Clear Sky Science · en
Xylosyltransferase engineering to manipulate proteoglycans in mammalian cells
Why Cell Coats Matter
Every cell in our body wears a molecular “coat” that helps it talk to its neighbors and sense its surroundings. A major part of this coat is made of proteoglycans—proteins decorated with long sugar chains that act like tiny antennae for growth factors, immune signals and structural cues. When this sugar decoration goes wrong, development can falter and tissues such as bone and muscle can be severely affected. Yet scientists have struggled to map exactly which proteins carry these chains and how individual components of the coat influence cell behavior. This study introduces a precise chemical strategy to tag and re‑build these sugar chains on living mammalian cells, opening a way to read and rewrite the information encoded on their surfaces.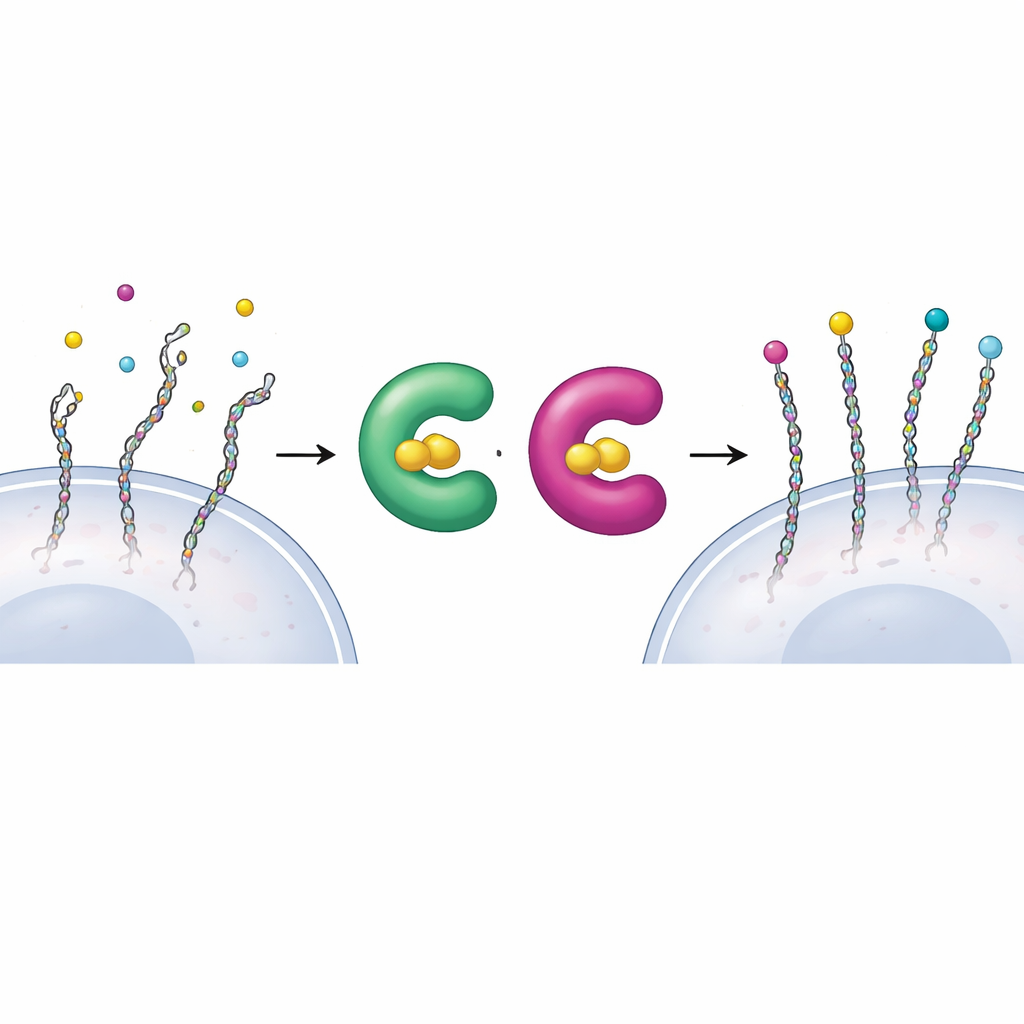 
Rewiring the First Step of Sugar Attachment
The construction of proteoglycans begins when a special enzyme family, xylosyltransferases XT1 and XT2, installs the first sugar onto a protein, marking the spot where a long chain will grow. Because both enzymes can perform this same first step, it has been difficult to tell which proteins each one works on inside cells. The authors used a “bump‑and‑hole” design—common in modern chemical biology—to solve this. They slightly reshaped the active site of XT1, creating extra space (the “hole”), and introduced a matching, slightly bulkier sugar building block, 6AzGlc (the “bump”). The engineered XT1 can now use this modified sugar, whereas the natural enzyme cannot, allowing researchers to tag only those proteins touched by the redesigned enzyme.
