Clear Sky Science · en
Chromosome-level genome assembly and annotation of the tropical sea cucumber Holothuria fuscocinerea
Why a humble sea cucumber matters
Sea cucumbers may look like little more than sand-covered sausages on the seafloor, but they are quiet workhorses of tropical reefs and a growing focus of fisheries and medicine. As demand for these animals rises and wild populations are stressed, scientists need detailed genetic information to manage them wisely and to tap their biochemical riches. This study delivers the first chromosome-level map of the genome of Holothuria fuscocinerea, a widespread tropical sea cucumber, creating a reference blueprint that future ecologists, breeders and drug discoverers can use.
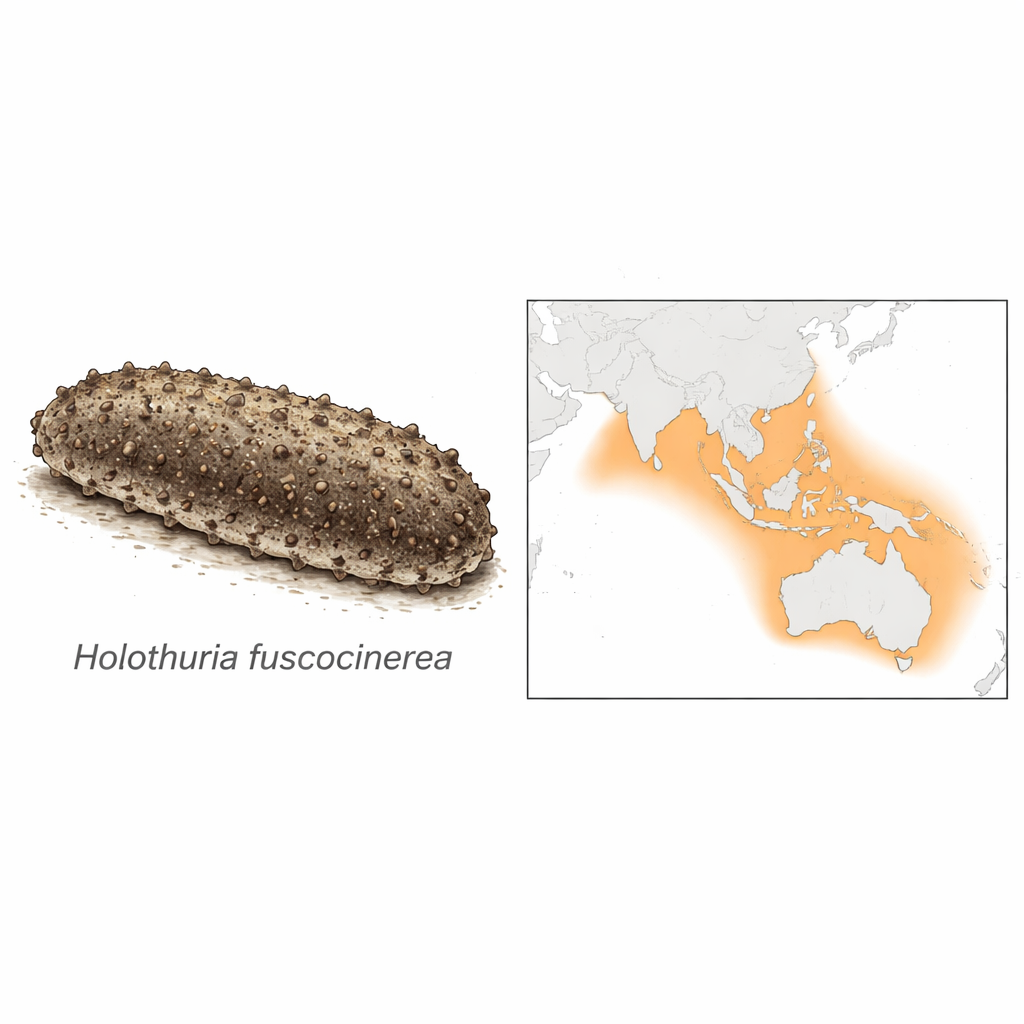
A widespread cleaner of tropical seas
Holothuria fuscocinerea lives across warm reefs from the Red Sea and East Africa through the Indian and Pacific Oceans to China, southern Japan, northern Australia and remote Pacific islands. It typically grows to about half a meter long, with an oval, slightly flattened body whose rough brown-gray upper surface often carries a layer of sand, helping it blend into the seabed. Like many of its relatives, it carries special defensive structures called Cuvierian tubules, which can be expelled when threatened. Though it currently has modest commercial value compared with some prized "sea cucumber" species, it is expected to become a target for fisheries as higher-value stocks decline, and it also produces bioactive compounds with antimicrobial, immune-stimulating and anticancer potential.
