Clear Sky Science · en
Reduced cyclin D3 expression in erythroid cells protects against malaria
How a Subtle Blood Difference Can Fight a Deadly Parasite
Malaria has shaped human evolution for millennia, favoring genetic tweaks that help people survive infection. This study uncovers one such tweak in a Sardinian population: a small DNA change that slightly alters how red blood cells are made. That change makes red blood cells fewer and larger, raises their internal stress chemistry, and in doing so quietly sabotages the malaria parasite that depends on them to grow.
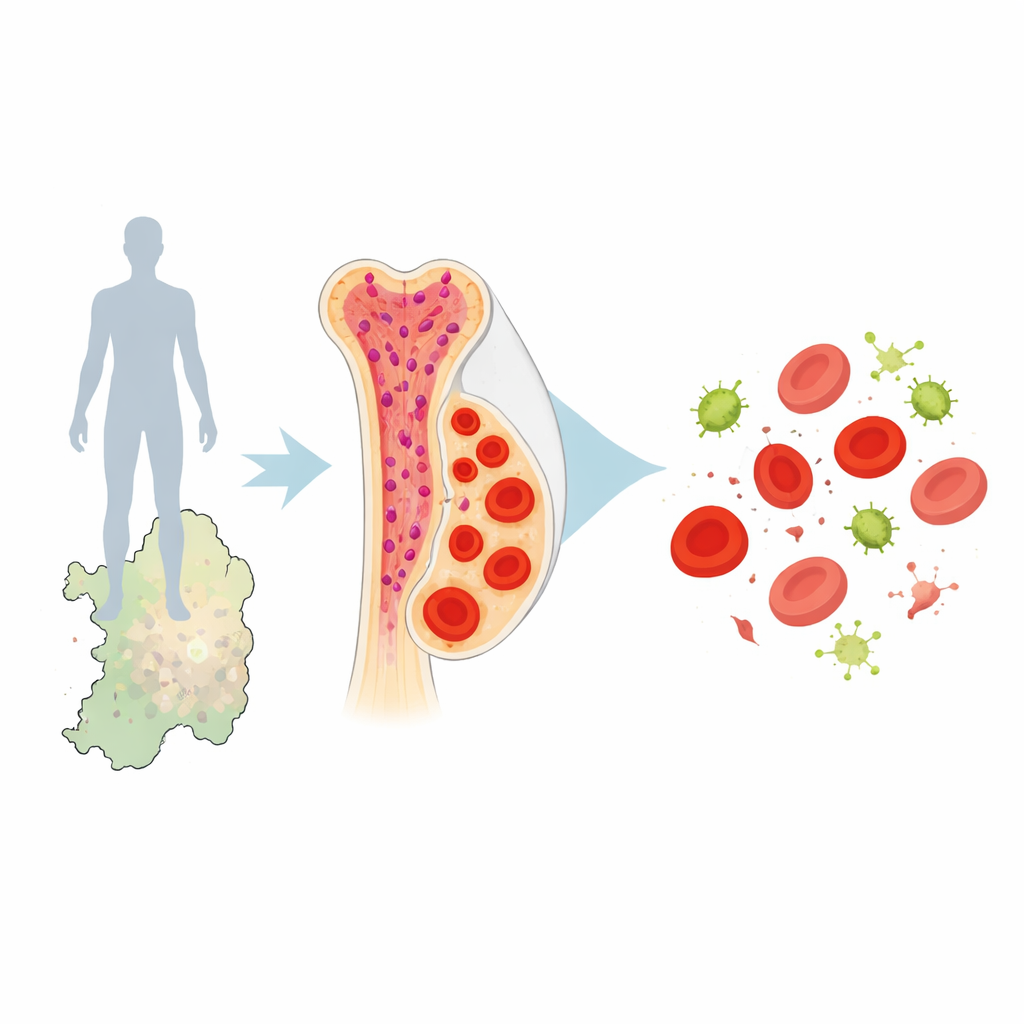
A Tiny DNA Change with Big Consequences
The researchers focused on a region of our DNA that controls a protein called cyclin D3, which helps immature blood cells divide. In earlier work, a genetic variant named rs112233623-T near the CCND3 gene was linked to red blood cells that are fewer in number but larger in size, and to higher levels of certain forms of hemoglobin. This variant is about ten times more common in people from Sardinia than in many other European groups, echoing the island’s long history as a malaria hotspot. The team asked a series of connected questions: how does this variant change blood cell development, why is it so common in Sardinia, and does it actually hinder malaria parasites?
