Clear Sky Science · en
Developmental convergence and divergence in human stem cell models of autism
Why this research matters for families and society
Autism spectrum disorder is famously complex: hundreds of different genes have been tied to increased risk, yet many autistic people share similar challenges in communication, social interaction and behavior. This study asks a deceptively simple question with big implications: when very different genetic changes increase the chance of autism, do they ultimately disturb the developing human brain in similar ways? Using tiny lab-grown models of the human cortex built from patients’ own cells, the researchers trace how early glitches in gene activity can funnel into common pathways that alter how brain cells are born, mature and connect.
Growing miniature human brain tissue in the lab
To tackle this question, the team collected skin or blood cells from 55 people: some had well-characterized rare mutations strongly linked to autism, others had autism without a known major mutation, and others were unaffected controls. These cells were reprogrammed into induced pluripotent stem cells, which can become almost any cell type. The scientists then guided them to form three-dimensional “cortical organoids” – tiny balls of tissue that mimic early development of the human cerebral cortex. Over 100 days, these organoids progressed from mostly stem-like progenitor cells to networks rich in young neurons. At four points along this timeline (days 25, 50, 75 and 100), the team measured which genes were switched on or off across hundreds of organoids, creating a detailed movie of molecular changes during early brain development.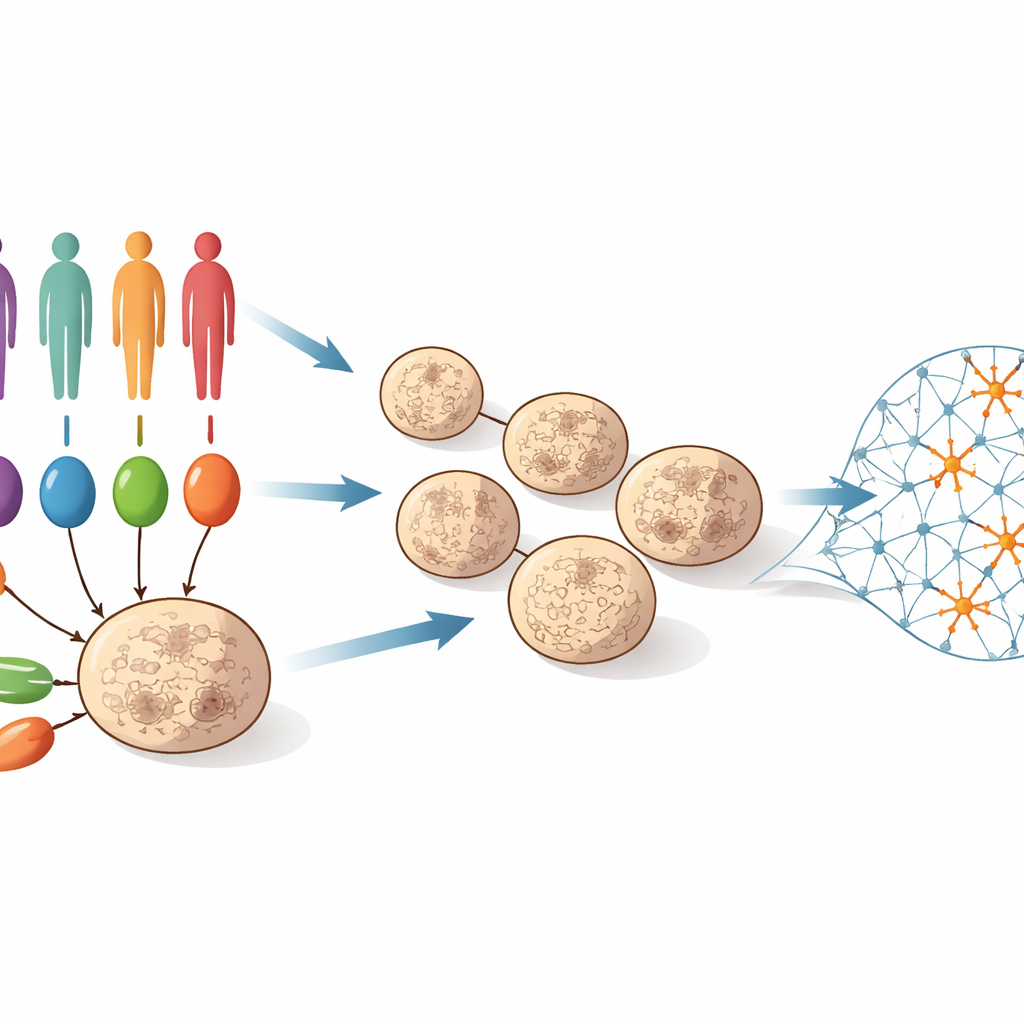
Many genetic paths, shared early detours
Although the autism-linked mutations were very different – including large missing or duplicated DNA segments and specific gene changes – organoids carrying the same mutation showed highly reproducible gene activity patterns. Early in development, especially around day 25, each mutation produced its own strong signature: hundreds to more than a thousand genes were misregulated compared with controls. Yet even at this early stage, several mutations altered overlapping sets of genes involved in how DNA is packaged and read inside cells. These shared changes affected known high-risk autism genes and other neurodevelopmental disorder genes, suggesting that different mutations begin to disturb similar cellular machinery very early in brain formation.
