Clear Sky Science · en
Semantic alignment of the German Human Genome-Phenome Archive metadata model in Europe’s genomics field
Why sharing genome data needs more than just files
Modern medicine increasingly relies on reading our DNA to diagnose diseases and tailor treatments. But the real power of genomics comes when data from many hospitals and countries can be combined. That only works if each dataset is described in a clear, compatible way and if privacy laws like Europe’s GDPR are strictly respected. This article explains how the German Human Genome-Phenome Archive (GHGA) is building a detailed "description system" for genomic studies so that valuable data can be found, understood, and safely shared across Europe.
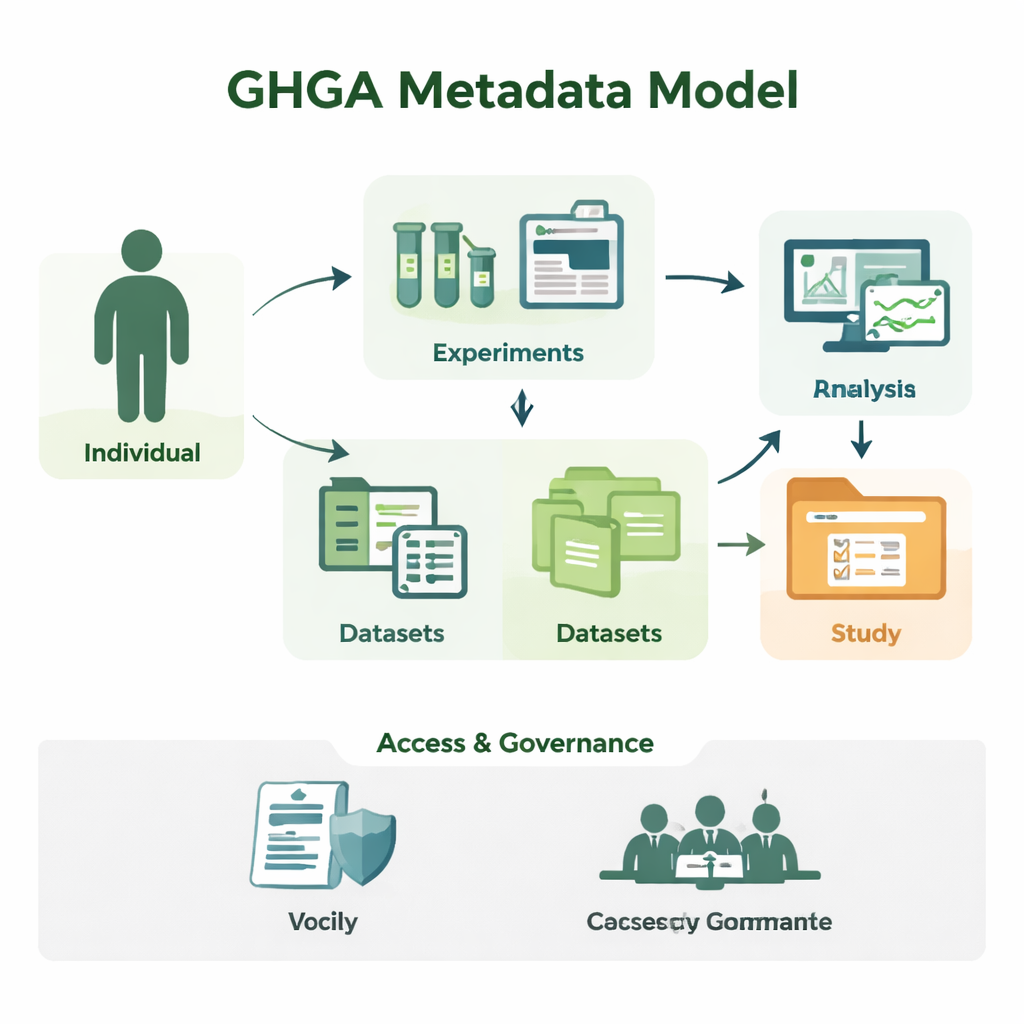
From raw sequences to understandable studies
Genomic research produces huge amounts of sequence data, but on its own, a file of DNA letters is meaningless. Researchers need to know who the sample came from, which tissue was used, how the experiment was done, and under what conditions they may reuse the data. GHGA captures this surrounding information as metadata. Its model arranges metadata into 16 building blocks, such as the person taking part in the study (the "Individual"), the sample taken, the experiment and analysis performed, the data files created, and the datasets and studies that bundle them. By separating scientific details from administrative ones like access conditions, the model mirrors how a real laboratory and data portal work, but in a way that computers can reliably process.
Keeping data useful but people unidentifiable
Because GHGA handles sensitive human health data, the team had to design the model to be scientifically rich without making it easy to identify any person behind the data. European GDPR rules say that information that could reasonably be linked back to an individual counts as personal data, even if names are removed. The article describes a careful privacy analysis that showed how combining details like age, postcode, and rare diagnoses can reveal identities. In response, GHGA’s public portal avoids fine-grained location data, groups ages into broad bands instead of exact years, and merges detailed diagnosis codes into coarser categories. This way, researchers still see whether a dataset may be relevant for their work, while the effort required to single out a person becomes unrealistic.
