Clear Sky Science · en
Transposable element–gene chimera cartography, origination and role in enhancing transcriptome plasticity
Hidden Passengers in Our DNA
Much of our DNA is made of ancient viral stowaways called transposable elements—bits of genetic code that once jumped around genomes like hitchhiking parasites. This study shows that these old viral fragments are not just silent junk: they can splice into our own genes to create hybrid messages, help cells adapt to changing conditions, and sometimes fuel disease. By mapping these hybrids across development, aging, and cancer, the authors reveal a surprising way in which viral DNA continues to shape human biology today.
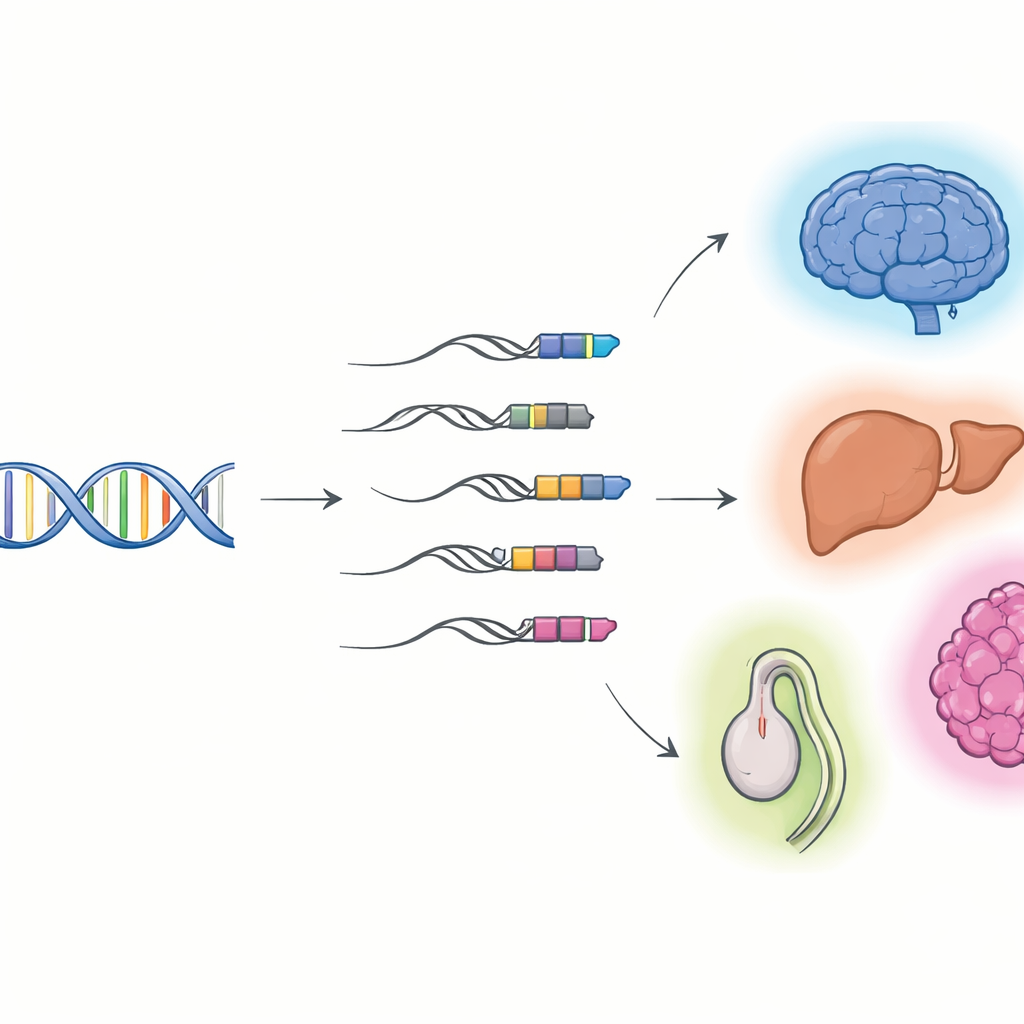
When Viral DNA Becomes Part of Our Genes
The researchers focused on “TE–gene chimeras”: RNA molecules that begin in a transposable element and then continue into a host gene. Using long-read sequencing, which can capture entire RNA molecules end to end, combined with standard short-read sequencing, they built detailed catalogs of these hybrids in mouse and human cells and organs. They found thousands of previously unknown chimeric transcripts, most of them noncoding RNAs rather than protein blueprints. Many start in a class of viral remnants called long terminal repeats (LTRs), which often sit at the front of genes and can act like alternative on–off switches. Different tissues and organs use different sets of these chimeras, with especially rich activity in the brain, liver, and testis, and their usage shifts as tissues develop from fetal to adult stages.
