Clear Sky Science · en
Genome-wide association analyses highlight the role of the intestinal molecular environment in human gut microbiota variation
Why Your DNA and Gut Bacteria Belong in the Same Story
Trillions of microbes live in our intestines and influence everything from digestion to metabolism and even the immune system. But why do some people naturally harbor different mixes of gut bacteria than others, even when they live in the same place and eat similar foods? This study, based on detailed genetic and gut microbe data from nearly 30,000 adults in Sweden and Norway, shows that our own DNA quietly helps script the community of microbes living inside us.
A Massive Look Inside Nordic Guts
To uncover how human genes shape the microbiome, researchers combined data from four large Swedish population studies, covering 16,017 adults, and checked their findings in 12,652 Norwegians. All participants provided blood for human DNA analysis and stool samples for deep sequencing of microbial DNA. Instead of focusing only on broad groups of bacteria, the team used high-resolution methods that can distinguish hundreds of individual species. They then scanned the human genome, variant by variant, to see which stretches of DNA tracked with overall microbial richness (how many different species are present) and with the presence or abundance of specific bacterial species.
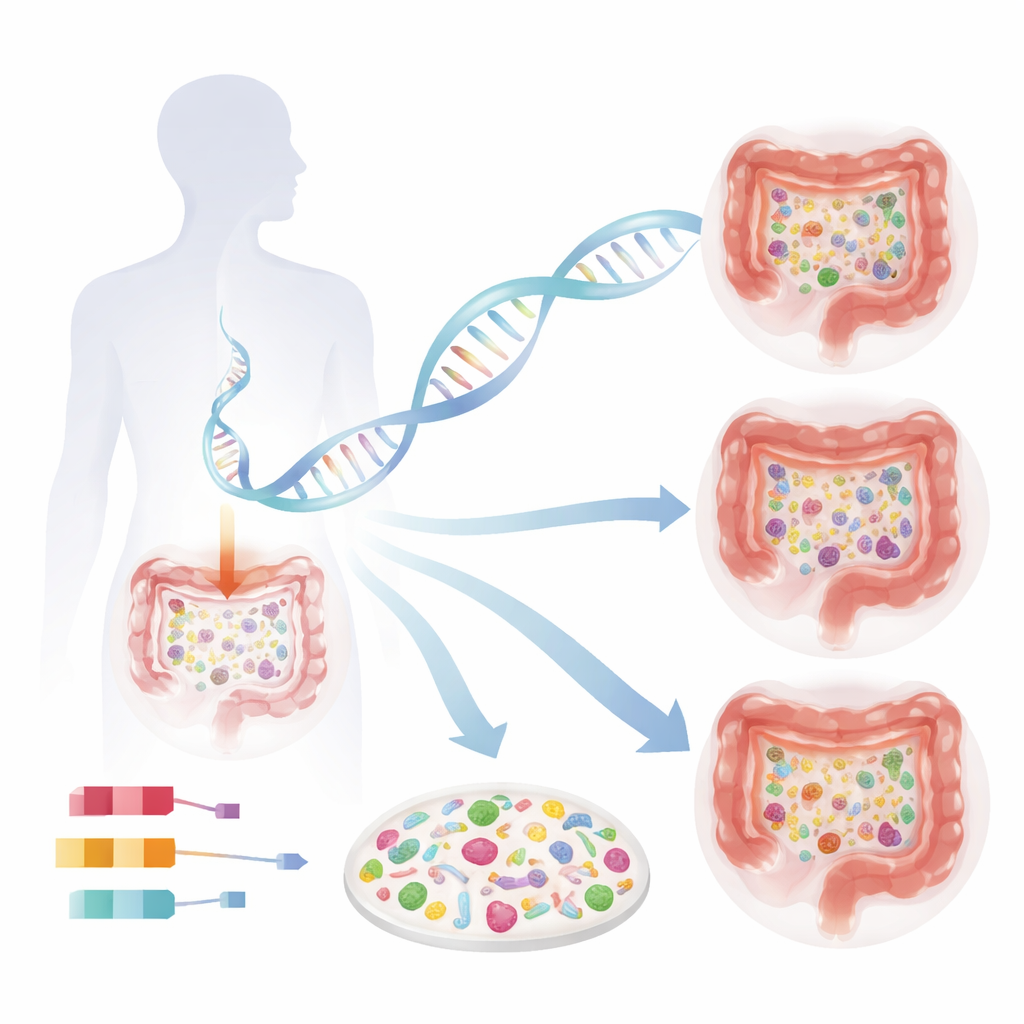
Genetic Switches That Tune Microbial Richness
One of the most striking discoveries was a region of the human genome containing two genes, OR51E1 and OR51E2, previously known as odor receptors. These receptors also sit on special hormone-producing cells in the gut lining and sense fatty acids made by microbes. People carrying a particular version of this DNA region tended to have fewer bacterial species in their intestines, and this pattern was confirmed independently in the Norwegian group. The finding suggests that how our gut cells sense microbe-derived fatty acids feeds back on the diversity of the microbiome itself, possibly by altering gut hormones that control motility, appetite or local immune responses.
