Clear Sky Science · en
Comprehensive Transcriptome Profiling of Sporadic Medullary Thyroid Carcinomas
Why this rare thyroid cancer matters
Medullary thyroid carcinoma is a rare form of thyroid cancer that behaves more aggressively than most thyroid tumors and often strikes people when it is already advanced. Because it is uncommon, doctors and researchers have had limited data to understand why some tumors grow and spread faster than others, or why some patients respond better to treatment. This study tackles that gap by taking a deep, genome-wide look at the activity of thousands of genes in 72 tumors, creating one of the most complete molecular snapshots so far of this disease.
Looking inside tumor cells
Instead of focusing only on DNA mutations, the researchers examined the tumor “transcriptome” – the full set of RNA messages produced by active genes in cancer cells. These RNA messages act like real-time status reports of what the cell is doing. By sequencing RNA from fresh-frozen tumor samples, the team could see which genes were turned up, which were turned down, and how gene messages were stitched together. This approach helps reveal hidden patterns that classic genetic tests may miss, such as changes in how genes are regulated or combined.
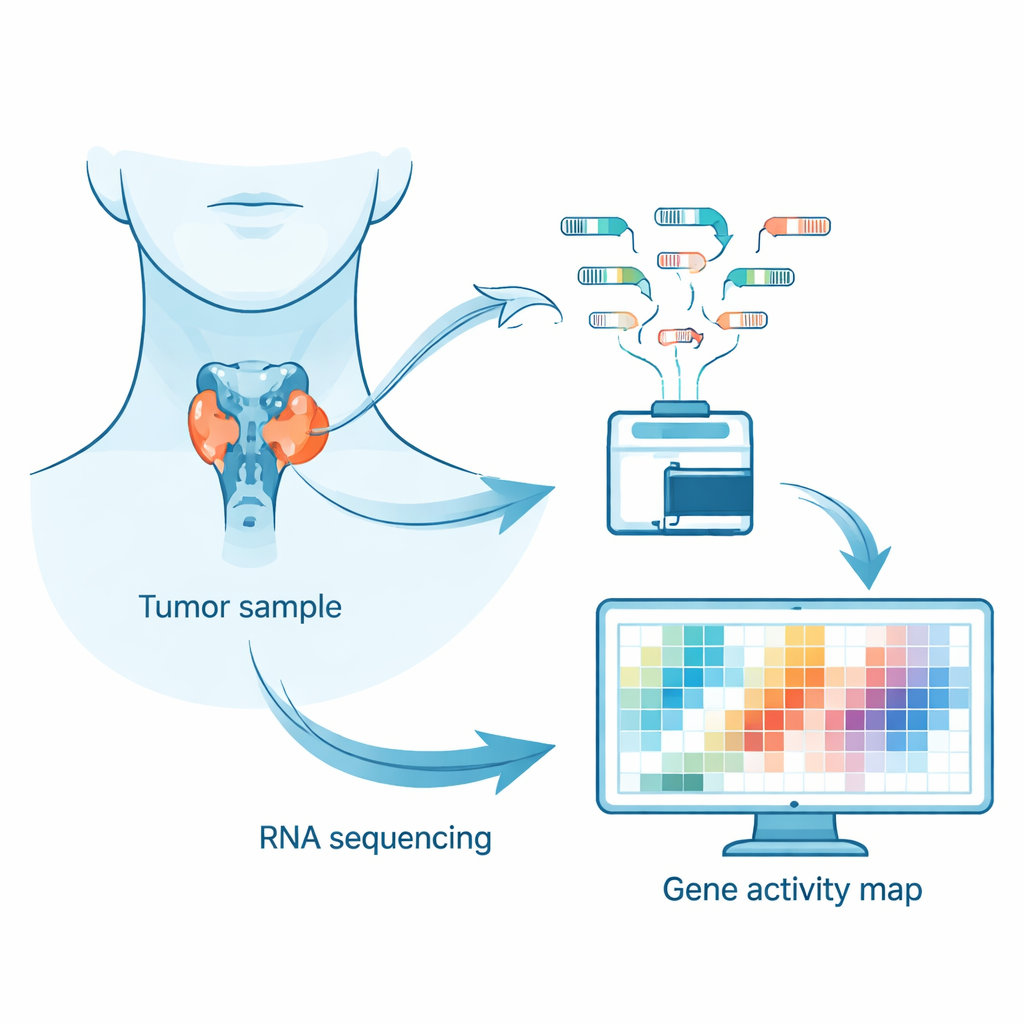
Who was studied and what was measured
The team analyzed tumors from 72 people with sporadic (non-familial) medullary thyroid carcinoma. Just over half were women, and the median age at surgery was 61. Around 58% of tumors carried changes in a gene called RET and about 24% had mutations in RAS family genes, both known drivers of this cancer. The scientists first checked that the tumor samples were of high quality and relatively free of normal thyroid tissue or immune cell contamination by looking at key markers such as calcitonin, which is high in these cancers, and thyroglobulin, which belongs to normal thyroid cells and stayed low. They then used well-established computational tools to align the RNA reads to the human genome, count gene activity, and search for differences between clinical and genetic groups.