The need for a full genetic blueprint
Sea cucumbers play multiple ecological roles: they churn and clean sediments, recycle nitrogen, help regulate seawater chemistry and support food webs on coral reefs and other coastal habitats. At the same time, many species are heavily harvested and recover slowly, prompting hatcheries to raise juveniles for release and to move animals between regions. Yet, without a full genome, it has been difficult to track population structure, detect hidden mixing between wild and farmed stocks, or understand how these animals evolved their unusual traits, such as soft tissues that can rapidly change stiffness and expendable defensive organs. Until now, work on H. fuscocinerea has focused on local abundance, identification, feeding and selected chemicals, while genetics relied mostly on short mitochondrial sequences, leaving the species’ evolutionary relationships partly unresolved.
Building the sea cucumber genome
The researchers combined several modern DNA sequencing strategies to assemble the genome at near-chromosome resolution. Short-read sequencing delivered large volumes of accurate, small DNA snippets; long-read sequencing provided continuous stretches that span repetitive regions; and Hi-C mapping captured how DNA is folded in 3D inside the cell nucleus, revealing which fragments belong on the same chromosome. Using specialized software, they stitched these data into 541 continuous pieces and then arranged and oriented them into 23 pseudochromosomes, together covering about 1.56 billion DNA letters. Quality checks showed that the assembly is highly complete and accurate, with few gaps and strong support from independent measures of gene content and error rate.
What the genome reveals inside
With the assembled chromosomes in hand, the team systematically catalogued the genetic elements. They identified nearly 30,000 protein-coding genes and verified their structures using RNA from ten different tissues, including skin, intestine, tentacles, defensive organs and reproductive organs. About 95 percent of these genes could be linked to known functions or families using major biological databases. Roughly half of the genome consists of repetitive DNA, especially mobile genetic elements known as transposons, which can shape genome size and evolution. The researchers also mapped thousands of noncoding RNAs and pinpointed many chromosome ends (telomeres) and probable centromere regions, showing that large parts of the genome are assembled end-to-end. They even reconstructed the complete mitochondrial genome as a separate, compact circular molecule.
A foundation for conservation and discovery
For non-specialists, the key message is that we now have a high-quality, nearly complete genetic reference for an ecologically important and increasingly exploited tropical sea cucumber. This reference will let scientists track genetic diversity in wild and farmed populations, design better breeding and restocking programs, and compare H. fuscocinerea with other sea cucumbers to uncover the genes behind their flexible bodies, unusual defenses and medically promising chemistry. In practical terms, the genome acts like an atlas for this species, guiding efforts to conserve reef ecosystems, sustain coastal livelihoods that depend on sea cucumbers and explore new marine-derived drugs.
Citation: Wang, X., Huang, Q., Qin, Z. et al. Chromosome-level genome assembly and annotation of the tropical sea cucumber Holothuria fuscocinerea. Sci Data 13, 281 (2026). https://doi.org/10.1038/s41597-026-06609-5
Keywords: sea cucumber genome, Holothuria fuscocinerea, marine conservation, chromosome-level assembly, tropical reef ecology