Clear Sky Science · en
High-resolution single-nematode transcriptomic datasets of plant-parasitic nematodes from juveniles to adults
Hidden Worms, Huge Harvest Losses
Many farmers fight an invisible enemy: microscopic worms in the soil that invade plant roots and quietly rob fields of yield. These plant-parasitic nematodes, especially the root‑knot nematode Meloidogyne incognita, damage staple crops like cotton, tomato, and soybean and cost the world an estimated 157 billion dollars every year. To outsmart these pests, scientists need to understand what their genes are doing at each step of their life inside the root. This study builds a detailed "gene activity atlas" for individual nematodes as they grow from invasive juveniles to egg‑laying adults, creating a public resource that researchers and breeders can use to develop smarter, more targeted control methods.
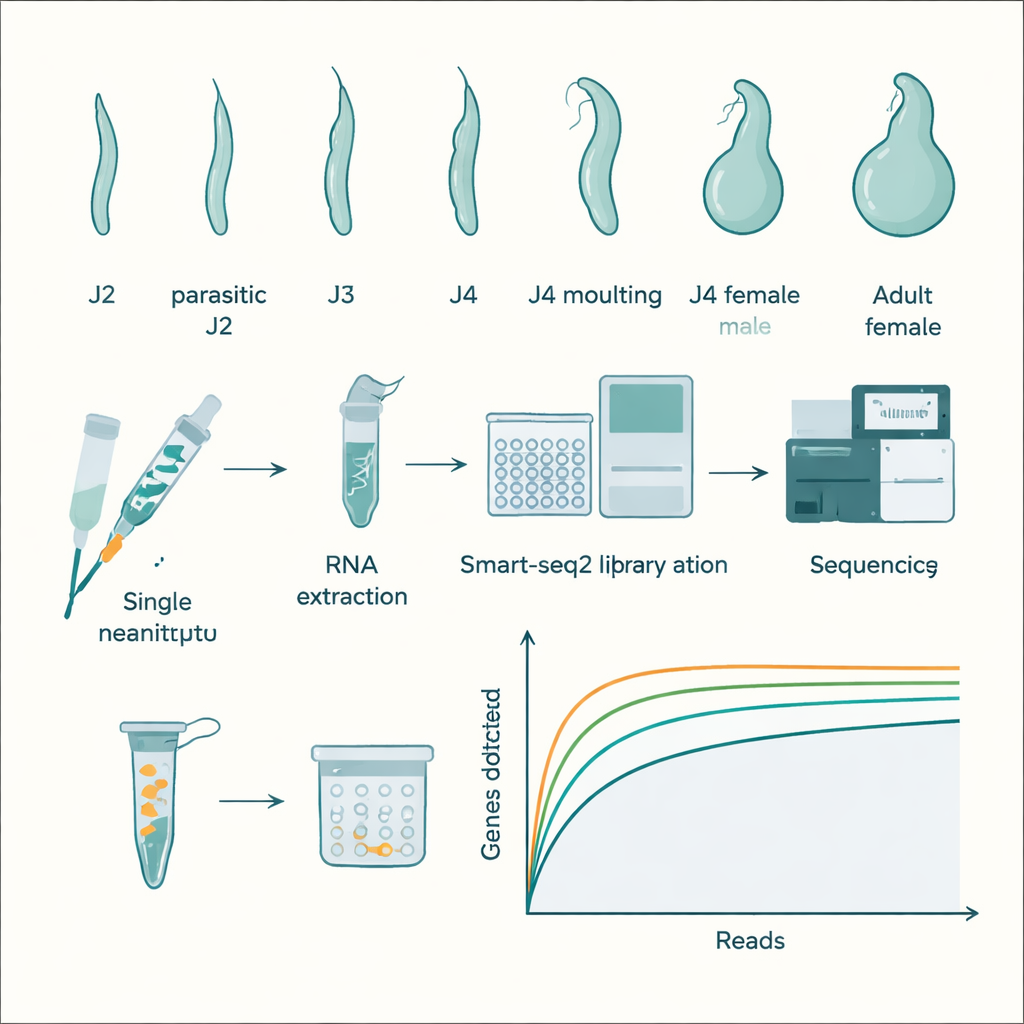
Worm Life Inside a Root
Meloidogyne incognita has a surprisingly complex life for a creature you can barely see. It hatches from an egg as a second‑stage juvenile, a slender, mobile form that seeks out plant roots. These juveniles pierce root cells with a needle‑like mouthpart and release enzymes that soften the plant cell wall, allowing them to burrow toward the plant’s water‑ and nutrient‑conducting tissues. Once they reach this inner region, they force the plant to create giant, nutrient‑rich feeding cells that serve as permanent food sources. Under good conditions, most worms grow into swollen, sedentary females that remain in the root and produce eggs, while some develop into slimmer males that leave the root and stop feeding. This rapid, flexible life cycle makes the nematode a persistent agricultural problem.
Why Gene Activity Matters
To control these pests without relying solely on chemical pesticides, scientists want to know which nematode genes switch on at each stage of infection. These genes may include the tools that help the worm invade roots, reprogram plant cells, or survive stress. However, collecting enough worms at exactly the same stage inside a plant has been technically difficult, especially for the middle juvenile phases. As a result, past studies often lumped different stages together, blurring the genetic signals that distinguish invasion, growth, molting, and sex differentiation. The current study tackles this challenge by analyzing one nematode at a time across eight carefully defined stages, from pre‑invasion juveniles raised in the lab to parasitic juveniles and multiple juvenile and adult forms within the plant root.