Gene activity linked to stage and mutation
When the researchers compared early-stage tumors (stage I–II) with more advanced ones (stage III–IV), they found 760 genes whose activity levels clearly differed between the two groups. This means gene expression patterns carry strong clues about how far the cancer has progressed. They also contrasted tumors carrying RET or RAS mutations with those lacking both, uncovering over 1,200 genes with altered activity. A separate analysis zoomed in on tumors with a specific RET change, called M918T, which has long been tied to especially poor outcomes. In these tumors, 529 genes were differentially active compared with RET-normal cancers, and the RET gene itself was more highly expressed, underscoring how strongly this single change reshapes the cell’s behavior.
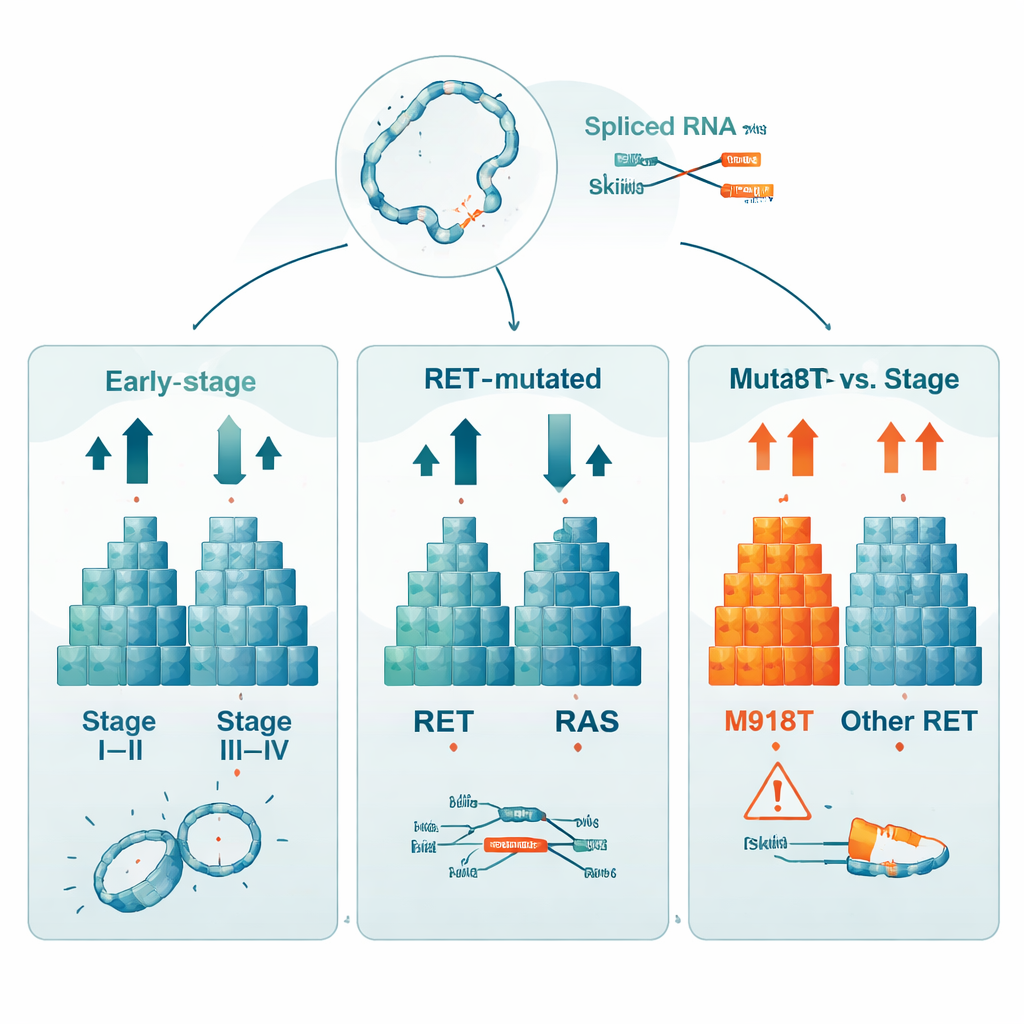
Hidden rearrangements and message rewiring
Beyond simple on–off changes in genes, the study searched for deeper forms of RNA disruption. The team detected 59 distinct gene fusion transcripts, where pieces of different genes become abnormally joined together at the RNA level. Most were rare, and the most frequent fusion may reflect a technical artifact, but the catalog shows that such rearrangements can be systematically mined from these data. The researchers also mapped thousands of alternative splicing events, where segments of gene messages are skipped, swapped or retained. While each clinical comparison (by stage, mutation type, or M918T status) showed similar overall counts of these splicing changes, the specific events largely differed, hinting at multiple, partly independent ways in which tumor cells rewire their gene messages.
What this resource offers patients and scientists
For patients, this work does not yet translate into an immediate new drug or test, but it lays essential groundwork. By making all of the processed RNA sequencing data publicly available, the authors provide a rich resource that other scientists can mine to discover new biomarkers to predict prognosis, refine how medullary thyroid cancers are classified, or spot weaknesses that could be targeted by future therapies, including the new generation of selective RET inhibitors. In simple terms, this study offers a detailed “activity map” of tumor genes across many real-world cases, giving researchers the tools they need to better understand why this rare cancer behaves so differently from person to person and how care might one day be tailored more precisely.
Citation: Palumbo, D., Melone, V., Palo, L. et al. Comprehensive Transcriptome Profiling of Sporadic Medullary Thyroid Carcinomas. Sci Data 13, 234 (2026). https://doi.org/10.1038/s41597-026-06550-7
Keywords: medullary thyroid carcinoma, cancer genomics, RNA sequencing, RET mutation, personalized oncology