Clear Sky Science · en
The HUNT study identifies host genetic factors reproducibly associated with human gut microbiota composition
Why Your Genes and Gut Bacteria Matter
Each of us carries trillions of microbes in our intestines that help digest food, train our immune system and may even influence our risk of disease. This study asks a deceptively simple question: how much of that inner ecosystem is written in our DNA? By scanning the genomes of tens of thousands of people and comparing them with the detailed makeup of their gut microbes, the researchers show that specific human genes consistently nudge the microbiome in particular directions—and that these shifts connect to conditions such as celiac disease, hemorrhoids and heart-related problems, as well as to body weight.
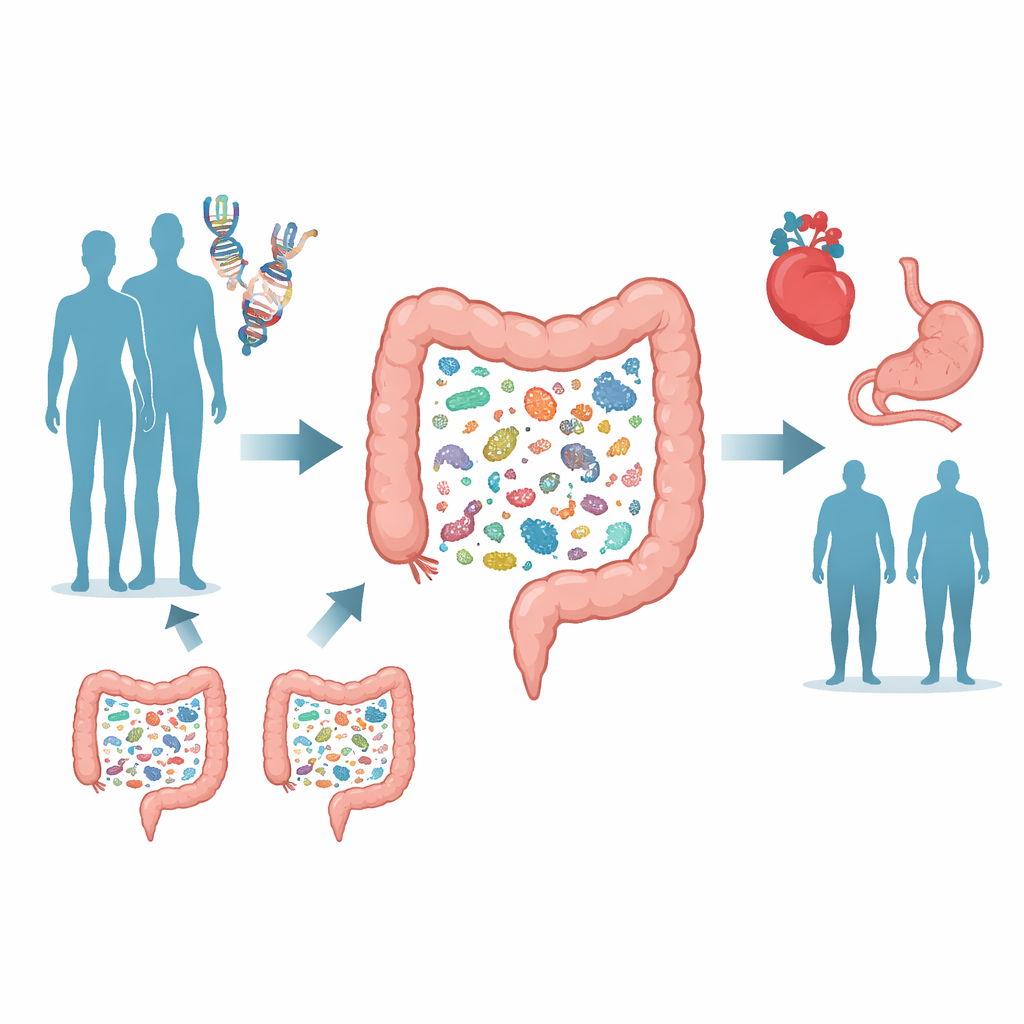
Looking for Patterns in a Crowd
To track down gene–microbe links, the team drew on the Trøndelag Health Study in Norway, where over 12,000 adults provided both blood for DNA analysis and stool samples for microbial profiling. Unlike many earlier projects that used relatively coarse bacterial fingerprints, this work relied on deep metagenomic sequencing, which reads much of the DNA from each sample and can distinguish hundreds of distinct bacterial species and their metabolic capabilities. The scientists then performed a genome-wide association study, scanning nearly eight million human genetic variants to see which ones lined up with differences in the relative abundance of 546 common gut species and with measures of overall microbial diversity.
Six Genetic Hotspots That Shape the Gut
The analysis revealed a surprisingly strong genetic imprint. Twelve robust associations emerged between human DNA variants and specific bacterial species, clustered in six regions of the genome. Two of these regions, near the LCT gene involved in lactose digestion and the ABO blood group gene, had been implicated before, but four—near HLA-DQB1, MUC12, SLC37A2 and FUT2—were new or newly confirmed. For example, people with the lactase-persistent version of LCT tended to carry fewer Bifidobacterium adolescentis, a species that thrives on milk sugar left over in the gut when lactose is not fully broken down. The FUT2 region, which affects whether blood group sugars are displayed on the gut lining, was linked to several bacterial species that appear to feed on those sugar-coated surfaces.