Surface Sugars, Mucus, and the Microbial Neighborhood
The study also pinpointed several genetic regions that govern the sugary and slimy environment at the gut surface—prime real estate for bacteria. Variants in the well-known lactase (LCT) gene, which determines whether adults can digest milk sugar, were tied to shifts in multiple species, including Bifidobacterium that thrive on lactose. Genes that define blood groups and related “secretor” status—ABO, FUT2 and FUT3–FUT6—modify fucose-containing sugars displayed on intestinal mucus and in secretions. Different genetic combinations here were linked to distinct sets of bacteria that can latch onto or feed on these sugars. Another key region lay within a mucin gene, MUC12, part of the scaffold for the mucus layer itself. Changes in this region tracked with the abundance of a species called Coprobacillus cateniformis and even shared a genetic signal with how often people move their bowels, hinting at intertwined effects on gut function and microbial makeup.
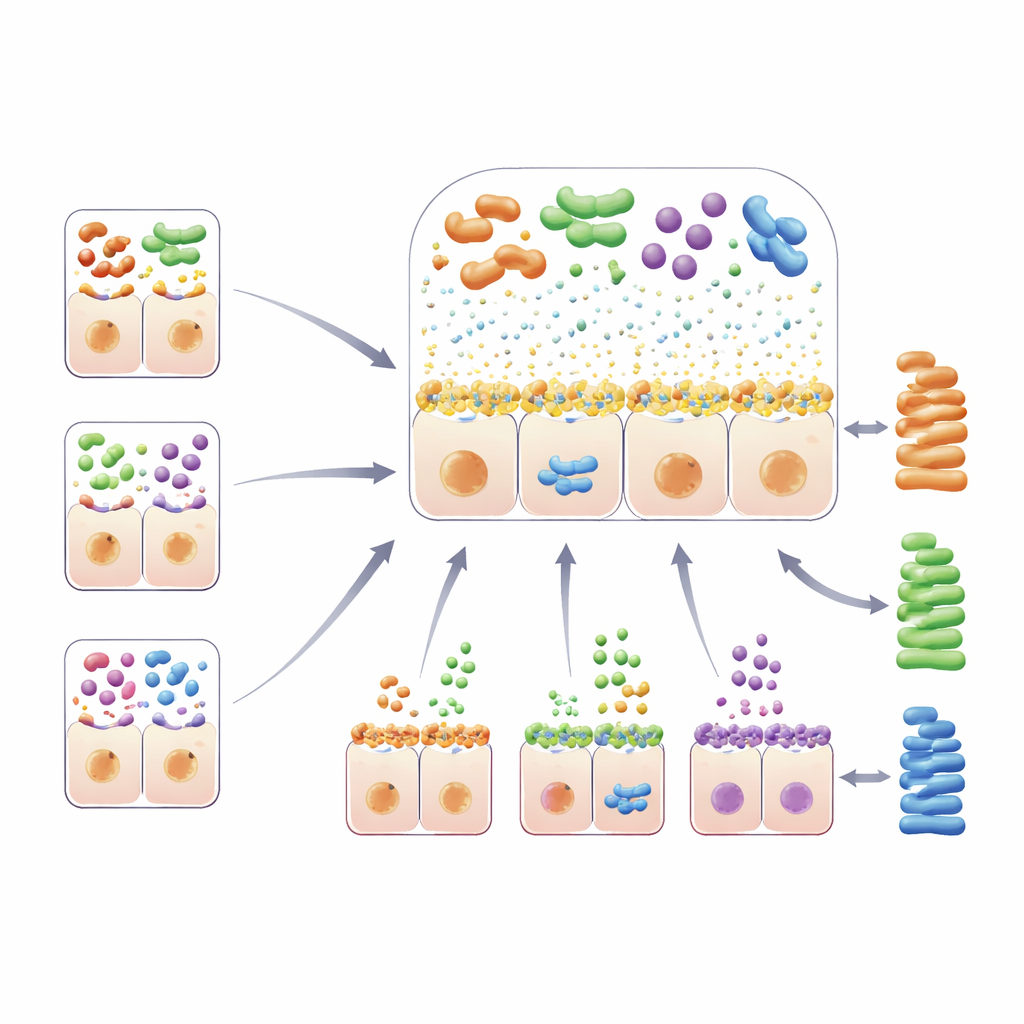
From Microbes to Metabolism and Body Shape
Beyond mapping “who lives there,” the team asked whether DNA regions tied to certain bacteria also overlapped with human traits such as blood cholesterol, bile acids and body fat distribution. In several cases the same pieces of genome were involved. Variants near the CORO7–HMOX2 and FOXP1 genes affected a cluster of bacteria including Turicibacter and Clostridium saudiense, and also lined up with differences in waist–hip ratio, bile acids and low-density lipoprotein (LDL) cholesterol. Using genetic tools designed to suggest cause-and-effect, the authors found hints that one