Clear Sky Science · en
Multimodal antigenic escape to GPRC5D-targeted T cell engagers in multiple myeloma
When Cancer Outsmarts Precision Immunotherapy
Powerful new immune-based drugs are transforming treatment for multiple myeloma, a blood cancer that remains incurable for most patients. These therapies work by redirecting the body’s own T cells to seek and destroy tumor cells marked by specific “flags” on their surface. One promising flag is a molecule called GPRC5D. Yet, as with many targeted therapies, tumors can evolve and slip past treatment. This study unravels, in unusual detail, how myeloma cells change or hide the GPRC5D flag in order to escape highly effective T cell–engaging drugs.
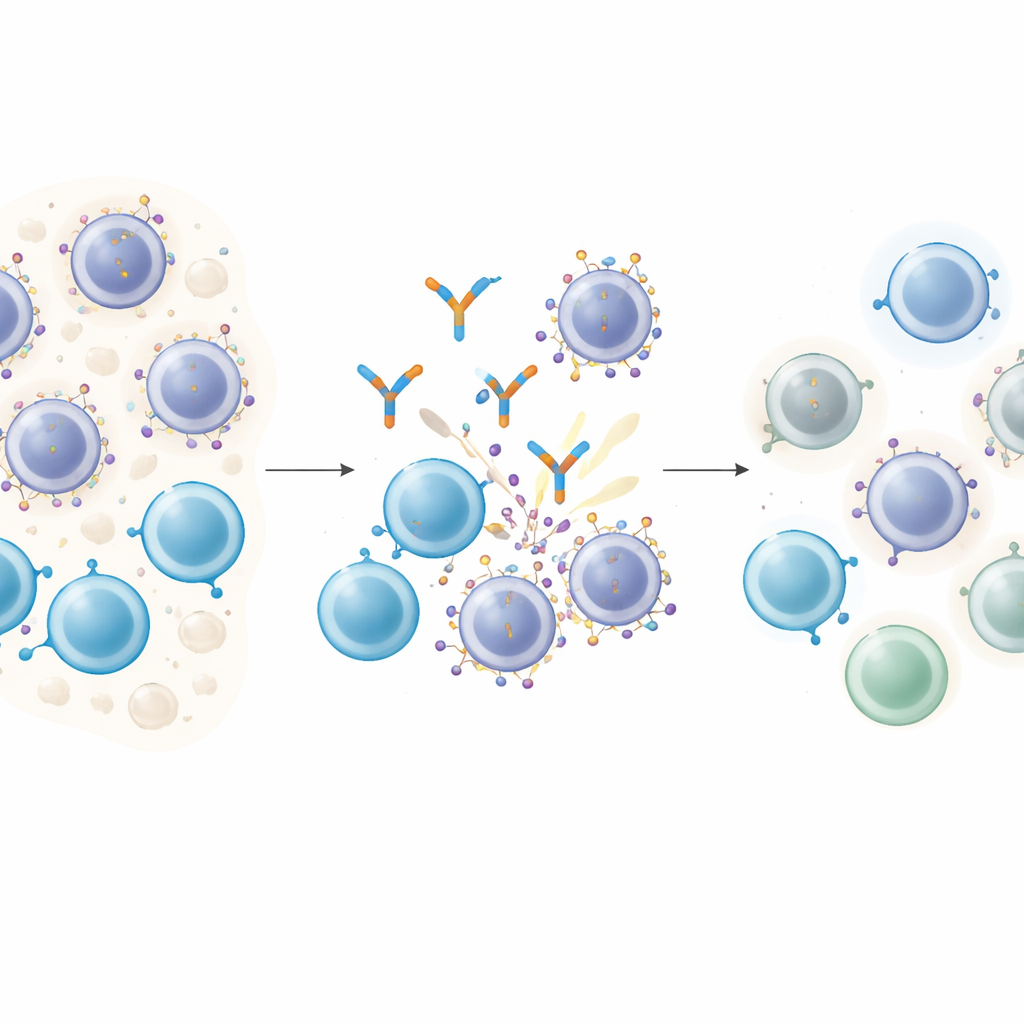
How Smart Drugs Lock Onto Cancer Cells
T cell–engaging antibodies act like molecular matchmakers: one end grabs a T cell, the other latches onto a target molecule on the cancer cell, bringing the two into direct contact so the T cell can kill its prey. In multiple myeloma, GPRC5D is an attractive target because it is abundant on malignant plasma cells but scarce on most normal tissues. Patients treated with a GPRC5D‑directed drug called talquetamab often experience deep remissions. However, nearly all eventually relapse, raising a critical question: are the cancer cells changing the very target that these drugs depend on?