From Microbes to Disease Risk
The story became more intriguing when the researchers overlaid these genetic results with large databases of human diseases. Variants in the HLA-DQB1 region that were tied to higher levels of an Agathobacter species were also associated with a lower risk of autoimmune conditions, including celiac disease. People with celiac disease in the Norwegian cohort tended to have especially low levels of this microbe, suggesting that the illness may, in part, reshape the gut community. Another region, near the MUC12 gene, was connected both to the abundance of a bacterium called Coprobacillus cateniformis and to a reduced risk of hemorrhoidal disease. Laboratory work showed that MUC12 is heavily produced in the cells lining the colon, hinting that subtle changes in this mucus barrier could influence which bacteria flourish and how they, in turn, affect delicate blood vessels and tissue in the rectum.
Microbial Functions, Heart Health and Body Weight
Beyond individual species, the team examined what the microbes were capable of doing by grouping their genes into functional modules, such as transport systems and regulatory circuits. The same human genetic regions—LCT, ABO and FUT2—also influenced these microbial functions, suggesting that our DNA shapes not just who is there in the gut, but what they are doing. At the FUT2 site, for instance, variants linked to "non-secretor" status went hand-in-hand with bacteria associated with potentially harmful metabolites and with an elevated risk of high cholesterol and high blood pressure. Finally, using a technique called Mendelian randomization, which leverages genetic variants as natural experiments, the researchers found evidence that a higher body mass index causally shifts the microbiome: people genetically predisposed to higher weight tended to have lower overall microbial diversity and consistent changes across many species.
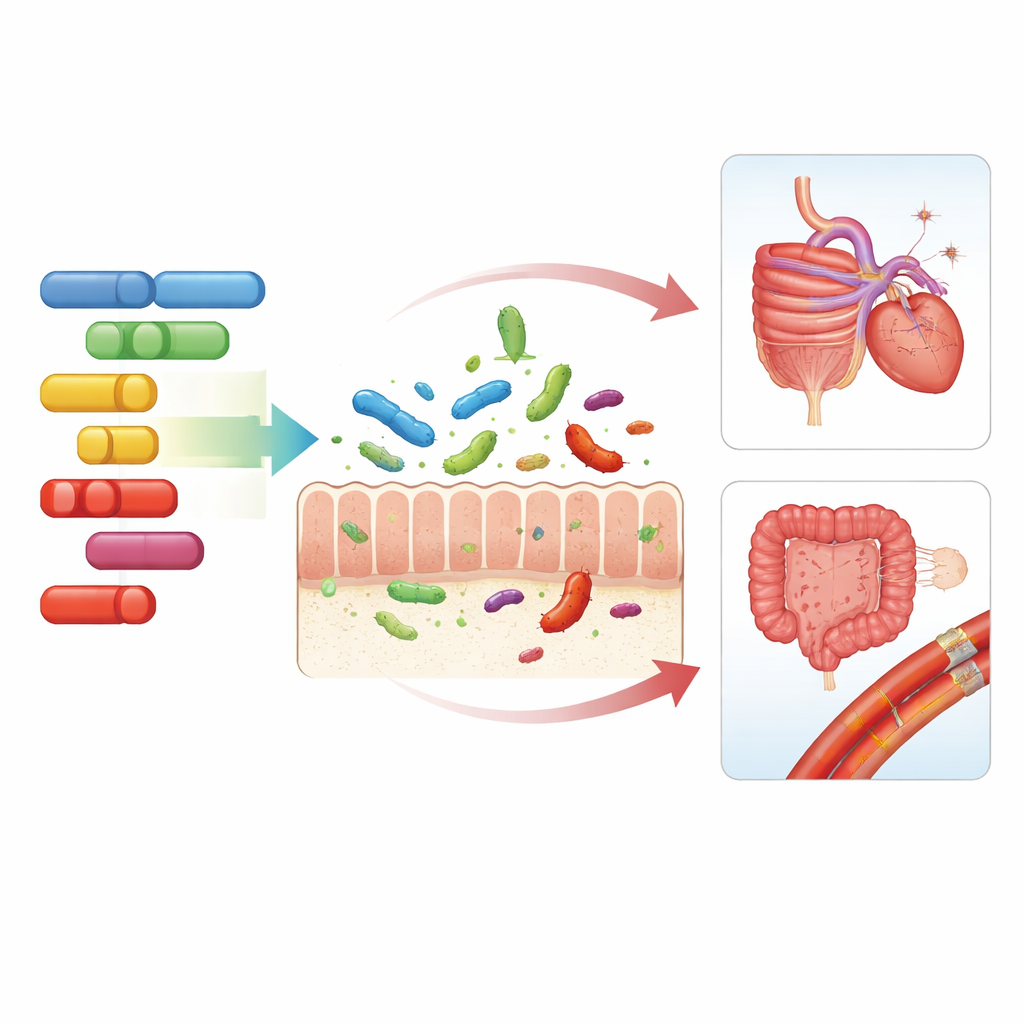
What It Means for Everyday Health
Taken together, these findings paint a picture of a three-way conversation between our genes, our gut microbes and our health. Certain stretches of human DNA subtly favor or discourage specific bacterial residents and microbial activities, which then intersect with risks for digestive illnesses, heart and vessel problems and the effects of excess body weight. While these genetic influences explain only a slice of the enormous variation in gut communities—and do not yet translate into clinical tests—they help clarify why people respond differently to the same diet or environment and point toward more personalized approaches to nutrition and disease prevention that consider both genome and microbiome.
Citation: Moksnes, M.R., Coward, E., Nethander, M. et al. The HUNT study identifies host genetic factors reproducibly associated with human gut microbiota composition. Nat Genet 58, 530–539 (2026). https://doi.org/10.1038/s41588-026-02502-4
Keywords: gut microbiome, human genetics, microbiota and disease, body weight and microbes, genome-wide association