Clear Sky Science · en
Temporal multiomics gene expression data across human embryonic stem cell-derived polyhormonal cell differentiation
How Cells Learn to Become Organs
Our bodies begin as tiny clusters of identical cells that somehow learn to become very different tissues, from brain to pancreas. This study follows that learning process in the lab, using human embryonic stem cells as they are guided to become early pancreatic cells. By tracking the activity of thousands of genes over time at multiple levels, the work creates a rich reference map that can help scientists better understand human development and, in the long run, improve strategies for treating diseases like diabetes.
Watching Stem Cells Choose a Path
Human embryonic stem cells are special because they can turn into almost any cell type in the body. In this study, researchers steered these stem cells toward a specific fate: a pancreatic lineage called polyhormonal cells, which can make key hormones such as insulin and glucagon. They mimicked early development in a dish by changing the growth medium step by step, moving cells first into a primitive gut-like state called endoderm, and then onward to hormone-producing pancreatic cells over 17 days. Samples were collected at ten carefully chosen time points to capture the entire journey from flexible stem cell to specialized hormone-producing cell. 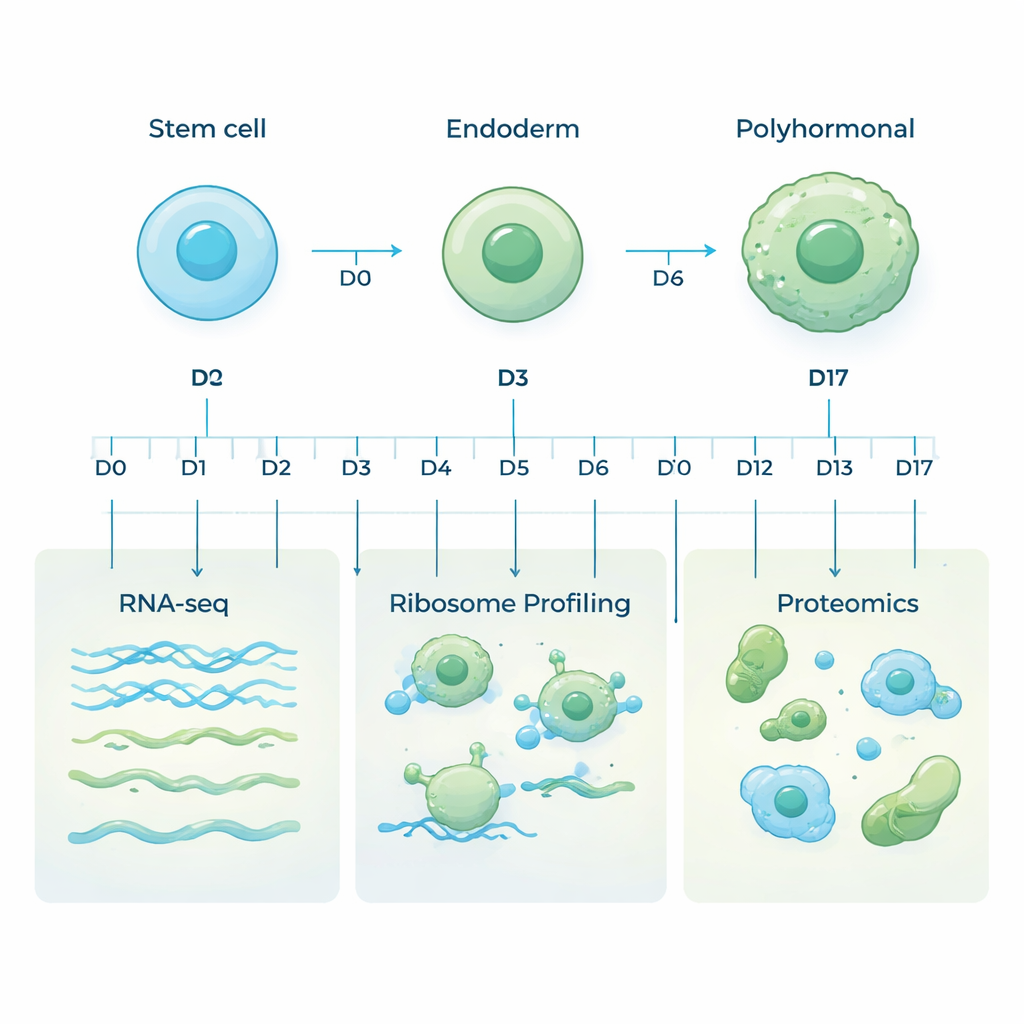
Looking Under the Hood at Three Levels
Most studies look only at messenger RNA (mRNA), the molecules that carry gene instructions. But mRNA is only part of the story: not every message is translated into protein, and proteins themselves can be made or destroyed at different rates. To get a fuller view, the team used three complementary approaches on the same samples. RNA sequencing measured which genes were being transcribed into mRNA. Ribosome profiling tracked which messages were actively being read by the cell’s protein-making machines. Mass spectrometry-based proteomics then measured the actual proteins present. Together, these layers reveal how gene activity is controlled as cells change identity.
