Clear Sky Science · en
SEA CDM: Study-Experiment-Assay Common Data Model and Databases for Cross-Domain Data Integration and Analysis
Why organizing lab data matters to all of us
Modern medicine is powered by mountains of experimental data—from vaccine trials and infection studies to cancer genomics. Yet these data are often locked away in incompatible formats, making it hard for scientists to combine results and spot important patterns, such as who responds best to a vaccine or why some people have more side effects. This article describes a new way to organize and connect diverse biomedical experiments so that researchers can ask richer questions and get faster, more reliable answers that ultimately influence how we prevent and treat disease.
A common language for experiments
Different research groups and databases tend to describe their studies in their own way, even when they are doing very similar kinds of work. One database might focus on vaccine trials, another on gene activity in single cells, and a third on clinical outcomes, each using different labels and structures. The Study–Experiment–Assay Common Data Model, or SEA CDM, offers a simple shared “grammar” for all these efforts. It breaks any biomedical project into three linked steps: the overall study that poses a question, the experiments carried out on people or animals, and the assays—such as blood tests or gene expression measurements—that generate results. Around these steps, the model also standardizes key elements such as who or what was studied, what samples were taken, which treatments were applied, and what analyses were done. 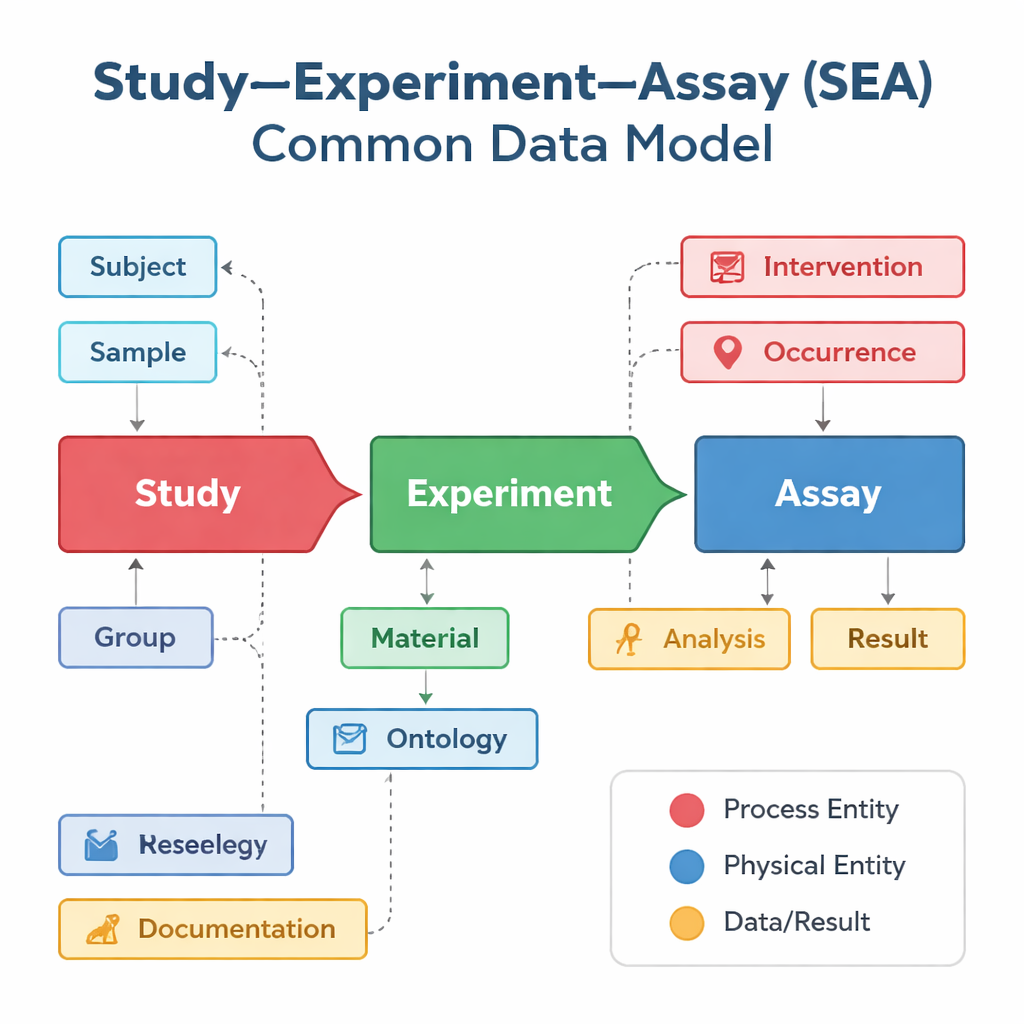
Ontologies: turning labels into knowledge
Simply lining up column headings is not enough; the same concept can be named differently in different places. SEA CDM leans on curated vocabularies known as ontologies to make sure that “flu shot,” “trivalent inactivated influenza vaccine,” and a brand name like “Fluzone” are all recognized as related ideas. These ontologies are structured like family trees of medical and biological terms. Because SEA CDM attaches an official identifier from an ontology to each variable—such as a disease, cell type, or vaccine—computers can automatically follow these trees, find all relevant records, and even infer relationships. For example, a short query can pull every study that used any trivalent influenza vaccine out of hundreds of named products, enabling powerful, semantic searches that go far beyond simple keyword matching. 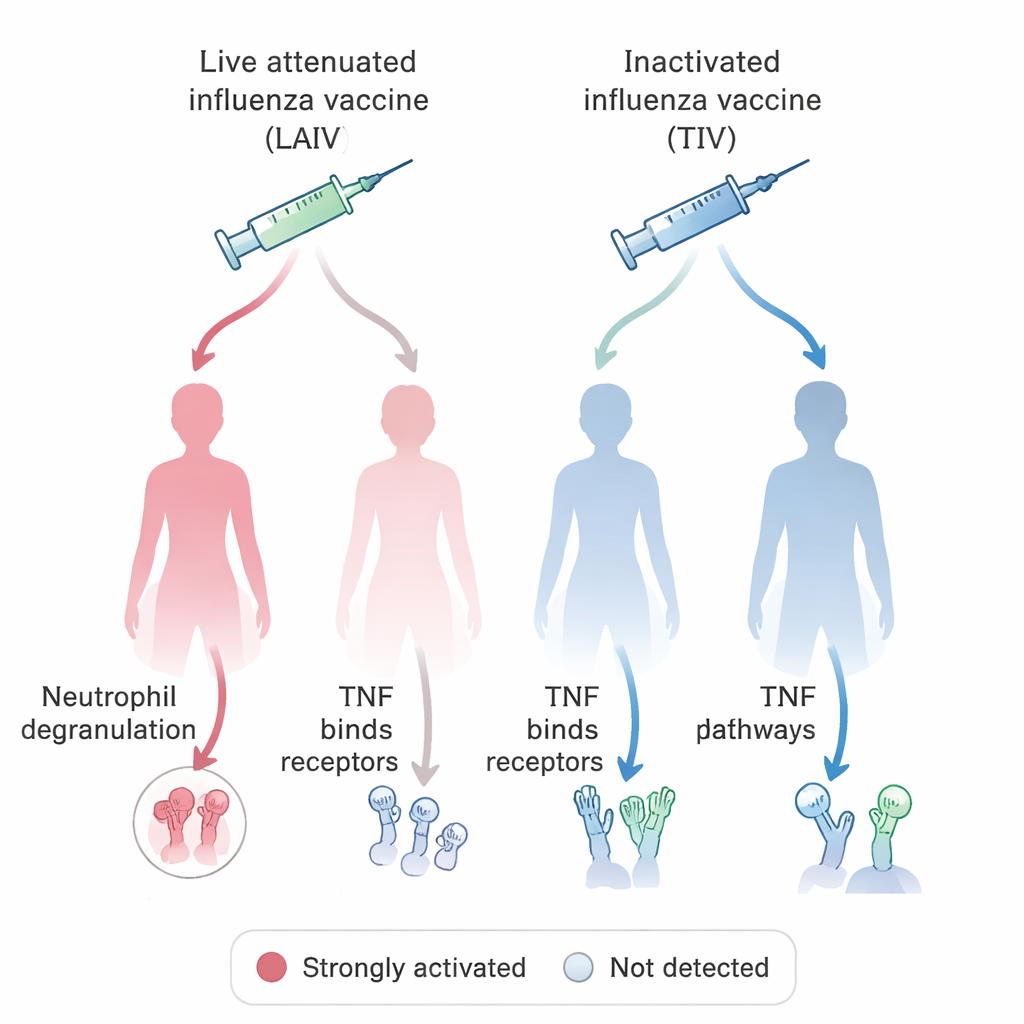
From scattered files to connected databases