Chimeras in Aging and Cancer
To see how these viral–host hybrids behave in real populations, the team mined large human datasets covering dozens of tissues from hundreds of people, as well as tumor samples. They identified several hundred high-confidence chimeras that recur across individuals. Their expression varies not only between organs but also with age: in tissues such as blood and brain, some LTR-driven chimeras decline in older adults, while in certain peripheral tissues they increase. In many types of cancer, these LTR-based hybrids are broadly turned up in tumors compared with nearby normal tissue, and patients whose tumors express more of them tend to fare worse. In datasets from people resistant to chemotherapy or immunotherapy, the genes affected by new LTR-driven exons often sit in pathways linked to how those drugs work, suggesting that TE chimeras may help tumors rewire themselves under treatment pressure.
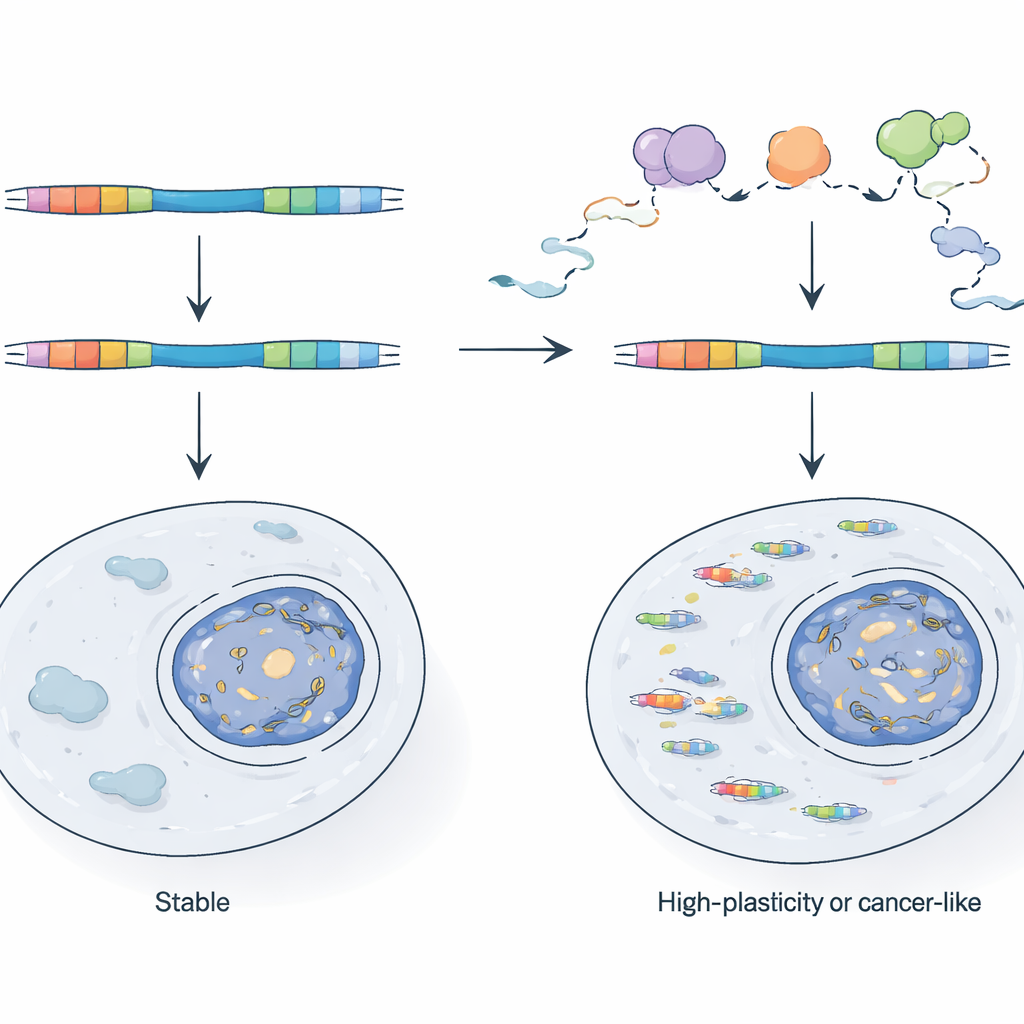
How Cells Keep Viral Hybrids in Check
Given their potency, why don’t TE chimeras overwhelm normal cells? The authors show that cells rely on an RNA quality-control system, centered on the nuclear exosome complex, to patrol and destroy many of these transcripts as they are being made. When they genetically cripple a key exosome component in mouse stem cells, LTR-driven promoters become far more active and chimeric RNAs surge. By comparing newly made RNA with stable RNA, they show that some LTRs are normally held in check because their transcripts are rapidly degraded, while others are controlled at the level of how often transcription starts. These active LTRs tend to sit in genome regions that are already accessible and frequently transcribed, indicating that simply being near an active gene or antisense transcript can help a viral fragment escape traditional epigenetic silencing.
Splicing, Stem-Cell Plasticity, and a Viral “Master Switch”
The story becomes more intricate when the cell’s splicing machinery is perturbed. The authors knock down several core splicing factors or chemically inhibit the spliceosome and observe a shared outcome with exosome loss: a strong boost in LTR-driven chimeras and in short, intron-poor genes. Among the winners is MERVL, an endogenous retrovirus active in very early mouse embryos. When MERVL’s