Convergence as the brain model matures
As the organoids developed toward more mature neurons (by day 75 and 100), the picture shifted. The mutation-specific differences became less distinctive, while the similarities grew stronger. Across almost all genetic forms of autism studied, the researchers saw reduced activity of genes important for electrical signaling at synapses – the communication points between neurons – and increased activity of genes tied to cell growth and protein production. Subtle shifts in the proportions and maturation stages of different cell types emerged, pointing to changes in the timing of neuron birth and maturation rather than wholesale loss of particular cells. By the latest time point, thousands of genes showed a shared pattern of dysregulation across the various rare mutations, even though the mutations themselves affected different original genes.
A central control hub that links many autism genes
Digging deeper, the team used network analyses to group genes that tended to turn on and off together into “modules.” One module in particular, called M5, stood out. Its genes are most active early in development and are heavily enriched for autism risk genes, especially those that control how other genes are switched on or off and how DNA is packaged. In organoids from several autism mutation groups, this module was consistently turned down. The researchers showed that many of the proteins in M5 physically interact, forming a regulatory hub. Using CRISPR-based tools to selectively damp down 26 of these M5 regulators in human neural progenitor cells, they confirmed that this hub directly controls large sets of downstream genes, including many additional autism and neurodevelopmental risk genes, and that disrupting it can alter pathways involved in neuron formation and synapse function.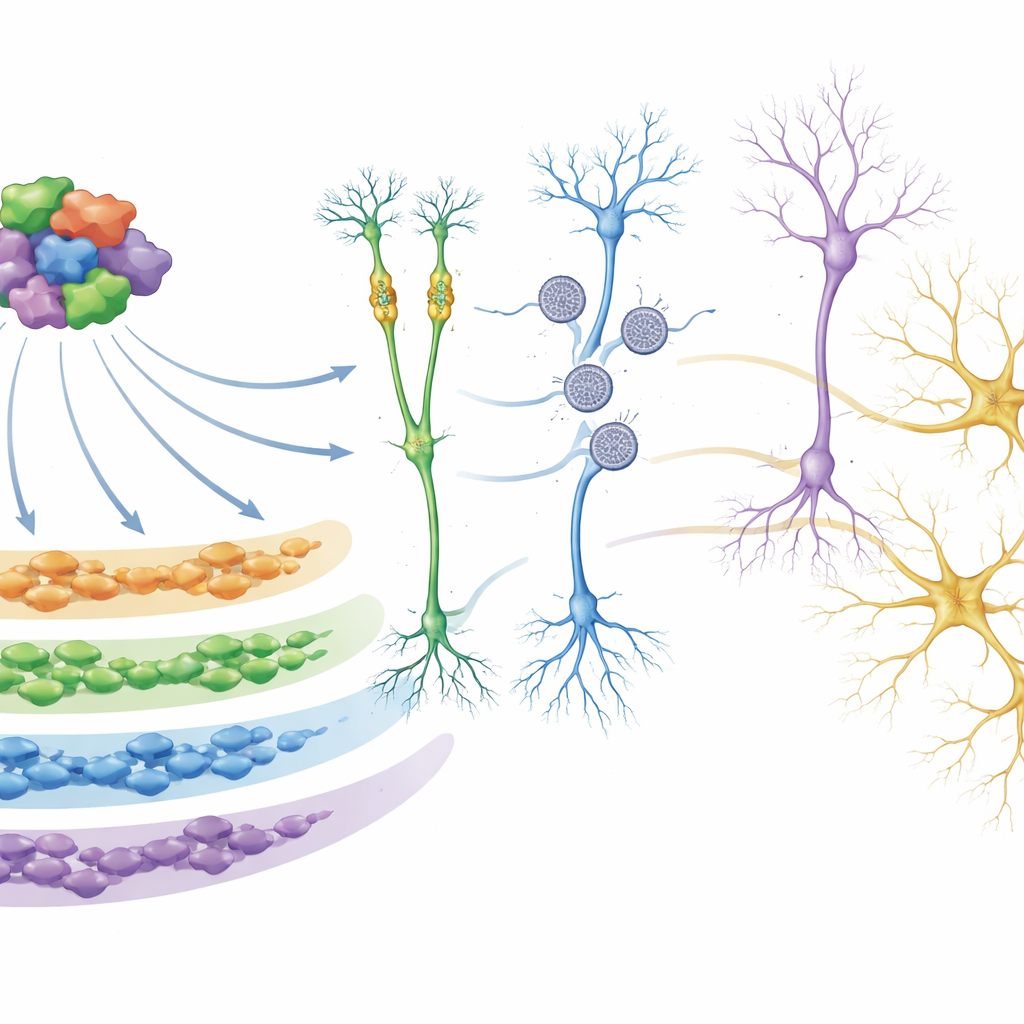
What this means for understanding autism
This work suggests a unifying picture: rare, powerful autism-linked mutations start by disturbing different pieces of the genome, but their effects ripple forward through a shared early control network that governs how cortical neurons are generated and wired. Over time, those ripples converge on common outcomes – delayed or altered neuronal maturation and disrupted synaptic programs – even though the initial genetic causes differ. For families, this means that very different diagnoses at the DNA level may still influence overlapping biological pathways in the developing brain. For researchers and clinicians, the identified regulatory network offers a focused set of molecular targets for future therapies aimed at restoring more typical patterns of brain development across a wide array of autism-related genetic conditions.
Citation: Gordon, A., Yoon, SJ., Bicks, L.K. et al. Developmental convergence and divergence in human stem cell models of autism. Nature 651, 707–719 (2026). https://doi.org/10.1038/s41586-025-10047-5
Keywords: autism spectrum disorder, brain organoids, stem cell models, gene regulation, neurodevelopment