Clear Sky Science · en
Mechanism and reconstitution of circadian transcription in cyanobacteria
Why body clocks matter, even for bacteria
All living things, from humans to tiny microbes, keep time. These internal clocks help organisms anticipate sunrise and sunset, scheduling activities like feeding, repair, and growth. This paper explores how a simple photosynthetic bacterium, Synechococcus elongatus, uses a protein-based clock to turn genes on and off with remarkable precision over a 24-hour cycle. By rebuilding this timing system in a test tube, the researchers reveal a stripped-down version of a biological clock that could inspire new tools for biotechnology and synthetic biology.
A tiny clock that runs on chemistry
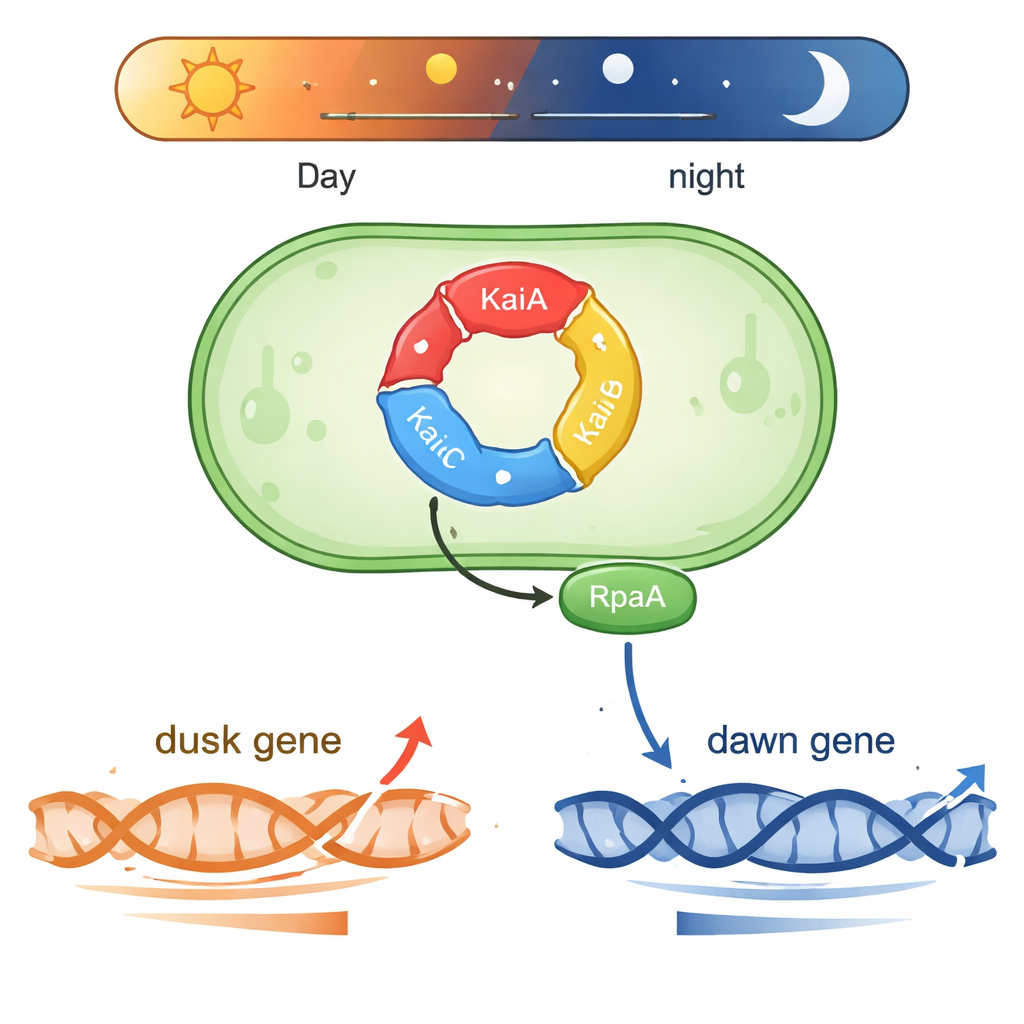
Synechococcus lives in sunlit waters and needs to predict daily changes in light. At its core is a three-part protein machine made of KaiA, KaiB, and KaiC. These proteins pass phosphate groups around in a repeating cycle that takes about 24 hours, forming a biochemical “gear train” that keeps time even outside the cell. Two other proteins, SasA and CikA, read the phosphorylation state of KaiC and use it to control another protein, RpaA. When RpaA is phosphorylated, it binds to DNA and acts as a master switch for hundreds of genes that rise and fall either around subjective “dawn” or “dusk” in constant light.