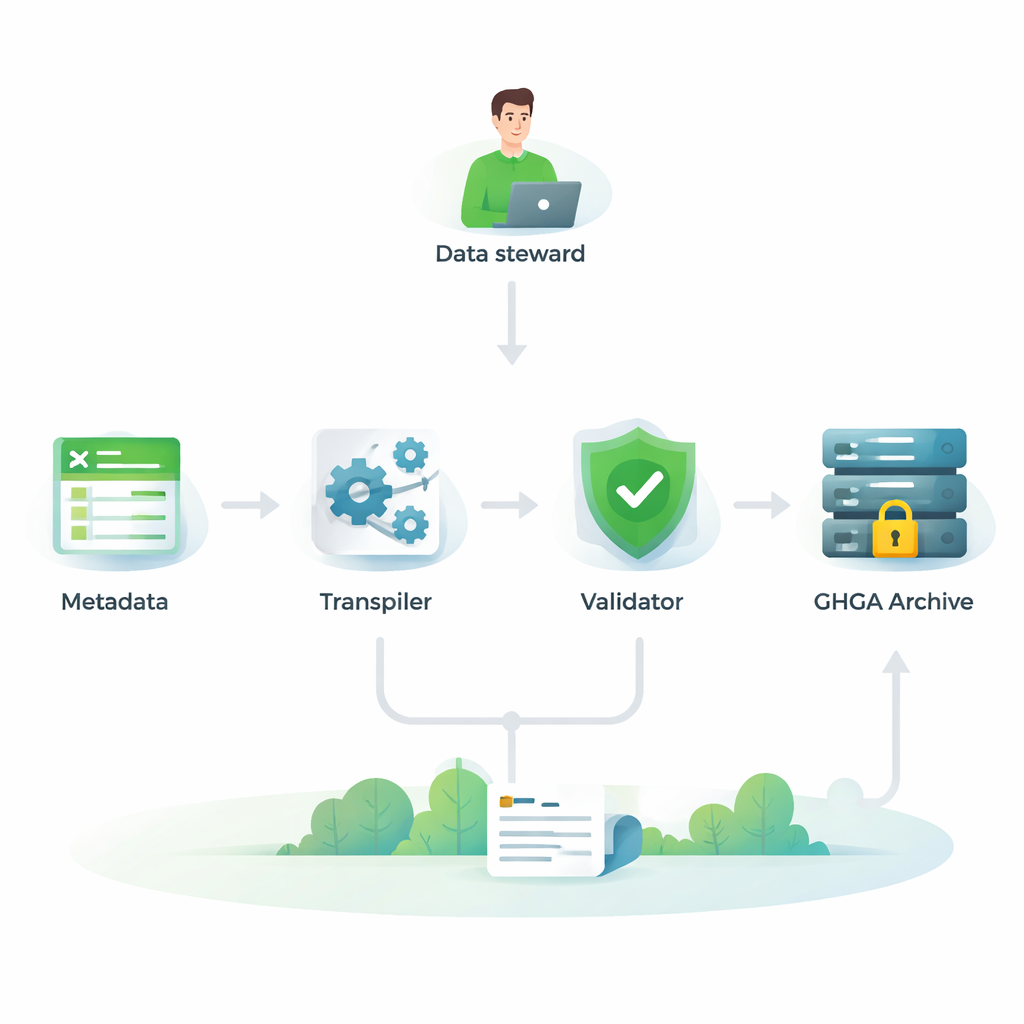
Checking compatibility with Europe’s genomics ecosystem
To be truly useful, GHGA’s metadata must fit into a wider European network of genomic archives and tools. The authors therefore compared their model, item by item, with four other widely used frameworks: two versions of the European Genome-phenome Archive (EGA), the ISA-tab standard, and the FAIR Genomes model from Dutch healthcare. They carried out a detailed “crosswalk” that asked, for each GHGA field, whether there was an equivalent in the other models and vice versa. They found that most of GHGA’s key properties have clear counterparts elsewhere, especially for describing studies, samples, experiments, analyses, and file formats. This means GHGA datasets can be understood and integrated alongside data stored in other European systems.
Finding common ground – and what is still missing
From this comparison, the team extracted 25 "consensus" metadata fields that appear in at least three of the five models. These cover essentials such as the sex and health status of participants, the tissue used, the type of sequencing and instrument, the analysis method, file formats, and basic study descriptions and contact details. These shared fields line up with existing minimum reporting guidelines and can serve as a core checklist for anyone designing new genomic data portals. At the same time, the analysis revealed information that some models collect but GHGA currently omits or only accepts in flexible, free-text form, such as the exact dates of sampling and sequencing, excluded diagnoses, and detailed contact names. Many of these omissions are deliberate trade-offs in favor of privacy and anonymity.
What this means for future health research
Overall, the study shows that GHGA’s metadata model is detailed, flexible, and closely aligned with international practice, while still staying within strict European privacy rules. It already covers all fields that other archives treat as mandatory, and it can be extended to new technologies such as single-cell and spatial omics. By offering a clear way to describe who and what a genomic study involves, how the data were produced, and under which conditions they can be reused, GHGA helps turn isolated data silos into a connected research resource. For patients, this improves the chances that their data, once donated, can safely contribute to discoveries and better treatments across borders for years to come.
Citation: Mauer, K., Iyappan, A., Parker, S. et al. Semantic alignment of the German Human Genome-Phenome Archive metadata model in Europe’s genomics field. Sci Data 13, 242 (2026). https://doi.org/10.1038/s41597-026-06575-y
Keywords: genomic data sharing, metadata standards, privacy and GDPR, GHGA, personalized medicine