Slowing the Cell-Cycle Factory for Red Blood Cells
To see what the variant does inside cells, the scientists grew red blood cell precursors from volunteers who either carried two copies of rs112233623-T or two copies of the usual version. In cells with the variant, cyclin D3 levels were clearly lower, and the cells moved more slowly through the stage of the cycle where DNA is copied and cells divide. As a result, each precursor cell went through fewer rounds of division before maturing, producing a blood profile with fewer but larger red blood cells—very similar to what had been seen in mice completely lacking cyclin D3. In genetic tests across thousands of Sardinian volunteers, the rs112233623-T variant emerged as the main driver of this blood-cell pattern.
Rewiring a Genetic Switch in Blood Precursors
The crucial DNA change sits in an “on–off” control region, or enhancer, that boosts CCND3 activity in developing red cells. The team showed that swapping in the rs112233623-T version sharply weakened this enhancer in laboratory reporter tests. By dissecting the surrounding sequence, they found that the normal DNA sequence forms a landing pad for a protein called SMAD3, which turns CCND3 on. The T version disrupts this landing pad and instead favors binding of GATA1, a protein that acts more like a brake in this context. In real blood precursors, SMAD3 bound strongly to the normal sequence but poorly to the variant, and drugs that blocked SMAD-type signals caused cyclin D3 levels to fall. Together, these experiments reveal a simple logic: fewer SMAD3 “go” signals and more GATA1 “stop” signals mean less CCND3, slower cell division, and altered red blood cell output.
An Evolutionary Signature of Past Malaria
Why did this seemingly disadvantageous variant become common in Sardinia? Population-genetic analyses provided a clue. Compared with other Europeans, Sardinians show unusually high frequency of rs112233623-T, long stretches of DNA around it with little variation, and patterns that are best explained by recent positive selection rather than chance. Using models of how gene variants rise in frequency over generations, the authors estimated that rs112233623-T has been strongly favored in Sardinia’s recent past. Because the island endured intense malaria transmission until the mid-twentieth century, the authors reasoned that protection against malaria was the most likely benefit.