One protein, two opposite gene schedules
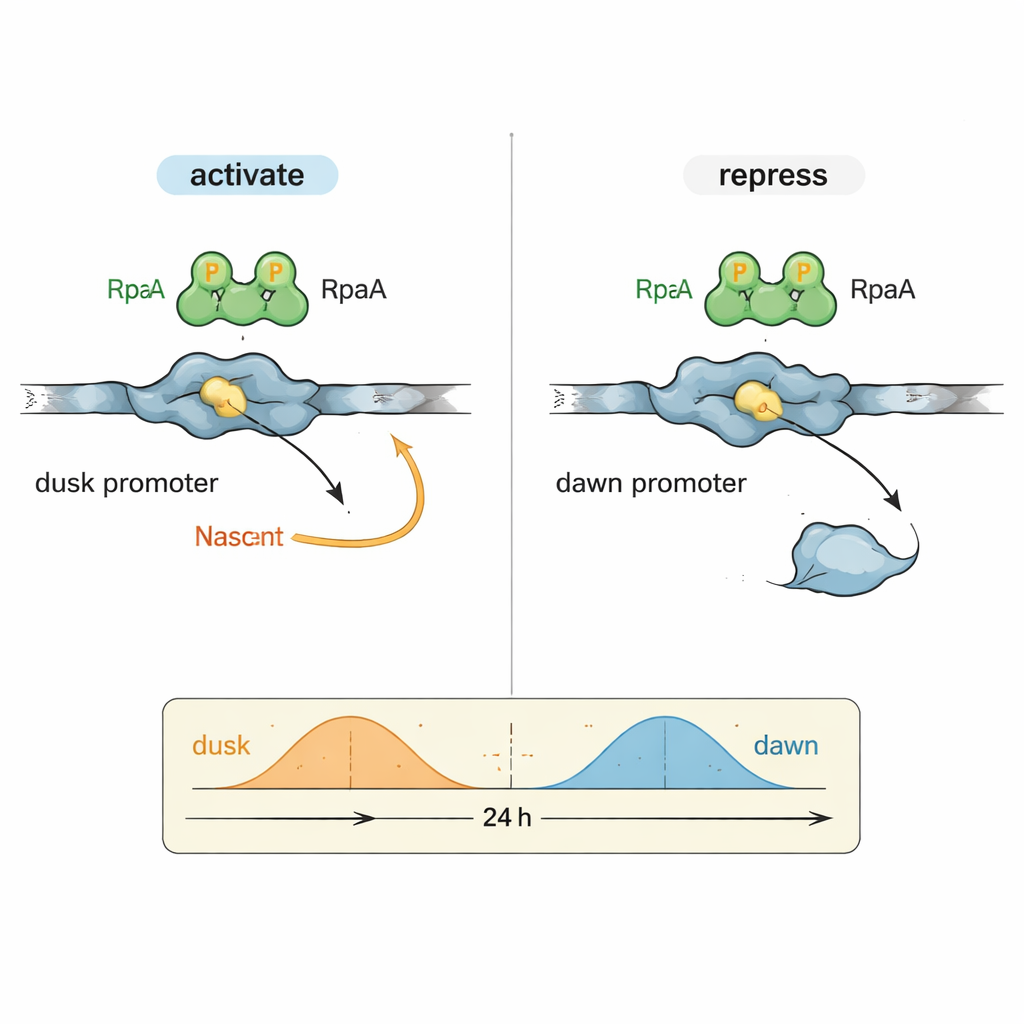
A puzzle in circadian biology has been how one output factor, RpaA, can coordinate genes that peak at very different times of day. The authors focused on two representative DNA control regions (promoters): kaiBC, which is most active at dusk, and purF, which peaks at dawn. In controlled reactions using purified cyanobacterial RNA polymerase and RpaA, they showed that phosphorylated RpaA boosts transcription from the kaiBC promoter while at the same time reducing transcription from the purF promoter. Detailed footprinting experiments mapped exactly where RpaA binds on each piece of DNA, revealing that its position relative to the standard promoter elements determines whether it acts as an accelerator or a brake.
Seeing the clock’s switch in atomic detail
To understand this dual behavior at the molecular level, the team used high-resolution cryo–electron microscopy to capture the structure of RpaA bound to the kaiBC promoter together with RNA polymerase. The images show RpaA gripping the DNA as an asymmetric pair and contacting two key parts of the enzyme: the alpha subunit’s tail and a region of the sigma factor that normally recognizes promoter sequences. These contacts bend the DNA and help position RNA polymerase at a slightly redefined start site for transcription. Carefully chosen mutations that weaken either of these contact points reduce or abolish activation of kaiBC, both in vitro and inside living cells, confirming that physical recruitment of RNA polymerase underlies the dusk-phase gene activation.
Building a clock-driven gene from scratch
Because the native cyanobacterial RNA polymerase is complex and hard to keep active for days, the researchers next turned to the much simpler T7 bacteriophage polymerase, widely used in laboratories. RpaA cannot recruit this unrelated enzyme, but it can still block it by sitting on the DNA. The team designed a synthetic DNA template where a T7 promoter drives a fluorescent RNA “Broccoli” reporter, with an RpaA binding site placed downstream. When they combined this template with the KaiA–KaiB–KaiC clock, the CikA kinase/phosphatase, RpaA, and T7 polymerase in a single optimized buffer, they observed waxing and waning transcription rates with a period of about 24 hours. The rhythm could be reset by changing the ATP-to-ADP ratio and ran with nearly the same period over a range of temperatures—classic hallmarks of a true circadian clock.
From simple clocks to engineered timekeeping
The study shows that only six proteins are sufficient to connect a chemical pacemaker to rhythmic gene readout: the three Kai clock proteins, CikA, RpaA, and an RNA polymerase. By changing where RpaA sits on DNA, the same molecule can make some genes peak at dusk and others at dawn, helping explain the complex waves of gene activity seen in cyanobacteria. Because the repression-based design works even with a foreign polymerase like T7, it should be possible to transplant this minimal clock module into other microbes or cell-free systems, allowing scientists to program genes to turn on and off with daily rhythms for research, industrial production, or future therapeutic applications.
Citation: Fang, M., Gu, Y., Leanca, M. et al. Mechanism and reconstitution of circadian transcription in cyanobacteria. Nat Struct Mol Biol 33, 275–281 (2026). https://doi.org/10.1038/s41594-025-01740-0
Keywords: circadian clock, cyanobacteria, transcription regulation, RpaA, synthetic biology