To test their model in the real world, the authors built a family of databases and tools under the umbrella name OSEAN. They converted three large public resources into the SEA CDM structure: ImmPort, which hosts immune-response study metadata; VIGET, which links vaccine studies to gene activity data; and CELLxGENE, which focuses on single-cell measurements. Using custom pipelines, they translated dozens of original tables and file formats into a consistent set of SEA CDM tables or graph nodes. This allowed them to store more than a thousand immune-related studies, over two million samples, and numerous descriptions of vaccines, diseases, and lab methods in one coherent framework that can be searched with the same software.
What unified data can reveal about vaccines and sex differences
With this unified system in place, the team asked a biological question of direct medical relevance: how do different influenza vaccines stimulate the immune system in women and men? By querying the VIGET-based OSEAN database and applying simple rules for what counts as a “stimulated” gene, they identified hundreds of genes whose activity increased after vaccination with either live attenuated flu vaccines (containing weakened virus) or inactivated, “killed” vaccines. They then compared the pathways these genes participate in, separating the data by sex. One striking pattern involved neutrophils, a type of white blood cell that attacks microbes by releasing toxic granules, and signaling through TNF, a key inflammatory molecule. In most groups, influenza vaccination was linked to signs of neutrophil degranulation, but this signature was missing in women who received the live attenuated vaccine. In contrast, TNF-related signaling was especially prominent in these women but not in parallel male groups. These findings echo animal studies suggesting that neutrophil behavior and vaccine responses can differ systematically between males and females.
Building an ecosystem for future discoveries
The authors argue that the real power of SEA CDM lies in making biomedical data more FAIR—findable, accessible, interoperable, and reusable. By giving experiments a shared structure and anchoring every important label to a well-defined ontology term, their system makes it far easier to combine data from different sources, trace how samples were handled, and reproduce analyses. The influenza case study shows that even relatively simple queries, run over a harmonized database, can uncover subtle, sex-specific patterns in vaccine response that might influence dosing or vaccine choice. As more resources adopt this common model and the accompanying tools, researchers will be better equipped to connect clues across diseases, technologies, and populations, turning fragmented datasets into a true integrative biodata ecosystem.
Citation: Huffman, A., Yeh, FY., Hur, J. et al. SEA CDM: Study-Experiment-Assay Common Data Model and Databases for Cross-Domain Data Integration and Analysis. Sci Data 13, 238 (2026). https://doi.org/10.1038/s41597-026-06558-z
Keywords: data integration, biomedical ontology, vaccine response, sex differences, knowledge graph