Clear Sky Science · en
Human brain MRI data of intrathecally injected tracer evolution over 72 hours for data-integrated simulations
Why this brain fluid study matters
Our brains constantly bathe in a clear liquid called cerebrospinal fluid, which helps cushion, nourish, and possibly clean the brain. Scientists suspect that this fluid may also be a powerful route for delivering drugs and removing waste linked to diseases such as Alzheimer’s and Parkinson’s. Yet watching how substances actually move through this fluid in a living human brain has been extremely difficult. This article presents a rare, open dataset that captures how a harmless tracer spreads through one person’s brain over three days, giving researchers worldwide a detailed playground for testing ideas and building computer models of brain fluid flow.
A single volunteer, many detailed scans
The dataset, nicknamed the “Gonzo” dataset, comes from an older healthy male volunteer who agreed not only to undergo an invasive imaging procedure but also to share all of his scans openly. A tiny dose of an MRI contrast agent was injected into the fluid-filled space around his spinal cord in the lower back. From there, the tracer mixed with the brain’s surrounding fluid and gradually entered brain tissue. The research team then scanned his head before the injection and at four later times over 72 hours, using multiple types of MRI. They also drew blood samples between scans to see how much tracer had entered the bloodstream. This combination of images and measurements lets scientists track when and where the tracer appears, and how fast it moves and clears.
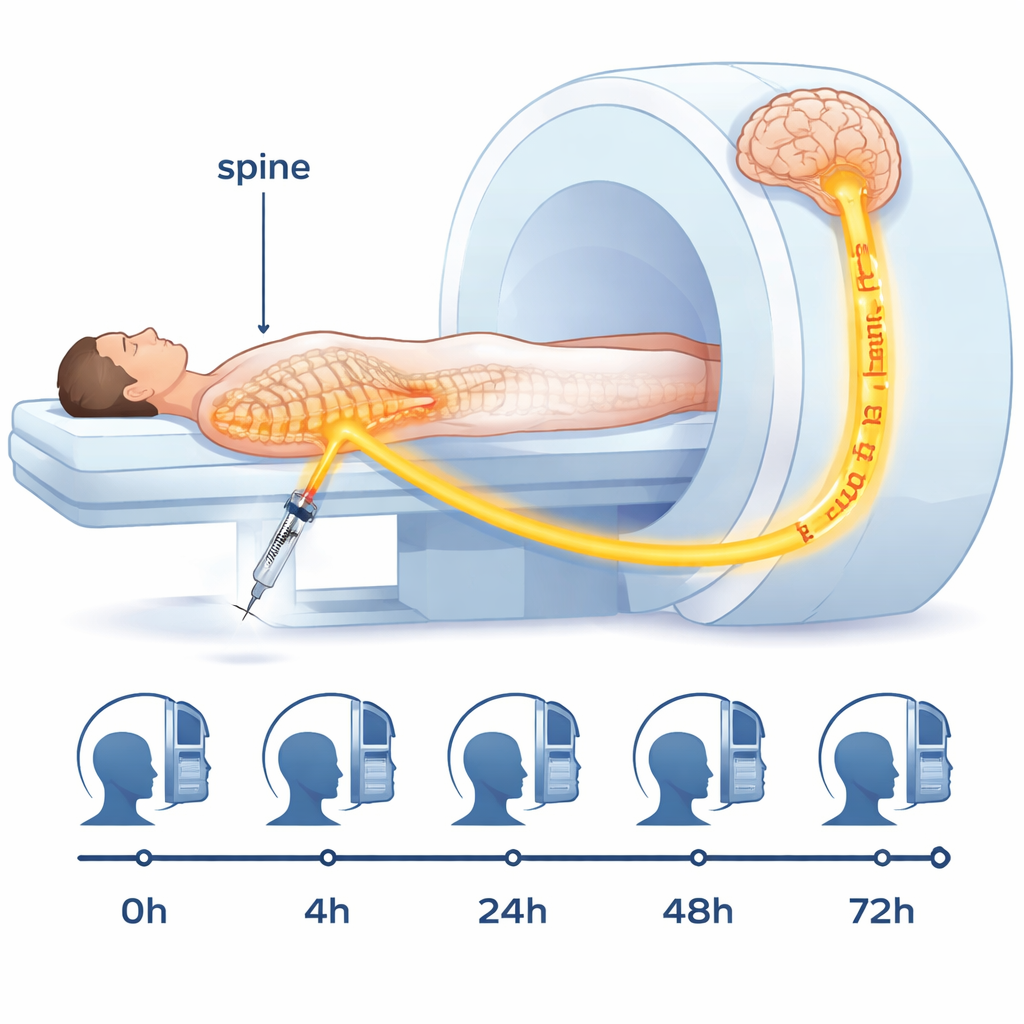
Turning raw images into usable brain maps
Modern MRI machines generate vast amounts of data, but to be useful for simulations and precise measurements, those raw images must be carefully processed. In this project, the team converted all scans into a common, well-documented file format and aligned them to the same three-dimensional reference frame so that every scan lines up with the same brain. They then used established software to segment the brain into regions, such as gray matter, white matter, and the fluid-filled spaces. From special MRI sequences, they calculated maps of physical properties like T1 relaxation time and diffusion, which are sensitive to how much tracer is present and how water moves in tissue. These steps turn fuzzy black-and-white pictures into precise, quantitative maps that can feed directly into mathematical and computer models.
