Clear Sky Science · en
Scaffolds with optimized quaternary symmetry for de novo cryoEM structure determination of small RNAs
Seeing the Smallest Shapes of RNA
Inside every cell, short strands of RNA fold into tiny three-dimensional shapes that switch genes on and off, sense cellular damage, or light up under a microscope. Many of these RNAs are so small that current imaging methods struggle to reveal their precise architecture. This article introduces a clever way to make these elusive molecules visible: by fastening them onto a larger, self-assembling RNA "frame" that can be seen clearly by cryo–electron microscopy, a powerful technique that images frozen biomolecules.
Building a Helpful RNA Frame
The authors began with an RNA segment from a virus that naturally tends to pair up into a two-part structure. They redesigned this segment so that, instead of forming pairs only a small fraction of the time, it now almost always assembles into highly regular two-part or four-part shapes in solution. These repeating arrangements create what is essentially an RNA frame, or scaffold, with built-in symmetry. Symmetry is valuable for cryo–electron microscopy because identical repeated units can be averaged together, sharpening the final picture.
Attaching Known RNAs as Test Guests
To test whether their scaffold could carry other RNAs into view, the researchers grafted well-studied molecules onto one region of the frame. One guest was a transfer RNA from bacteria, a classic L-shaped molecule that delivers amino acids during protein synthesis. Another was Mango-III, a tiny engineered RNA that binds a dye and glows, widely used as a fluorescent tag. In both cases, the combined molecules folded and paired as designed, and cryo–electron microscopy produced detailed maps of the overall shapes. For the transfer RNA, the images were sharp enough to spot subtle differences between the unmodified form used here and previously studied, chemically modified versions. For Mango-III, the maps showed that the aptamer becomes much more rigid when its dye is bound, explaining how binding switches fluorescence on. 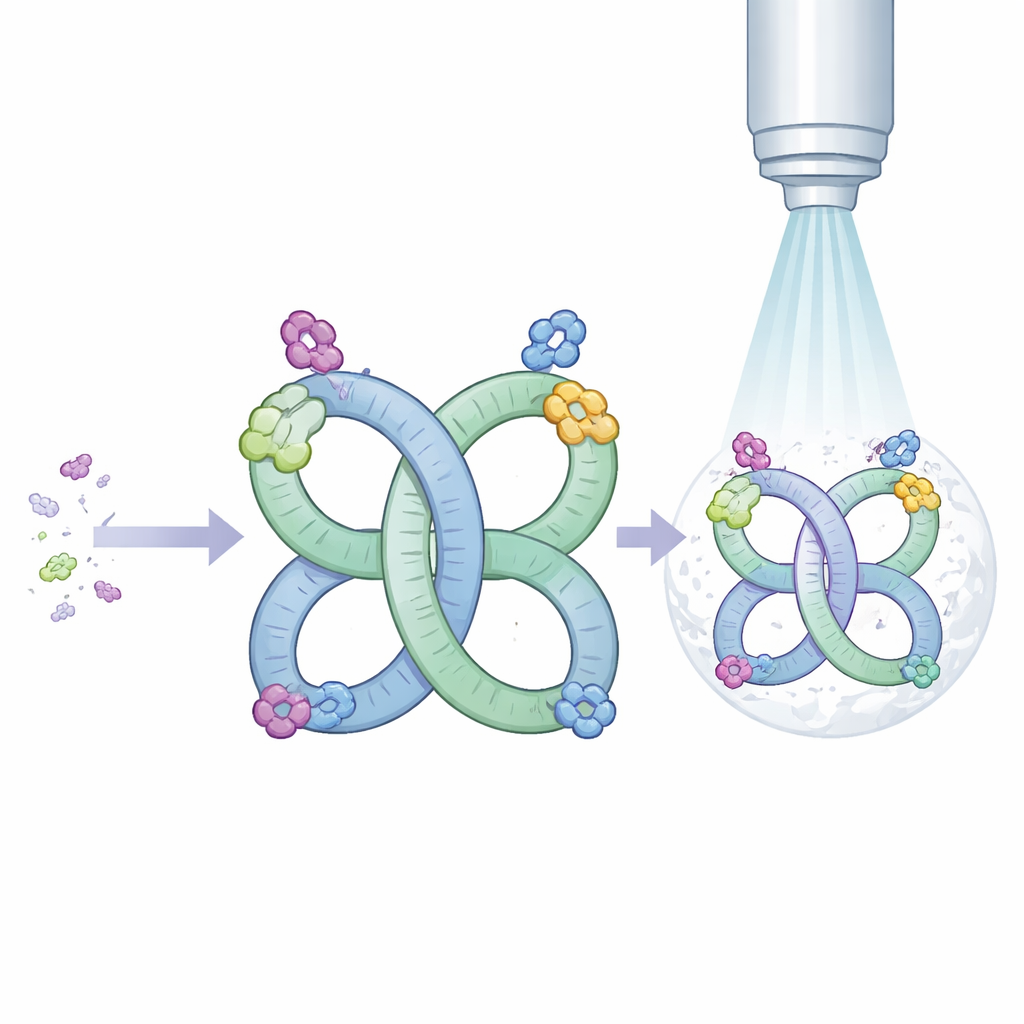
Revealing How Designer RNAs Grasp Small Molecules
The team then moved beyond test cases to RNAs whose full structures had not yet been seen. They attached two small aptamers—short RNAs selected in the lab to bind specific small molecules—to the scaffold. One aptamer recognizes the drug quinine; the other senses 8-oxoguanine, a damaged form of a genetic letter that signals oxidative stress in bacteria. Thanks to the scaffold, cryo–electron microscopy delivered exceptionally high-quality maps, fine enough to trace each RNA chain from end to end and to see where metal ions and water molecules sit. In the quinine aptamer, the binding pocket cradles the drug mainly through snug stacking and shape complementarity, with surprisingly few direct hydrogen bonds. In contrast, the 8-oxoguanine aptamer wraps its ligand in an intricate web of hydrogen bonds that contact almost every chemically distinct site on the damaged base, explaining its sharp discrimination between 8-oxoguanine and normal guanine.