Following Key Signals of Cell Identity
To check that the cells really followed the intended developmental route, the scientists monitored well-known marker genes. Early on, classic stem cell markers such as OCT4 and NANOG were high and then faded as differentiation progressed. As cells entered the endoderm stage, markers like KIT and SOX17 rose. By the final stages, polyhormonal markers insulin (INS) and glucagon (GCG) appeared strongly at both the RNA and protein level, confirming that the cells had adopted a pancreatic-like hormone-producing identity. Although one of the biological replicates moved through these stages slightly more slowly than the other, both followed the same overall path, reflecting small natural differences rather than technical problems. 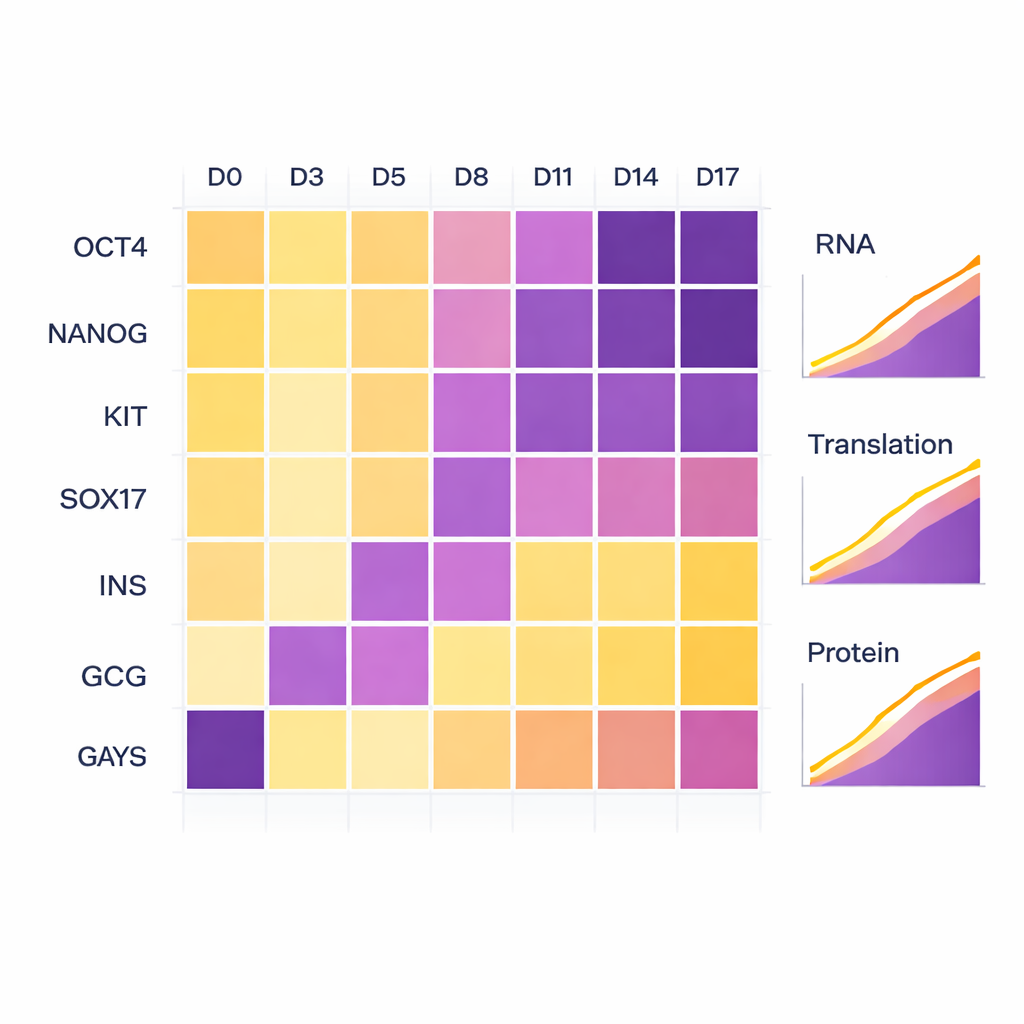
Quality Checks for a Trusted Resource
Because this work is meant to serve as a community resource, the authors devoted extensive effort to checking data quality and consistency. For each of the three methods, they assessed sequencing and measurement accuracy, coverage of genes, and how well repeated experiments agreed. Stem cell and polyhormonal stages showed clear and reproducible differences across RNA, translation, and protein levels. Principal component analyses—statistical maps that group similar samples together—showed that time points lined up in order, with early and late stages clearly separated and biological replicates clustering tightly. The proteomics data alone reliably tracked nearly 7,500 proteins across all time points, with relatively few missing measurements, underscoring the depth of the dataset.
A Foundation for Future Discoveries
The authors make all raw and processed data publicly available, along with analysis scripts and reference files, so that other researchers can reuse and reanalyze the dataset. Beyond describing how one cell type becomes another, this work offers a detailed time-resolved picture of how gene messages, their translation, and resulting proteins interact during a major developmental transition. For non-specialists, the key takeaway is that cell fate is controlled by several layers of regulation working together over time, and that this dataset provides a high-resolution “movie” of those changes. Scientists can now use this resource to probe why some genes change early and others late, how different organs may follow similar or distinct rules, and how to better guide stem cells toward medically useful fates.
Citation: Keskin, A., Shayya, H.J., Patel, A. et al. Temporal multiomics gene expression data across human embryonic stem cell-derived polyhormonal cell differentiation. Sci Data 13, 278 (2026). https://doi.org/10.1038/s41597-026-06606-8
Keywords: stem cells, pancreas development, gene expression, multiomics, polyhormonal cells