Clear Sky Science · en
Cereblon induces G3BP2 neosubstrate degradation using molecular surface mimicry
Turning Cellular Cleanup into a Smart Targeting System
Modern medicines increasingly try to fix disease not just by blocking troublesome proteins, but by getting rid of them altogether. This paper explores a clever way to rewire one of the cell’s own "cleanup" machines so that, when paired with a small molecule, it can recognize and destroy a protein linked to cancer and other disorders. The work reveals an unexpected trick: the cleanup machine reshapes part of its surface to imitate one of the protein’s natural partners, allowing it to latch on and send the protein to the cellular trash.
A Cell’s Garbage Collector with Hidden Flexibility
Our cells constantly tag worn-out or unwanted proteins for destruction, using a system built around E3 ligases—large protein complexes that decide what should be thrown away. One such complex relies on a component called cereblon, which acts like a sensor that recognizes specific features ("degrons") on other proteins. Certain approved drugs already exploit cereblon by acting as "molecular glues": they bind to cereblon and create a new docking surface that pulls in disease-related proteins, which are then tagged and degraded. Until now, most known targets used a particular structural pattern, meaning the range of proteins that could be removed in this way seemed limited.
Discovering a New Target Outside the Old Rules
In the new study, the authors screened a collection of cereblon-focused molecular glues in human cells and identified a compound called MRT-5702. This small molecule caused rapid loss of a protein named G3BP2, which helps manage RNA and forms part of stress granules—assemblies linked to cancer, heart problems and neurodegenerative disease. Importantly, G3BP2 lacks the usual degron pattern that cereblon recognizes. Follow-up cell experiments, including sensitive light-based readouts of protein proximity, confirmed that MRT-5702 pulls cereblon and G3BP2 together into a three-part complex that leads to G3BP2 degradation, while sparing its closely related sibling protein G3BP1 unless its key domain is swapped.
Imitation as a Binding Strategy
Digging deeper, the researchers asked how G3BP2 could bind cereblon without the standard recognition motif. Instead of looking for similarities between G3BP2 and known cereblon targets, they flipped the question: could cereblon resemble one of G3BP2’s usual binding partners? G3BP proteins commonly interact with other molecules through a hotspot on a region called the NTF2-like domain, which normally recognizes short sequence motifs. Using structural models of G3BP2’s natural partner USP10, the team computationally scanned cereblon’s surface and found a patch on a lesser-understood region, the LON domain, that closely mimics the shape and chemistry of USP10’s binding motif. Mutating just a few key amino acids on this patch, or in the G3BP2 hotspot, weakened the three-part complex, suggesting that cereblon is effectively impersonating USP10 to dock onto G3BP2.
Capturing the New Interface in Atomic Detail
To visualize this unusual interaction, the team used high-resolution cryo–electron microscopy to solve the structure of a complex containing cereblon, another core ligase component, MRT-5702, and the NTF2-like domain of G3BP2. The images revealed that a flexible loop in cereblon’s LON domain bends and reshapes to form nearly half of the contact surface with G3BP2. The active mirror-image form of MRT-5702 sits in cereblon’s usual drug-binding pocket but, together with the loop, helps create a broad landing pad that cradles one half of the G3BP2 dimer. Strikingly, this arrangement barely uses the canonical cereblon site that handles previous glue targets, showing that cereblon can engage very different regions of its surface depending on the glue and the protein involved. At the same time, the untouched half of the G3BP2 dimer remains available to bind other partners, offering a plausible route for “collateral” degradation of proteins that happen to be attached.
