Clear Sky Science · en
Metabarcoding and metagenomic data across aquatic environmental gradients along the coasts of France and Chile
Hidden Life in Shifting Seas
Along the world’s coasts, from tranquil lagoons to dramatic fjords, microscopic life is constantly adapting to changing conditions. These tiny organisms help drive the cycles of carbon and nutrients that underpin fisheries, water quality, and even climate. Yet many coastal waters, especially complex places like lagoons and fjords, have barely been explored at the genetic level. This study set out to change that by creating a rich, open dataset of coastal microbes from France and Chile, offering a new window into how marine life responds to an ever-changing environment.
Coastlines as Natural Test Labs
Coastal waters are rarely steady. Rainstorms, river inflow, evaporation, and tides can swing temperature and salt levels over short distances and time spans. Nutrients that feed microscopic algae can spike or crash, and human activities add further disruption. Such variation creates a patchwork of habitats that select for different microbial communities and encourage new adaptations. To capture this complexity, the researchers sampled 26 locations along the French and Chilean coasts, including lagoons, estuaries, fjords, harbors, beaches, nearshore waters, and one offshore site. Some French sites were visited monthly over a year to track the seasons, while Chilean sites were sampled in the southern autumn, providing a broad snapshot of contrasting coastal worlds. 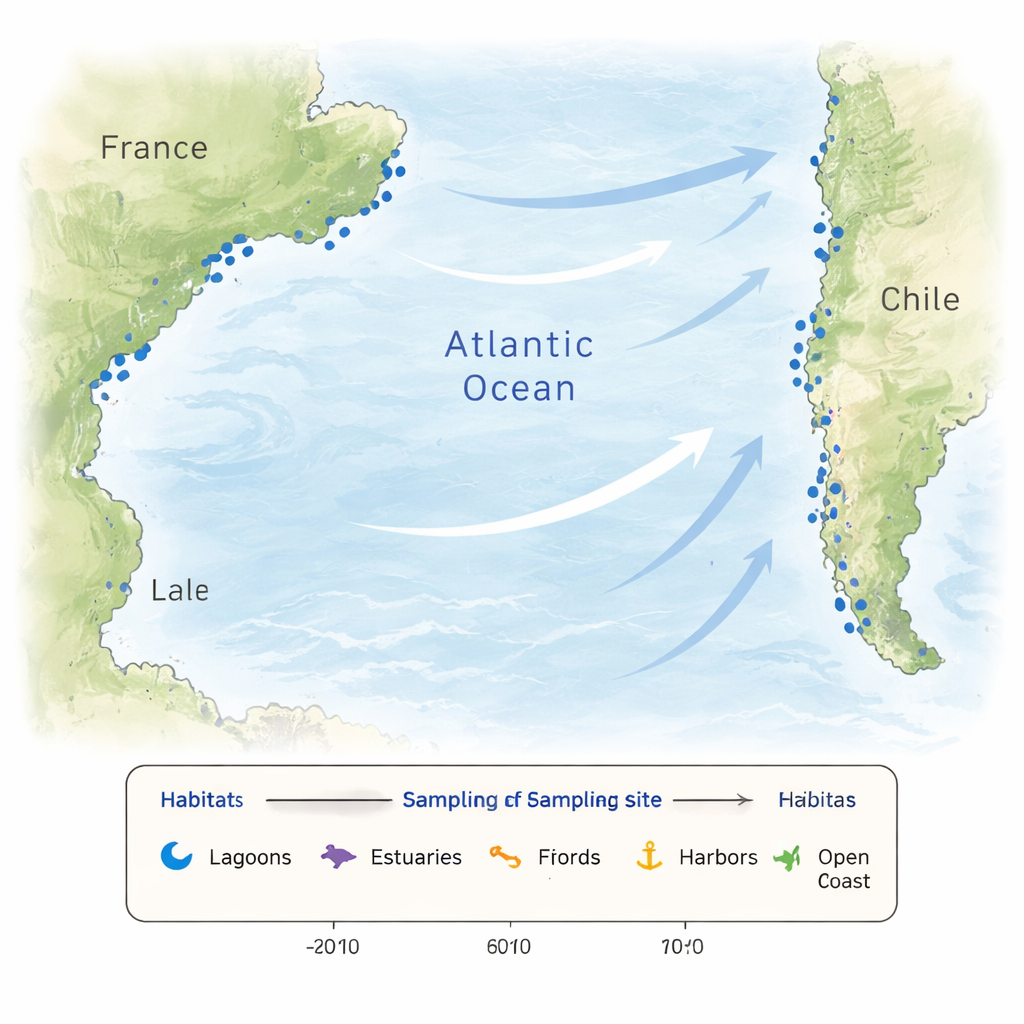
From Buckets of Water to DNA Fingerprints
At each site, the team collected large volumes of seawater, mostly from the sunlit surface but also from deeper layers in selected fjords and offshore waters. They measured basic environmental conditions such as temperature, salinity, and nutrients, alongside indicators of biological activity like chlorophyll (a proxy for algae), dissolved organic matter, bacterial production, and respiration. Back in the lab, microbes were concentrated on fine filters, and their DNA was extracted. One set of tests focused on a standard marker gene (16S rRNA), which acts like a barcode for identifying bacteria and archaea. This metabarcoding approach revealed more than 53,000 distinct DNA variants and showed that some samples shared as few as three, underscoring how different neighboring communities can be.
