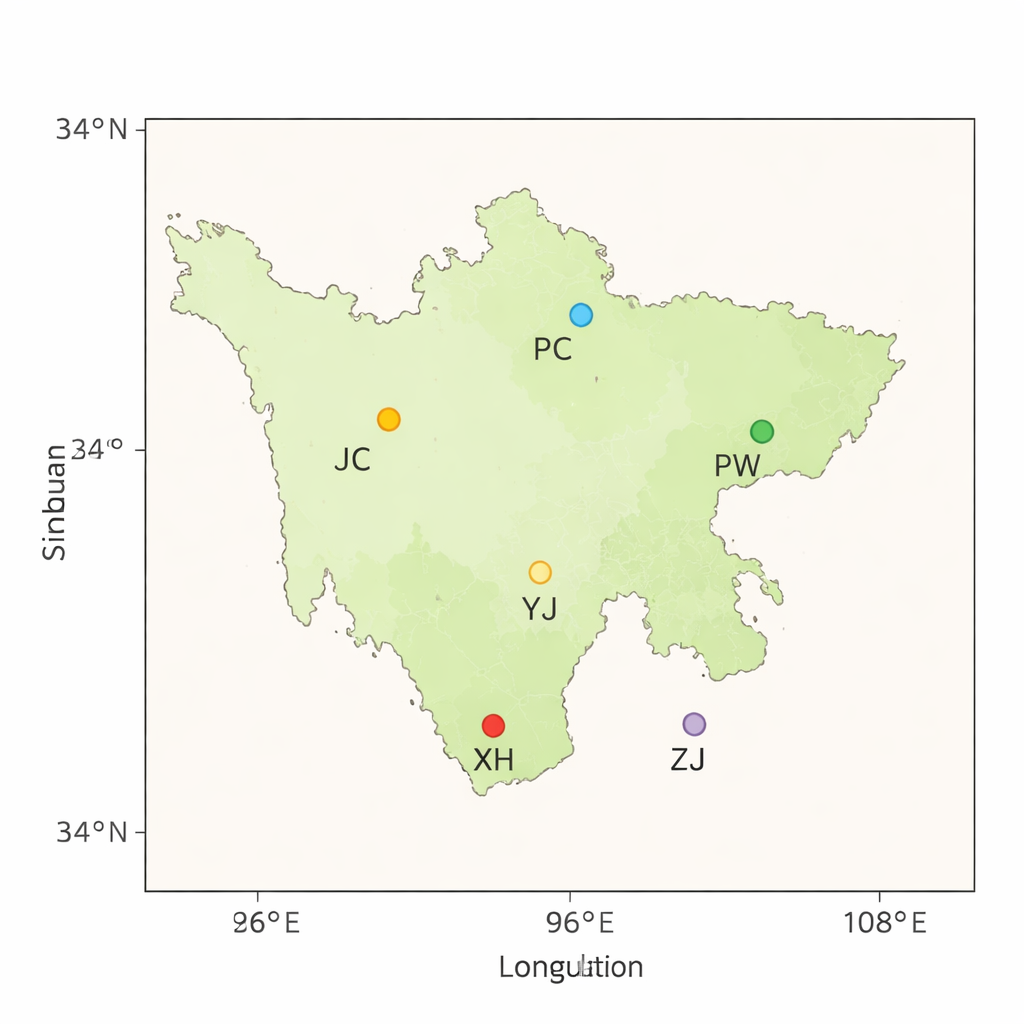
Sichuan is one of China’s hot spots for cattle diversity, with millions of animals spread across basins, hills, and highlands. Local yellow cattle have been shaped by centuries of survival in rugged environments and by careful selection from farmers who valued strength, disease resistance, and the ability to live on rough forage. Yet these same breeds now face shrinking numbers and genetic erosion. Uncontrolled crossbreeding, seasonal movement of herds, disease outbreaks, floods, and the rapid mixing of animals through trade all blur the boundaries between traditional breeds. Without deliberate conservation, the distinct biological strengths of these cattle could disappear.
Reading the Whole Genetic Story
To understand and safeguard this diversity, the researchers collected blood from 56 yellow cattle across five Sichuan counties, each representing a local breed. They also included DNA from 10 high-altitude yaks as a comparison group, or outgroup, to place the cattle’s genetics in a broader evolutionary context. Using modern high-throughput sequencing, they generated about 2.3 terabytes of DNA data, reading each animal’s genome roughly 13 times over to ensure accuracy. The reads were then aligned to a standard cattle reference genome, with more than 99% of them landing in the right place, a sign of very high data quality.