Smuggling a Designer Sugar into Cells
To make this approach work in living cells, the team had to deliver the bumped sugar in a form that cellular metabolism could recognize. Instead of relying on normal sugar uptake pathways—which do not handle xylose analogs well—they built a caged version of 6AzGlc carrying a hidden phosphate group. Once inside the cytosol, cellular enzymes unmask this group and convert the molecule into UDP‑6AzGlc, the activated form required by the engineered xylosyltransferases. Careful chromatography confirmed that cells treated with the correctly configured caged compound produced substantial amounts of the activated designer sugar, whereas a mirror‑image control produced almost none. 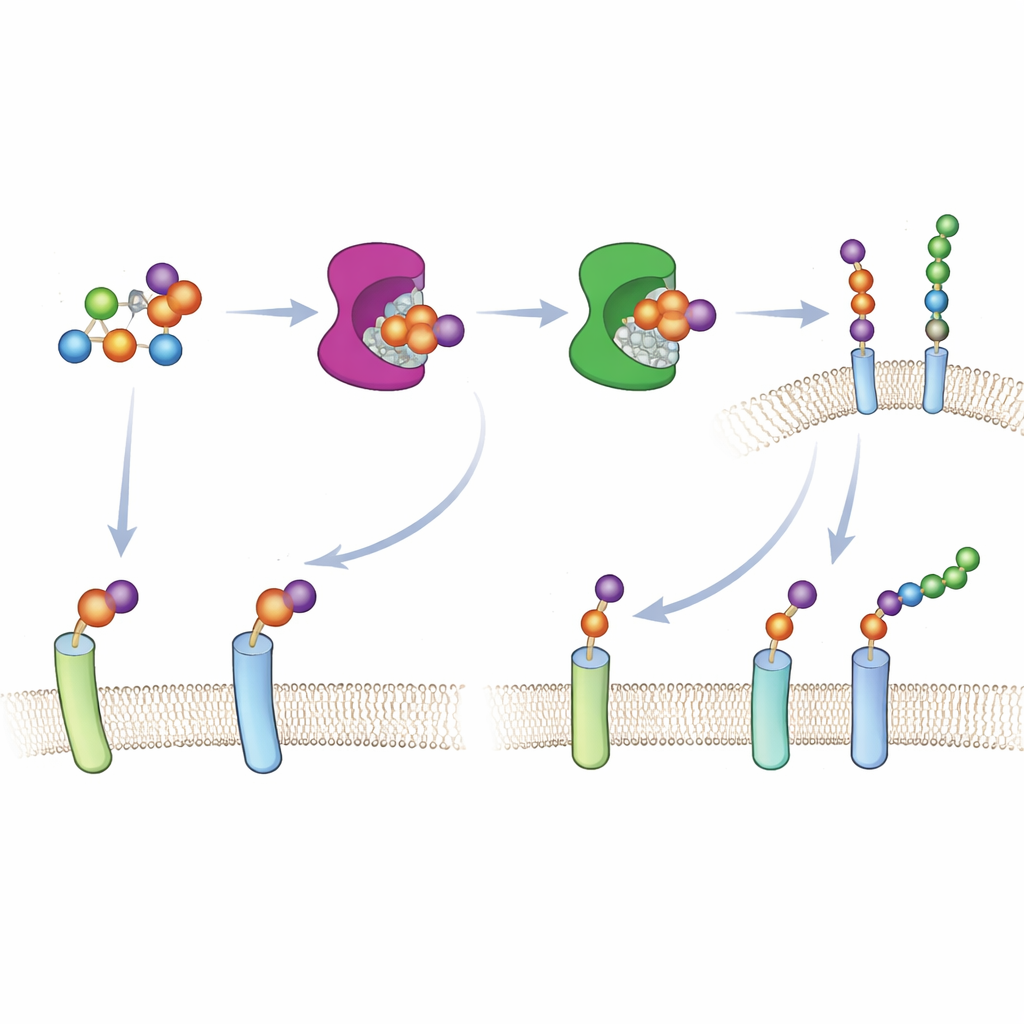
Tagging and Identifying the Hidden Players
With both the modified enzyme and the activated sugar in place, the researchers showed that only cells expressing the engineered XT1 or XT2 incorporated the 6AzGlc tag into their surface proteoglycans. The azide group on 6AzGlc serves as a tiny chemical handle that can be “clicked” to fluorescent dyes or biotin, enabling visualization and enrichment of tagged proteins. Mass spectrometry of enriched samples revealed a rich collection of known proteoglycans—including decorin, multiple glypicans, syndecan‑4, CD44 and versican—confirming that the system marks genuine sugar‑bearing proteins. Importantly, the engineered enzymes preserved the same sequence preferences as their natural counterparts, meaning that the chemical tag is installed at authentic attachment sites rather than at new, artificial positions.
Building Designer Proteoglycans
An unexpected bonus of the 6AzGlc tag is that, unlike the natural starting sugar, it cannot be extended into a full‑length chain by downstream enzymes. This effectively caps the chain and simplifies the molecular structure that must be analyzed by mass spectrometry. The authors turned this feature into a tool: after XT1 installs 6AzGlc at a natural attachment site, they use click chemistry to bolt on synthetic heparin fragments, creating “designer proteoglycans” with precisely defined sugar chains. In breast cancer cells lacking their own syndecan‑1, adding back such designer versions restored normal spreading on a protein‑coated surface, proving that the chemically rebuilt proteoglycan can substitute functionally for the natural one.
What This Means for Biology and Medicine
This work delivers a powerful set of tools to selectively label and manipulate the sugar chains that control how cells communicate. By splitting the roles of XT1 and XT2 and tagging only their direct targets, researchers can now map which proteoglycans operate in specific tissues and disease states. The ability to stop natural chain growth and replace it with custom‑built sugars also allows scientists to dissect how much of a proteoglycan’s function is carried by its protein backbone versus its sugar coat. In the long term, such precision engineering may help decode complex signaling at the cell surface and inspire therapies that repair or reprogram faulty cellular coats in developmental disorders, cancer and other diseases.
Citation: Li, Z., Chawla, H., Di Vagno, L. et al. Xylosyltransferase engineering to manipulate proteoglycans in mammalian cells. Nat Chem Biol 22, 612–621 (2026). https://doi.org/10.1038/s41589-025-02113-w
Keywords: proteoglycans, glycosaminoglycans, xylosyltransferase, chemical biology, cell surface signaling