Rebuilding Genomes from a Genetic Soup
The second line of analysis took a more ambitious route: shotgun metagenomics, in which all DNA in a sample is sequenced at once. Using advanced assembly and binning methods, the team reconstructed 1,372 draft genomes, known as metagenome-assembled genomes or MAGs. Many of these genomes could not be matched to known species, and only about 4% aligned with formally described microbial species. In some groups, such as certain bacteria and archaea adapted to unusual conditions, more than half of the predicted proteins had no known function. The researchers also built a massive gene catalogue of over 23 million non-redundant genes, finding that roughly 31% lacked matches in major reference databases. This indicates a deep reservoir of uncharacterized biology residing in coastal waters.
Extreme Habitats, Novel Tools
Some sites, especially hyper-salty lagoons in France, stood out as hotspots of genetic novelty. There, salinity levels swung from nearly normal seawater to more than twice as salty as the ocean in just a few months. These stressful conditions may favor so-called extremotolerant microbes equipped with enzymes that still function under high salt or temperature. Such enzymes are increasingly sought after for industrial uses in detergents, biofuels, pollution cleanup, and food processing. By tying together detailed environmental measurements with gene and genome data, this dataset helps pinpoint where such unusual organisms and biochemical tools are most likely to be found. 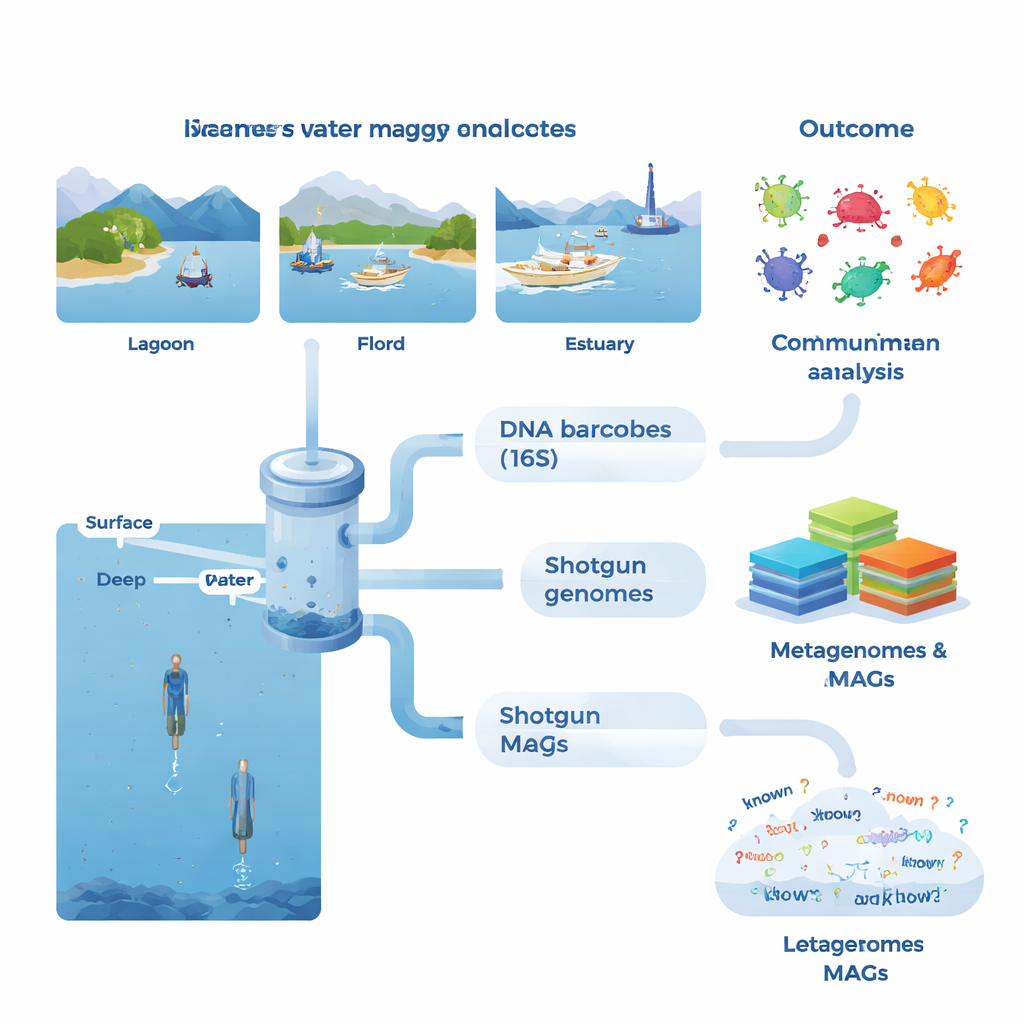
A Public Resource for the Ocean’s Future
Instead of presenting a single discovery, this work delivers a carefully validated, open resource that other scientists can mine. All DNA sequences, genome reconstructions, gene catalogues, and environmental measurements are publicly archived, along with the computer code used to process them. For non-specialists, the key message is that coastal microbes are both diverse and full of surprises, especially in overlooked environments like lagoons and fjords. As researchers tap into these data, they will be better equipped to understand how microscopic life along our shores responds to warming, pollution, and disturbance—and to harness novel genes that could benefit biotechnology and environmental management.
Citation: Maeke, M.D., Hassenrück, C., Aguilar-Muñoz, P. et al. Metabarcoding and metagenomic data across aquatic environmental gradients along the coasts of France and Chile. Sci Data 13, 29 (2026). https://doi.org/10.1038/s41597-026-06572-1
Keywords: coastal microbiomes, metagenomics, lagoons and fjords, marine biodiversity, environmental DNA