Following the tracer through brain fluid and tissue
Using these processed maps, the authors estimated the tracer concentration in each tiny volume of the brain and surrounding fluid at each time point. Early on, most tracer stays in the fluid spaces that wrap around the brain, but over the first day it spreads more widely and enters the tissue itself. After 24 hours, almost half of the injected tracer is found in the head, divided fairly evenly between brain tissue and surrounding fluid. By 48 and 70 hours, the total amount has started to decline and becomes more evenly distributed, reflecting both clearance out of the brain and continued mixing. The team also extracted measurements of how easily water diffuses through different tissues, which helps characterize the microscopic structure of white and gray matter and may influence how substances spread.
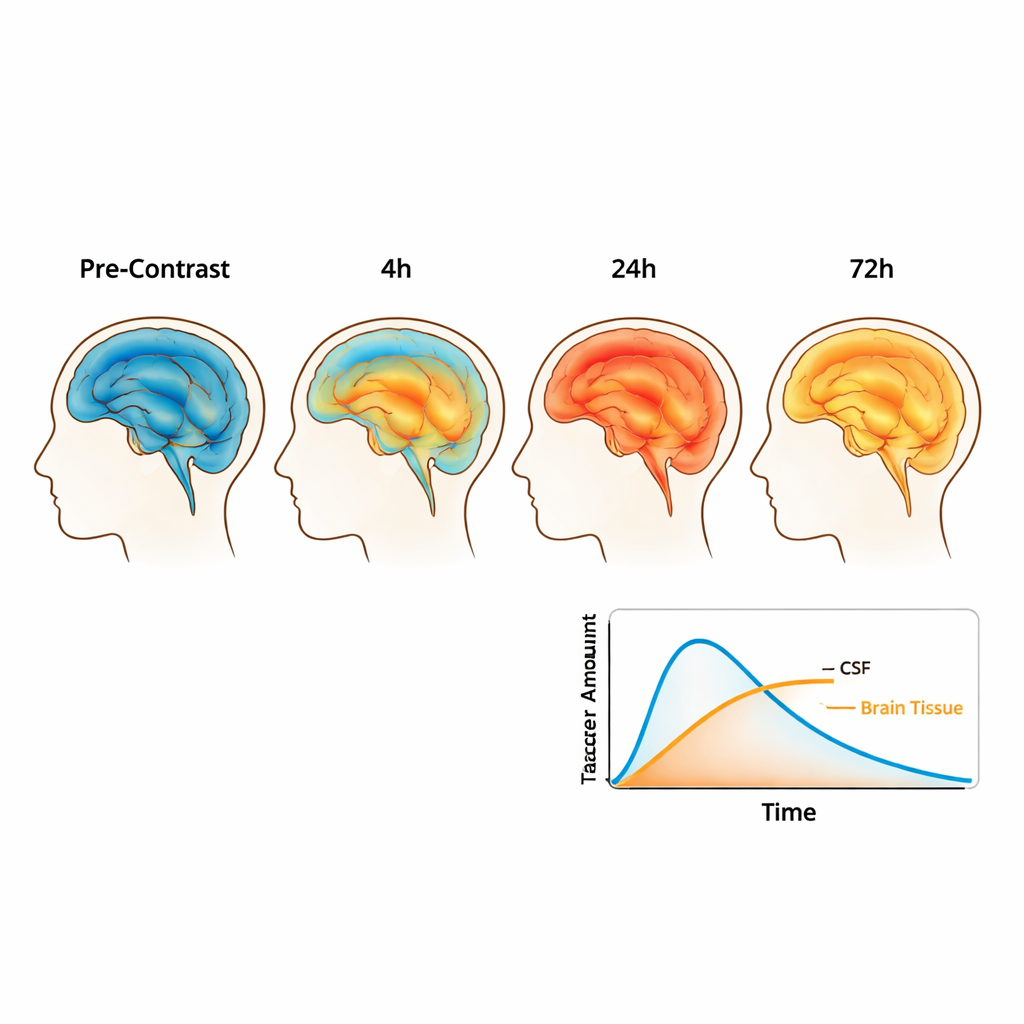
Building a 3D brain model for simulations
Beyond images, the study provides ready-to-use three-dimensional computer models of the volunteer’s brain. The researchers built detailed meshes—networks of small tetrahedral elements—that approximate the shape of the brain, its fluid spaces, and key internal structures. They then mapped tracer concentrations and diffusion properties from MRI onto this mesh. This allows engineers and mathematicians to run realistic simulations of how molecules move through brain tissue and along fluid pathways, test competing theories about brain “cleaning” mechanisms, and design new analysis methods, all without having to redo the heavy image-processing work. The dataset is organized into several downloadable bundles, from raw scans to fully prepared meshes, so users can choose the level that fits their expertise.
What this means for future brain research
The authors are clear that data from a single person cannot answer medical questions or support broad statistical claims about disease. People’s brain fluid flow patterns vary widely, so this dataset is best seen as a high-quality testbed rather than a population sample. Its true value lies in giving researchers a common, openly available reference case: a deeply characterized human brain with tracer evolution mapped over time. By making every processing step transparent and sharing code alongside the data, the study lowers the barrier for others to develop and validate models of brain fluid transport. In the long run, such models may help clarify how the brain clears waste, how that process fails in disease, and how we might better deliver drugs directly via the brain’s own fluid highways.
Citation: Riseth, J.N., Koch, T., Lian, S.L. et al. Human brain MRI data of intrathecally injected tracer evolution over 72 hours for data-integrated simulations. Sci Data 13, 245 (2026). https://doi.org/10.1038/s41597-026-06564-1
Keywords: cerebrospinal fluid, glymphatic system, brain MRI, tracer transport, brain simulation