Tracking Tumor Evolution in Real Time
The researchers followed 21 people with hard‑to‑treat myeloma who received talquetamab. They collected bone marrow samples before treatment in some patients and again when the cancer came back, then used multiple layers of DNA, RNA and chromatin analysis to examine the tumor cells. This ultra‑detailed view revealed that, in about two‑thirds of relapsed patients, the cancer had altered GPRC5D in ways that would blunt or block the drug’s effect. Strikingly, several different resistant subclones—genetically distinct branches of the tumor—often coexisted in the same person, showing that evolution toward escape can follow multiple paths at once.
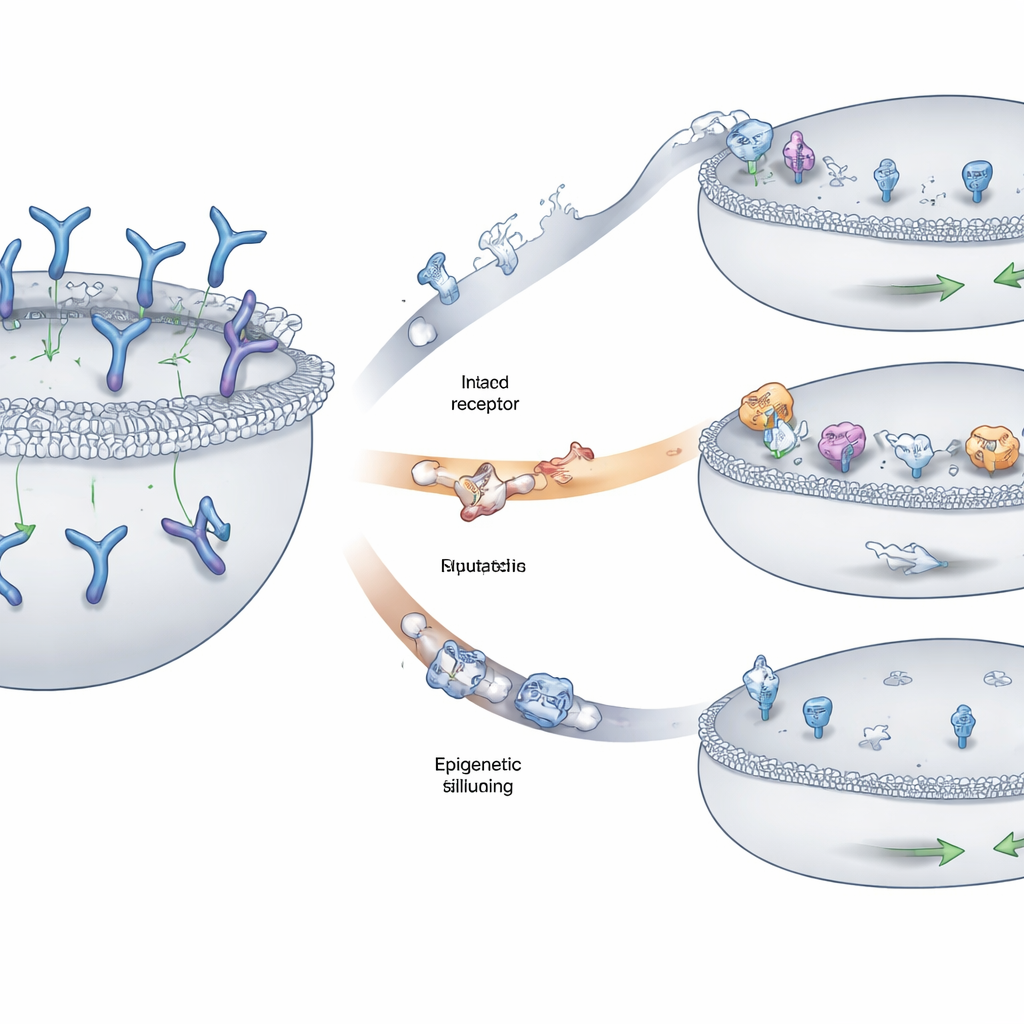
Three Main Ways Tumors Hide the Target
The team found three recurring patterns by which myeloma cells evaded recognition. In some patients, pieces of the chromosome carrying the GPRC5D gene were deleted on both copies, erasing the flag entirely from the cell surface. In others, smaller changes—single‑letter DNA swaps or tiny insertions and deletions—distorted key regions of the GPRC5D protein. Many of these changes disrupted parts of the protein needed to travel properly from inside the cell to the outer membrane, causing GPRC5D to get stuck in an internal compartment instead of being displayed where the drug could see it. A third route involved epigenetic silencing: the DNA surrounding GPRC5D became tightly packaged and chemically marked so that the gene was barely read, even though the underlying sequence was largely intact. Tumor cells using this strategy made little or no GPRC5D at all.
When the Flag Is Present but the Lock No Longer Fits
Not all escape routes removed GPRC5D from the cell surface. In some cases the protein looked normal to standard lab tests but no longer fit the drug like a key in a lock. The team recreated specific patient‑derived mutations in laboratory cell lines and tested how well different GPRC5D‑targeted antibodies could bind and kill these cells. One mutation subtly altered the exact spot where talquetamab grips the receptor, sharply reducing the drug’s ability to bind and to trigger cell death, even though the receptor was still visible with routine staining. Intriguingly, a different GPRC5D‑directed drug that holds onto the receptor at two sites remained effective against several of these mutants, highlighting how drug design—such as using multivalent or multi‑epitope binding—can influence whether resistant clones survive.