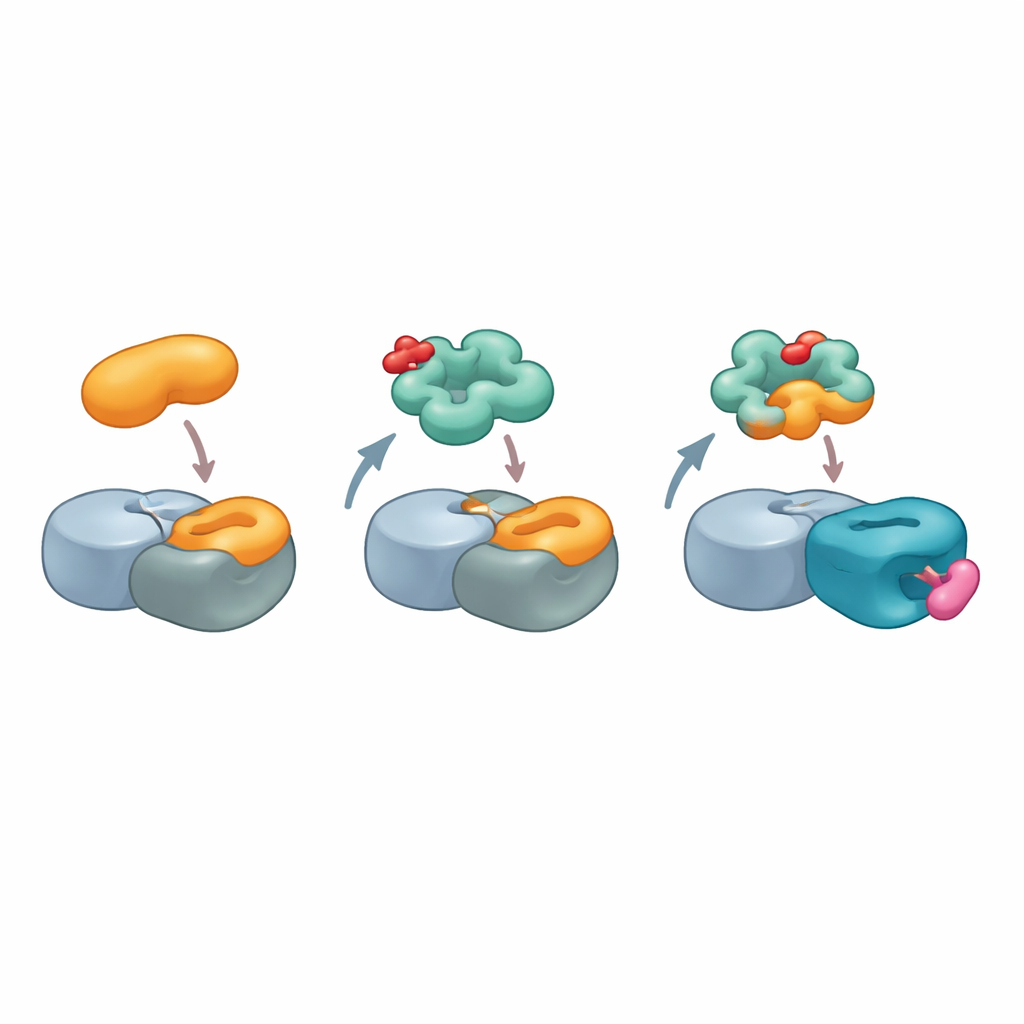
Glueprints for Future Drug Design
Taken together, the findings paint cereblon as a far more versatile matchmaker than previously appreciated. Rather than insisting that targets carry a single kind of degron, cereblon can, when armed with the right molecular glue, mold a composite surface that imitates natural protein–protein contacts elsewhere in the cell. The authors propose that such combined protein-and-glue surfaces, which they call "glueprints," could be mapped and designed to mimic existing interaction hot spots on many disease-linked proteins. For a lay reader, the key message is that drug developers may not need to discover brand-new binding pockets on troublesome proteins; instead, they can teach the cell’s own recycling machinery to recognize those proteins by cleverly copying the shapes of their usual partners, greatly widening the menu of targets that can be safely and selectively removed.
Citation: Annunziato, S., Quan, C., Donckele, E.J. et al. Cereblon induces G3BP2 neosubstrate degradation using molecular surface mimicry. Nat Struct Mol Biol 33, 479–487 (2026). https://doi.org/10.1038/s41594-025-01738-8
Keywords: molecular glue degraders, cereblon, targeted protein degradation, G3BP2, protein–protein interactions