Clear Sky Science · en
Structure and function of the yeast amino acid-sensing SEAC–EGOC supercomplex
How Cells Decide When to Grow
Every cell must constantly decide whether it can afford to grow or should conserve resources. That decision depends heavily on amino acids, the building blocks of proteins and a key source of nitrogen. This paper uncovers, in remarkable structural detail, how yeast cells sense amino acid availability through a giant molecular assembly and use that information to turn a major growth controller, called TORC1, on or off. Because the same circuitry is conserved in human cells and tied to cancer and metabolism, insights from yeast offer a window into how our own cells judge when conditions are right for growth.
An Intricate Hub on the Cell’s Recycling Center
In yeast, a central decision-making complex called SEAC sits on the membrane of the vacuole, an organelle that acts as the cell’s recycling center. There it meets another complex, EGOC, which relays information about amino acid levels. Together they feed signals to TORC1, the main growth switch. SEAC has two functional halves: SEACIT, which can shut down TORC1, and SEACAT, long thought to keep SEACIT in check. The authors used high-resolution cryo–electron microscopy to visualize the full SEAC bound to EGOC, capturing a snapshot of this molecular hub exactly where signaling happens, on the vacuole surface. 
Seeing the Molecular Architecture Up Close
The structure reveals SEAC as a large core made of SEACAT subunits, from which two flexible "wings" of SEACIT extend. Each wing can bind one EGOC complex, meaning a single SEAC can engage two signaling inputs at once. Strikingly, EGOC attaches only to the wings and not to the core, and the overall shape of SEAC barely changes when EGOC is present. This shows that SEACAT does not physically block the active site of SEACIT and does not prevent EGOC from binding. The orientation of the complex on the vacuole membrane also positions the SEAC core facing the cytoplasm, where it can interact with additional regulators, while the wings lie close to the membrane, ideally placed to contact EGOC and other membrane-anchored partners.
How SEAC Picks the Right Signal
The authors zoomed in on the contacts between SEAC and EGOC and found that SEAC recognizes only the "active" version of EGOC, in which one of its small GTPase proteins, Gtr1, carries a GTP-like nucleotide, while its partner, Gtr2, holds GDP. SEACIT provides a catalytic "finger" that reaches into Gtr1’s nucleotide pocket and accelerates the breakdown of GTP, flipping EGOC from an active to an inactive state. Detailed cryo-EM maps and mutational experiments show that this catalytic finger and a neighboring helper residue are essential for the interaction and for correctly reading the nucleotide state. When the authors altered these key residues, SEAC no longer localized properly to the vacuole, and EGOC could not engage productively.
Fast Versus Slow Control of the Growth Switch
To understand what this means for the living cell, the researchers tracked the activity of TORC1 over time when amino acids were removed and then reintroduced. In normal cells, TORC1 quickly turns off within a few minutes of nitrogen starvation and then rapidly turns back on when amino acids return. When the catalytic finger in SEACIT was disabled, or SEACIT subunits were deleted, TORC1 hardly responded in this early time window. The same was true when both Gtr1 and Gtr2 were removed, showing that the SEAC–EGOC pair is the dominant, fast pathway for amino acid control of TORC1. However, at later times—around half an hour—TORC1 activity slowly adjusted even in these mutants, indicating that a separate, slower route to TORC1, independent of this GTPase system, also operates in cells.
A Surprise Role for a Distant Subunit
A particularly intriguing finding concerns Sea2, a SEACAT component with a flexible "propeller" domain that juts out from the core. Removing Sea2, or just its propeller, produced signaling defects almost identical to those seen when SEAC’s catalytic activity was lost: TORC1 responded sluggishly to both amino acid withdrawal and refeeding. Yet Sea2 sits far from the catalytic site and does not directly touch the SEACIT wing, making a simple blocking mechanism unlikely. Genetic experiments combining Sea2 loss with a catalytically dead SEAC variant suggested that having the GAP activity locked fully on or fully off both disrupts the rapid response, whereas intermediate regulation is needed. The authors propose that Sea2’s propeller acts as a docking platform for an unknown factor that tempers SEACIT’s activity, allowing cells to fine-tune TORC1 rather than simply switching it permanently off.