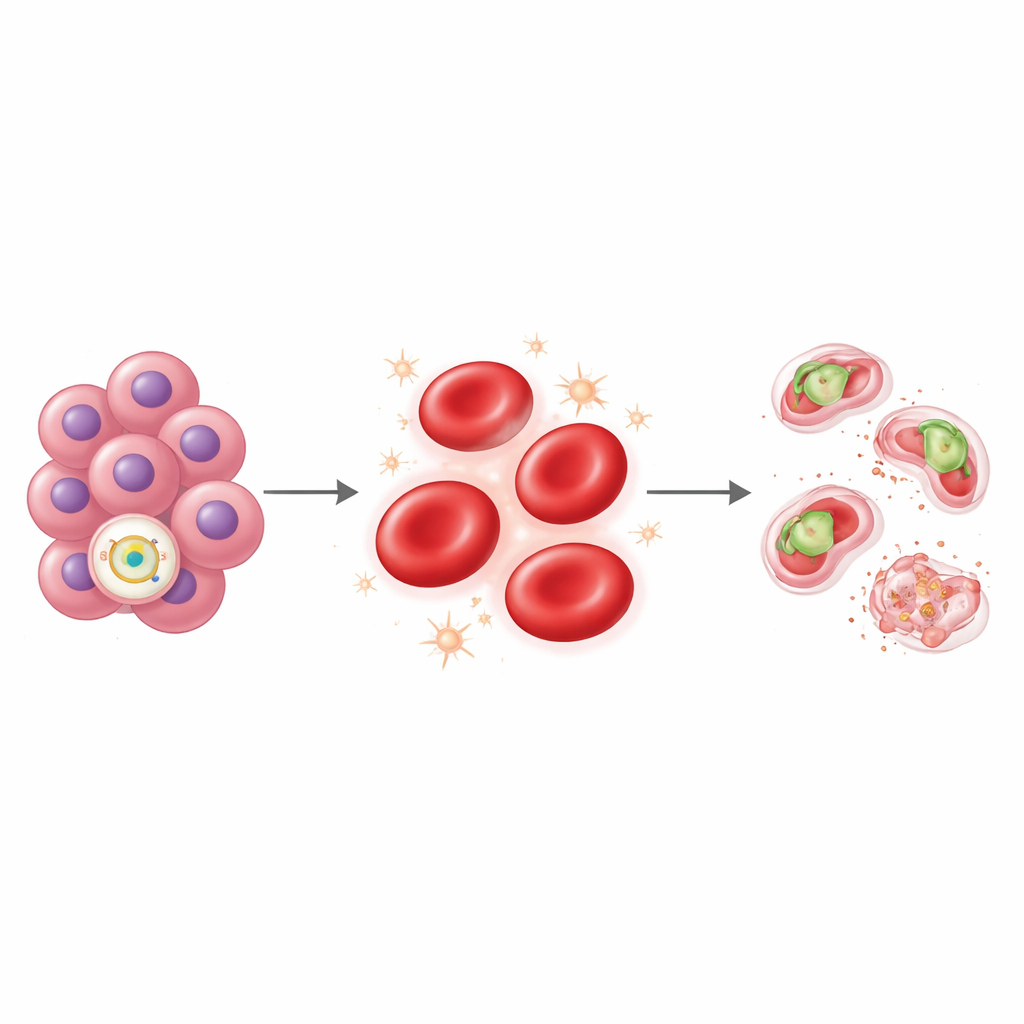
Stressing the Parasite to Death Inside the Cell
To test that idea directly, the team infected red blood cells from carefully genotyped Sardinian volunteers with Plasmodium falciparum, the parasite that causes the deadliest form of malaria. Red cells from people carrying the rs112233623-T variant allowed much poorer parasite growth over several cycles than cells from those without it. Parasites in these cells often stalled and died instead of completing their usual developmental stages. Measuring the chemistry inside the red cells, the researchers found higher levels of reactive oxygen species—molecules that cause oxidative stress—in variant carriers. The higher the oxidative stress, the less the parasites grew, forming a tight inverse relationship. Strikingly, this same stress-based handicap resembled what is seen in people with a well-known protective trait: deficiency in the enzyme G6PD, which has long been linked to resistance against severe malaria.
What This Means for Future Malaria Control
In plain terms, the study shows that dialing down cyclin D3 in red blood cell precursors makes the resulting cells a harsher environment for malaria parasites, largely by boosting internal “rusting” chemistry that the parasite cannot fully withstand. This gentle, inherited slowdown in red cell production appears to have been rewarded by natural selection in Sardinia because it reduced the risk of severe, high-parasite infections. The work suggests that drugs that temporarily mimic this genetic effect—by inhibiting CCND3 in the bone marrow—could complement existing antimalarial treatments, tipping the balance further against the parasite while staying within the limits of what the human body can tolerate.
Citation: Marini, M.G., Mingoia, M., Steri, M. et al. Reduced cyclin D3 expression in erythroid cells protects against malaria. Nature 651, 698–706 (2026). https://doi.org/10.1038/s41586-026-10110-9
Keywords: malaria resistance, red blood cells, human evolution, genetic variant, oxidative stress