Reading the Genes of Single Worms
The researchers grew root‑knot nematodes on tobacco plants in a greenhouse and collected roots at different times after infection. They gently ground the roots, filtered the material through fine sieves, and then picked individual worms under a microscope based on their shape and size: pre‑invasion juveniles, root‑invading juveniles, third‑ and fourth‑stage juveniles, molting juveniles, and developing males and females, plus fully formed adult females. Each single worm was placed into a tiny tube, broken open in a special lysis solution, and its fragile RNA—the molecule that reflects which genes are active—was converted into stable DNA using a sensitive method called Smart‑seq2. These DNA copies were then amplified, turned into sequencing libraries, and read on a high‑throughput Illumina machine, generating billions of base pairs of data.
Checking Quality and Patterns
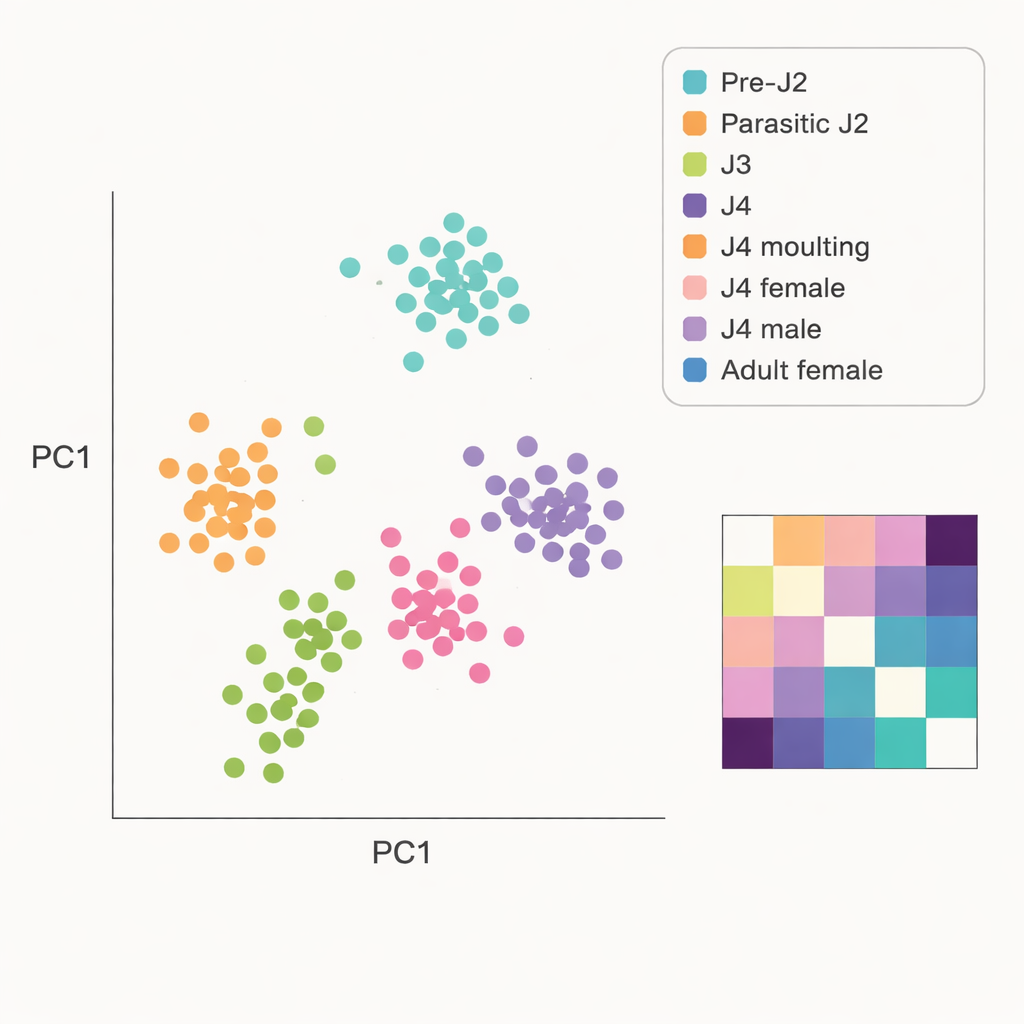
From this process, the team produced 75 high‑quality gene expression profiles, with at least five separate worms measured for each stage. They showed that around 10 million sequencing reads per worm were enough to capture most of the active genes, and that the data passed standard quality checks. Using statistical tools, they compared worms from different stages and found that their gene activity clustered by developmental phase: early juveniles grouped together, as did later juveniles, males, and adult females. Some neighboring stages, such as the third and fourth juveniles, blended smoothly into one another, reflecting the gradual nature of development. The researchers also counted how many genes were active at each stage, revealing shifts in the overall complexity of gene expression as the nematode moves from invasion, through growth and molting, to reproduction.
A New Map for Smarter Control
This study does not test new pesticides or resistant crops directly, but it provides a powerful foundation. By making all raw and processed data freely available in public databases, the authors offer the community a stage‑by‑stage map of which nematode genes turn on as the worm invades and exploits its host. Future researchers can mine this resource to pinpoint genes that are essential only at key stages—such as invasion or feeding—and then design targeted strategies to block them, from breeding plants that interfere with these genes to developing precise biological or chemical treatments. In simple terms, this work gives agriculture a detailed playbook of the nematode’s moves, opening the door to more effective and environmentally friendly ways to protect crops.
Citation: Han, X., Yang, Y., Wang, F. et al. High-resolution single-nematode transcriptomic datasets of plant-parasitic nematodes from juveniles to adults. Sci Data 13, 273 (2026). https://doi.org/10.1038/s41597-026-06599-4
Keywords: root-knot nematode, plant-parasitic nematode, RNA sequencing, crop protection, Meloidogyne incognita