Clear Sky Science · en
Construction of complex and diverse DNA sequences using DNA three-way junctions
Building New Genetic Stories
Modern biology can read and edit DNA with astonishing speed, yet actually writing long, custom-made genetic sequences still lags behind. This gap slows everything from designing new medicines to creating greener materials. This study introduces “Sidewinder,” a new way to stitch DNA pieces together that aims to make writing complex, tailor‑made genes as reliable and scalable as reading them.
Why DNA Assembly Needs a Rethink
Every cell runs on DNA, a long string of chemical letters that encode life’s instructions. Scientists can chemically make only short stretches of DNA, so longer genes must be assembled from many small pieces, like sentences built from cut-up words. Existing methods all use matching edges on these pieces to guide which ones stick together. But those matching edges become part of the final DNA, which means they cannot be freely optimized for flawless assembly without also changing the gene itself. As designs get longer and more intricate, this built‑in compromise leads to more mistakes, lower yields and practical limits on what can be built.
A Side Path That Guides Without Leaving a Trace
Sidewinder tackles this problem by adding a third helper strand of DNA that never appears in the finished product. DNA pieces are prepared with two features at their ends: short “toeholds” that will eventually form the seamless final connection, and longer “barcodes” that are designed only to talk to their intended partners. When mixed together at a controlled temperature, the barcodes on neighboring pieces find one another and wrap into a temporary side helix, forming a three‑way junction that pulls the matching toeholds into place. An enzyme then seals the main DNA pieces together. Finally, the helper barcodes are stripped away, leaving behind a clean, continuous sequence with no extra scars or tags. 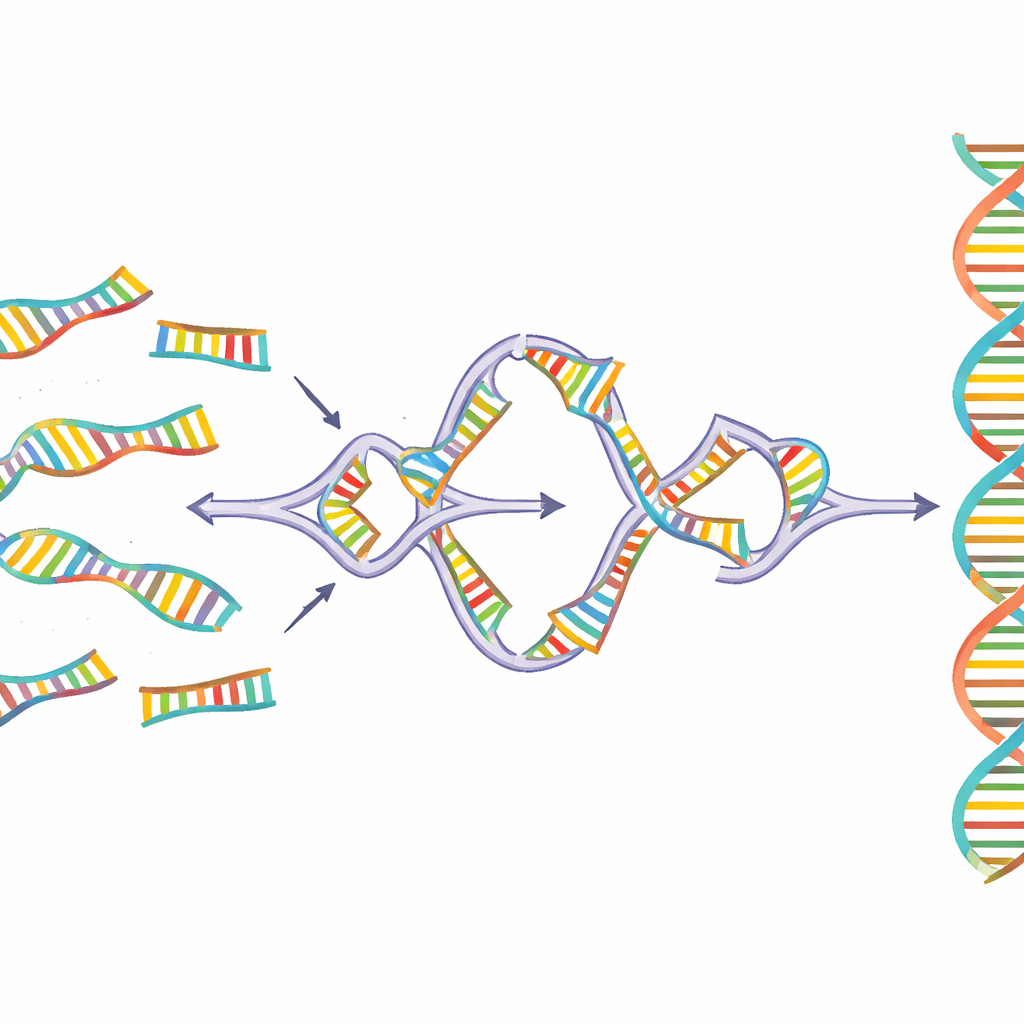
From Dozens of Pieces to Difficult Genes
To show what Sidewinder can handle, the authors built DNA constructs from 5, 10, 20 and even 40 separate fragments in a single reaction. Competing state‑of‑the‑art methods faltered beyond just a handful of pieces, producing messy mixtures or failing outright, while Sidewinder consistently yielded a single, correctly sized product. Long‑read sequencing confirmed that, in a 40‑fragment test, more than 96% of reads were true Sidewinder products and every one of those was assembled in the perfect order. The team then pushed the method on “hard‑mode” sequences: a human gene with extremely high content of the letters G and C, and a silk‑like protein segment packed with repeats. Such sequences often defeat standard assembly because they stick to themselves in confusing ways. Sidewinder still produced near‑perfect assemblies, even when all junctions deliberately shared the same toehold sequence, something that would be almost impossible to control with older techniques.