What This Means for Cell Growth Decisions
Overall, the study recasts SEAC not as two opposing assemblies but as a single, integrated machine whose catalytic wing (SEACIT) and structural core (SEACAT) work together to sense amino acids and rapidly adjust the growth switch TORC1. SEACIT’s enzymatic action on EGOC provides the fast, precise control, while Sea2’s propeller likely recruits regulators that modulate this activity. At the same time, slower, parallel pathways ensure that TORC1 still eventually reflects nutrient status even if the primary sensor is disturbed. Because closely related complexes carry out the same job in human cells, these insights from yeast illuminate how our own cells balance growth with nutrient supply, with implications for understanding diseases where TORC1 signaling goes awry.
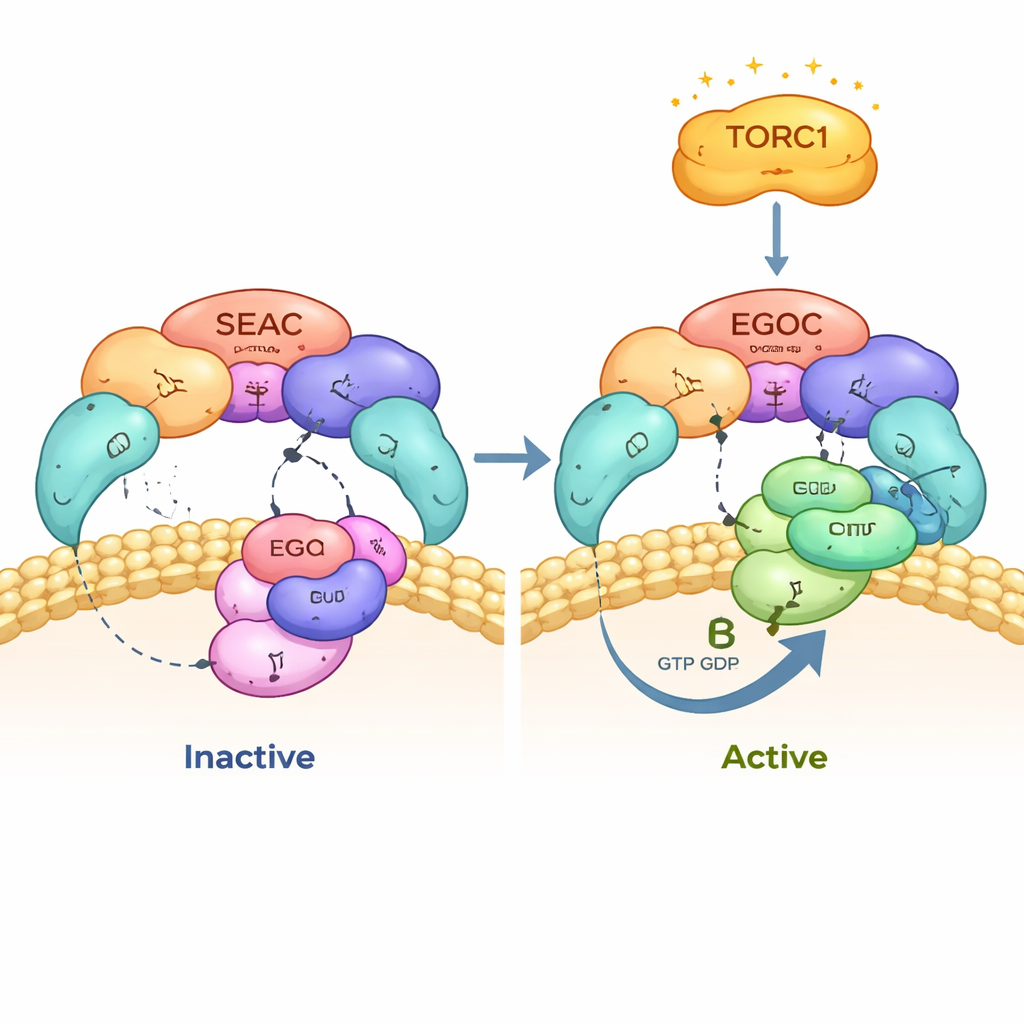
Citation: Tafur, L., Bonadei, L., Zheng, Y. et al. Structure and function of the yeast amino acid-sensing SEAC–EGOC supercomplex. Nat Struct Mol Biol 33, 488–498 (2026). https://doi.org/10.1038/s41594-026-01746-2
Keywords: amino acid sensing, TORC1 signaling, SEAC GATOR complex, cell growth control, cryo electron microscopy