Millions of Differences in the DNA Code
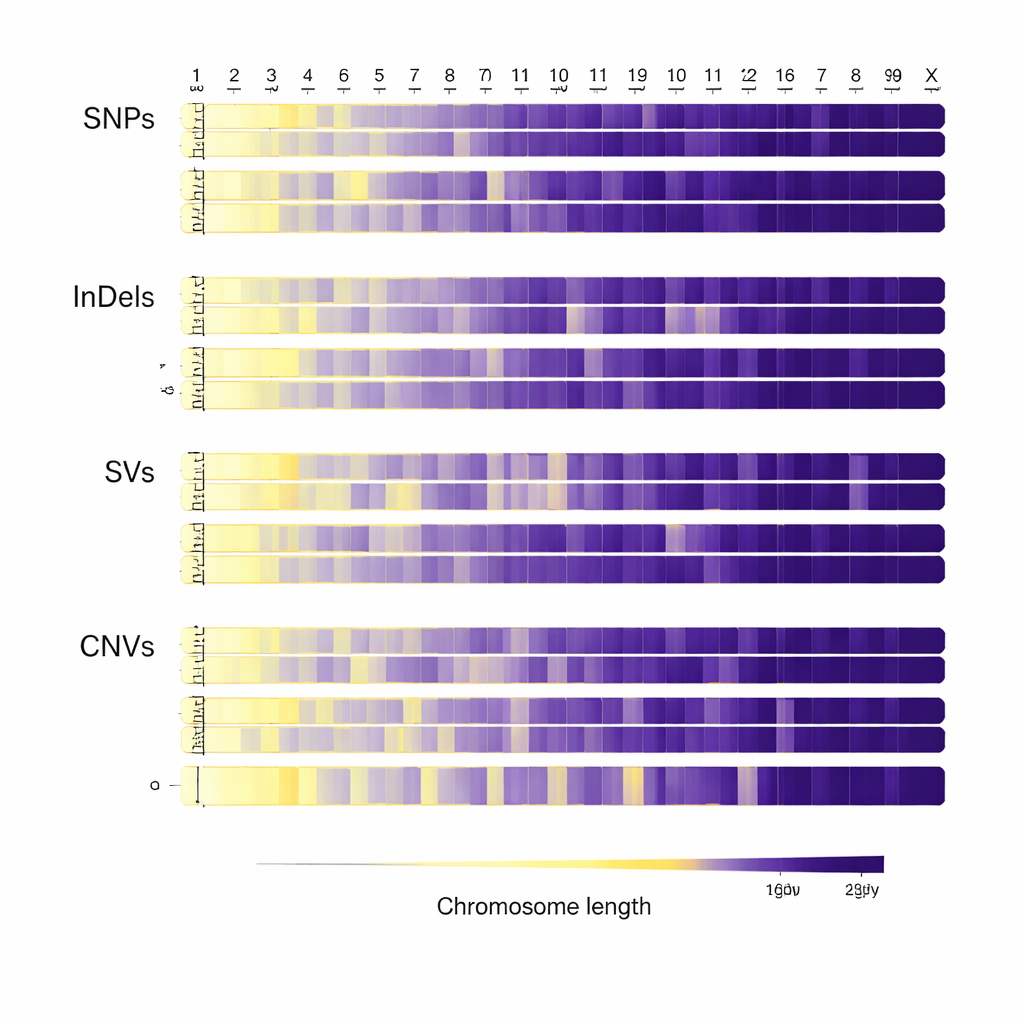
Once the genomes were aligned, the team scanned them for different kinds of genetic variation. They found about 31 million single-letter changes in the DNA sequence, as well as more than two million tiny insertions and deletions, tens of thousands of larger structural changes, and over 240,000 regions where chunks of DNA are missing or duplicated. Most of these differences lie outside the protein-coding parts of genes, but a substantial number fall within or near genes that could affect important traits. The pattern of changes across the chromosomes, especially the distinct behavior of the X chromosome, reflects both the biology of cattle and the evolutionary pressures they have faced over time.
Turning Raw Data into Biological Insight
Crucially, the researchers did more than just count DNA changes—they annotated them to predict possible effects on gene function. Using widely adopted bioinformatics tools, they linked each variant to its position in or near genes and highlighted changes that might disrupt proteins or alter gene regulation. They then made all of the raw sequence data and processed variant lists freely available in international databases. This openness means that scientists worldwide can mine the data to search for genes tied to heat and humidity tolerance, disease resistance, efficient use of poor-quality feed, and meat quality, or to compare Sichuan cattle with other breeds.
From Hidden Code to Practical Benefit
For non-specialists, this work can be viewed as taking a high-resolution genetic snapshot of some of China’s toughest cattle. By cataloging their DNA differences in detail and sharing the results openly, the study lays essential groundwork for breeding programs that keep the resilience of traditional animals while improving productivity. In a future of climate stress and shifting diseases, such information will help farmers and breeders maintain hardy, locally adapted cattle that support sustainable beef production and protect an irreplaceable piece of agricultural heritage.
Citation: Wang, W., Li, L., Chen, Y. et al. Whole-genome resequencing and genetic diversity of five indigenous cattle breeds from China. Sci Data 13, 282 (2026). https://doi.org/10.1038/s41597-026-06610-y
Keywords: cattle genetics, indigenous breeds, genetic diversity, whole-genome sequencing, livestock conservation