Many Genes at Once and Oceans of Variants
Because Sidewinder’s barcodes uniquely define who can pair with whom, multiple genes can be built in the same tube without cross‑contamination. The researchers mixed fragments for three differently colored marker proteins and assembled them in one pot. With the right primers, they could selectively amplify any one gene or the mixture, and sequencing showed that incorrect crossings between designs were extremely rare. 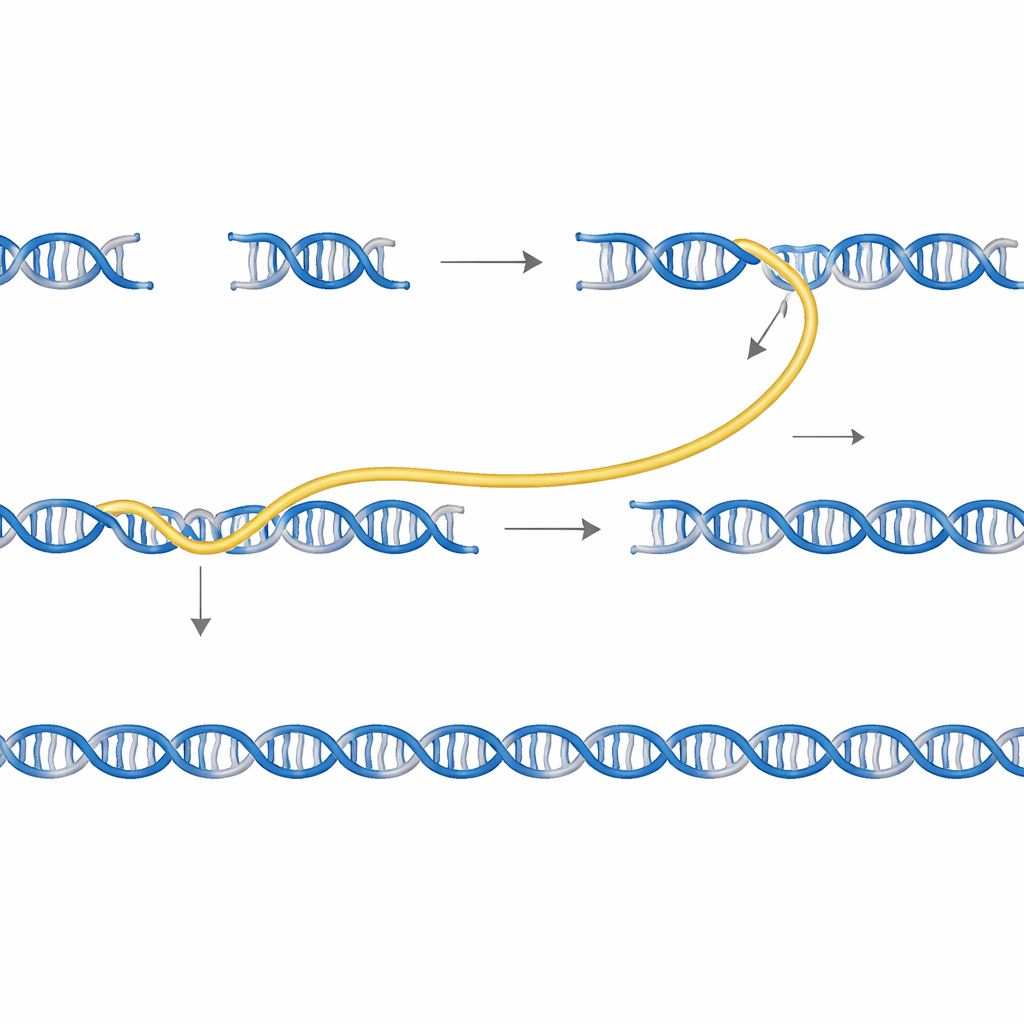
What This Means for Future Bioengineering
Sidewinder’s key achievement is to separate the “assembly instructions” from the final DNA story. By moving the guiding information onto a removable side strand, scientists can design joins that are extraordinarily specific and reliable without being forced to compromise the gene itself. The result is a general‑purpose way to build long, tricky and highly varied DNA sequences with accuracy that rivals, and in some respects improves upon, the quality of the starting pieces. As tools like AI increasingly propose bold new genetic designs, techniques like Sidewinder may become essential for turning those designs into real molecules for medicine, materials, agriculture and beyond.
Citation: Robinson, N.E., Zhang, W., Ghosh, R. et al. Construction of complex and diverse DNA sequences using DNA three-way junctions. Nature 651, 491–500 (2026). https://doi.org/10.1038/s41586-025-10006-0
Keywords: DNA assembly, synthetic biology, gene libraries, DNA nanotechnology, protein engineering