Flexible Symmetry for Clearer Pictures
Interestingly, the same RNA scaffold can assemble into either pairs or four-part structures depending on conditions and the attached guest. When a four-part arrangement forms, the repeated geometry further improves image quality. In one case, the scaffold adopted a four-part shape even though its sequence was identical to the two-part version, highlighting how small shifts in base pairing can reorganize the entire assembly. The authors also explored practical aspects of cryo–electron microscopy data collection, such as how stage tilting can overcome preferred orientations of particles on the grid, and how imposed symmetry during image processing modestly but consistently sharpens the resulting structures.
A New Window into Tiny RNA Machines
Overall, this work shows that a compact, symmetric RNA frame can turn otherwise invisible small RNAs into excellent cryo–electron microscopy targets, enabling structures beyond atomic-level detail in favorable cases. By attaching an unknown RNA to the scaffold through a simple helical connector, researchers can now determine its three-dimensional fold, see exactly how it grips a small-molecule partner, and spot ordered metal ions and water molecules that tune its behavior. For a general audience, the key message is that we now have a practical tool to look closely at some of the smallest and most versatile RNA machines in nature and biotechnology, paving the way for rational design of new RNA-based sensors, drugs, and molecular devices. 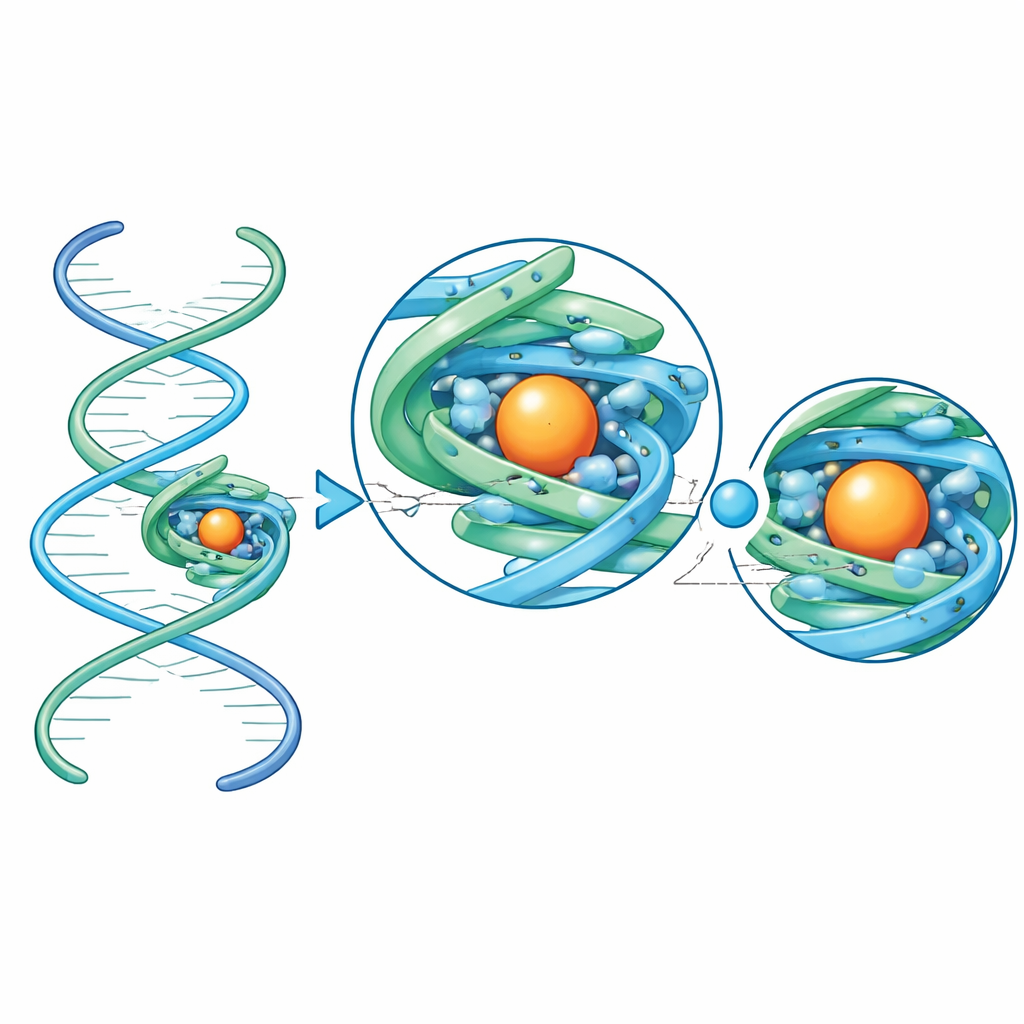
Citation: Jones, C.P., Ferré-D’Amaré, A.R. Scaffolds with optimized quaternary symmetry for de novo cryoEM structure determination of small RNAs. Nat Methods 23, 609–616 (2026). https://doi.org/10.1038/s41592-026-03016-x
Keywords: RNA structure, cryo electron microscopy, aptamer, riboswitch, molecular scaffolds