internal sequence and its downstream transcription factors are upregulated, stem cells shift toward a more “totipotent-like” state, resembling early embryonic cells that can form all tissues. Conversely, degrading MERVL RNA with antisense oligonucleotides dampens this shift and reduces LTR promoter activity, indicating that MERVL acts as a master switch that can trigger a broader network of TE chimeras and cell-state plasticity.
Viral Innovation Across Evolution
Looking across many mammalian genomes, the team traces when different TE families began donating exons to host genes. In both humans and mice, relatively young LTR families are especially likely to form chimeras in the “sense” orientation that can be read as part of host transcripts. This pattern suggests that new waves of TE insertions have repeatedly fed raw material into gene regulatory networks over evolutionary time. Rather than being purely harmful, these insertions can be tested by the cell’s transcription and decay machinery: most are silenced or degraded, but a minority find themselves in the right place and context to be co-opted as useful regulatory elements or new gene starts.
Why These Viral Remnants Matter
To a non-specialist, this work shows that a large and once-dismissed portion of our genome is actively shaping how genes are read, how cells change identity, and how diseases progress. Ancient viral sequences can be turned into alternative gene “front doors,” producing hybrid RNAs that are normally policed by cellular cleanup systems. When that policing or normal splicing falters, these hybrids can surge, nudging stem cells into more flexible states or helping tumors adapt and resist therapy. Over millions of years, evolution has repeatedly tinkered with these viral fragments, turning many from genomic parasites into building blocks of regulatory complexity—and leaving us with a transcriptome far more plastic and dynamic than a simple list of genes would suggest.
Citation: Cheon, Y., Alvstad, E.G., Torre, D. et al. Transposable element–gene chimera cartography, origination and role in enhancing transcriptome plasticity. Nat Struct Mol Biol 33, 448–463 (2026). https://doi.org/10.1038/s41594-026-01757-z
Keywords: transposable elements, chimeric transcripts, RNA surveillance, stem cell plasticity, cancer evolution