Beyond the Target: Other Limits to Immune Attack
While most relapses showed some form of GPRC5D loss or alteration, a minority of patients relapsed without obvious changes in the target itself. In these cases, the problem appeared to lie in the T cells: they were less fit and less capable of killing cancer cells in lab tests, suggesting that long‑term continuous exposure to engager therapy may exhaust the immune system. The study also uncovered that a particular genetic subtype of myeloma (involving a chromosomal swap called t(11;14)) naturally keeps the GPRC5D gene in a more “closed” chromatin state, with lower baseline expression. This raises the possibility that some patients may be predisposed to poor responses because their tumors start out with muted levels of the target flag.
What This Means for Patients and Future Treatments
For patients and clinicians, this work explains why even highly potent GPRC5D‑targeted therapies are not yet cures: myeloma cells can delete, reshuffle, misroute or switch off the very flag these drugs rely on, and they often do so in several ways at once. The findings suggest that monitoring tumor DNA, rather than just measuring protein levels, will be important for detecting emerging escape mutations that standard lab tests can miss. They also point toward next‑generation strategies—such as drugs that bind GPRC5D more avidly at multiple sites, combinations that hit more than one target at once, or agents that reopen silenced genes—to stay ahead of tumor evolution. In essence, this study maps the escape routes myeloma uses so that future therapies can be designed to block them, keeping T cell–based treatments effective for longer.
Citation: Lee, H., Ahn, S., Gonzales, G.A. et al. Multimodal antigenic escape to GPRC5D-targeted T cell engagers in multiple myeloma. Nat Med 32, 964–977 (2026). https://doi.org/10.1038/s41591-025-04175-8
Keywords: multiple myeloma, T cell engagers, GPRC5D, antigen escape, cancer immunotherapy