microbe, an Intestinibacter species, may push LDL cholesterol higher, and that Turicibacter might influence where body fat is stored. Another region, SLC5A11, was linked to a butyrate-producing bacterium, Agathobaculum butyriciproducens, which has shown protective effects in animal models of brain disease. Here, the human DNA variant appeared to lower blood levels of a small molecule called myo-inositol while favoring the growth of this potentially helpful microbe.
What This Means for Health and Future Treatments
Taken together, these results show that human genes involved in gut sensing, mucus composition, and surface sugars help determine which microbial species can successfully settle in our intestines. The effects are modest for any single gene, and the picture so far is clearest for relatively common bacteria in people of European ancestry. Still, the work expands the list of human DNA regions reliably tied to specific gut microbes from just a couple to at least eight, and links several of them to metabolic traits such as cholesterol and body fat patterning. For a layperson, the key message is that the gut microbiome is not shaped by diet and environment alone: our own genetic blueprint builds the habitat that microbes encounter, nudging the community toward some residents and away from others. As larger and more diverse studies arrive, understanding this two-way relationship between genes and microbes could help tailor dietary advice, predict disease risks and perhaps guide therapies that combine drugs, diet and targeted manipulation of the microbiome.
Citation: Dekkers, K.F., Pertiwi, K., Baldanzi, G. et al. Genome-wide association analyses highlight the role of the intestinal molecular environment in human gut microbiota variation. Nat Genet 58, 540–549 (2026). https://doi.org/10.1038/s41588-026-02512-2
Keywords: gut microbiome, human genetics, intestinal mucus, bile acids, metabolism