Clear Sky Science · en
Chromosome-level genome assemblies of two maize inbred lines with contrasting plant architectures
Why corn shape matters for feeding the world
From the roadside to supermarket shelves, corn is everywhere. But not all corn plants look—or perform—the same. Some grow tall with wide, spreading leaves, while others are shorter and more upright. These differences in “plant shape” help determine how many plants can be packed into a field and, ultimately, how much food we can harvest from each hectare. This study decodes, in remarkable detail, the DNA of two inbred corn lines with contrasting shapes, creating a reference map that plant breeders and scientists can use to design future high-yield, climate-ready crops.
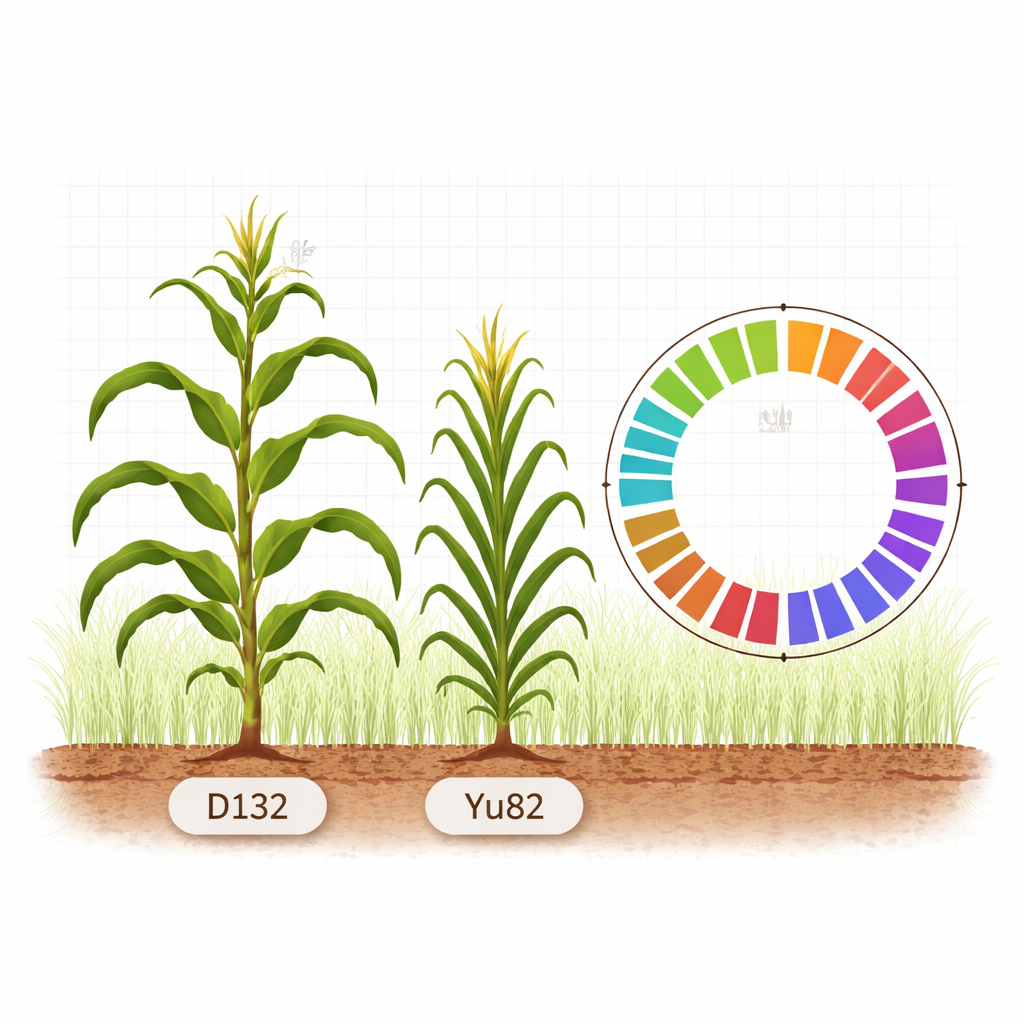

Two corn plants, two very different silhouettes
The researchers focused on two inbred maize lines, D132 and Yu82, that differ strikingly in the way they grow. D132 has an open, sprawling structure: it is taller, its ear (the cob-bearing part) sits higher, and its leaves spread out more widely. Yu82, by contrast, is compact. It is shorter, the ear is closer to the ground, and the leaves are more upright and narrow. These traits are not cosmetic. A compact structure lets farmers plant more stalks per square meter without excessive shading or competition, a key requirement for boosting yield in crowded fields. By comparing the complete DNA “instruction books” of these two lines, the team aims to uncover the genetic foundations of plant architecture in maize.
Building near-complete DNA maps chromosome by chromosome
To capture these instruction books, the team combined several cutting-edge DNA sequencing technologies. Long-read platforms, which can read very long stretches of DNA at once, were used to assemble the basic pieces of each genome. Short-read platforms, which produce many highly accurate small fragments, were then used to polish and correct errors. Hi-C technology, which measures how different parts of the DNA physically touch inside the cell, allowed the researchers to stitch the pieces together into full-length chromosomes. For Yu82, they also used optical mapping, which images extremely long DNA molecules to help order and join fragments. The result is two chromosome-level genome assemblies: D132 at about 2.17 billion DNA letters and Yu82 at about 2.19 billion, with more than 90–99% of their sequences placed cleanly onto the ten maize chromosomes.
What lies inside: genes, repeats, and shared structure
Once the genomes were assembled, the scientists cataloged their contents. Each line carries roughly 41,000 protein-coding genes—segments of DNA that provide instructions for building proteins. They also found that more than four-fifths of each genome is made of “jumping DNA,” known as transposable elements. These repetitive sequences, while often dismissed as genomic clutter, strongly influence genome size and can affect how genes are switched on or off. To check accuracy, the team compared their assemblies to several existing maize reference genomes and to thousands of well-known plant genes. The new maps showed high completeness and closely matched the structure and gene order seen in other well-studied maize lines, confirming that they are reliable foundations for further research.
From raw DNA to useful breeding clues
Beyond simply listing genes, the authors used large collections of RNA data—snapshots of which genes are actively used in different tissues—to refine gene models and attach functional hints to most genes in both genomes. They then examined how the D132 and Yu82 genomes line up with each other and with other maize varieties, identifying long stretches where gene order is conserved. Such comparisons highlight regions where the DNA is stable, as well as hotspots where structure or gene content differs. Those variable regions are prime suspects for harboring genes and switches that shape plant height, leaf angle, ear placement, and root systems—the very traits that separate open, sprawling plants from compact, high-density-ready ones.
How this work helps grow more corn on less land
For non-specialists, the take-home message is that this study delivers two detailed, high-quality DNA maps of corn plants that grow in very different ways. These maps act like reference blueprints: breeders and geneticists can now more easily pinpoint specific genes and DNA changes that control plant architecture, test how they influence performance in crowded fields, and combine favorable versions into next-generation hybrids. In a world where demand for grain is rising but farmland is limited, the ability to design corn plants that thrive when planted close together—and to do so using precise genetic information—could play an important role in producing more food while using land and resources more efficiently.
Citation: Yao, W., Li, S., Ren, J. et al. Chromosome-level genome assemblies of two maize inbred lines with contrasting plant architectures. Sci Data 13, 276 (2026). https://doi.org/10.1038/s41597-026-06603-x
Keywords: maize genome, plant architecture, crop breeding, high-density planting, genome assembly