Clear Sky Science · en
Language model-guided anticipation and discovery of mammalian metabolites
Hidden Chemistry Inside Our Bodies
Every drop of blood or urine contains thousands of tiny molecules that reflect what we eat, how we live and whether we are getting sick. Yet for most of these molecules, scientists do not know their names or what they do. This paper introduces DeepMet, an artificial intelligence system that reads the "language" of these molecules and predicts which ones are missing from our current maps of human and animal chemistry. By guiding experiments toward the most promising candidates, DeepMet helps researchers uncover this chemical dark matter and better understand how our bodies work.
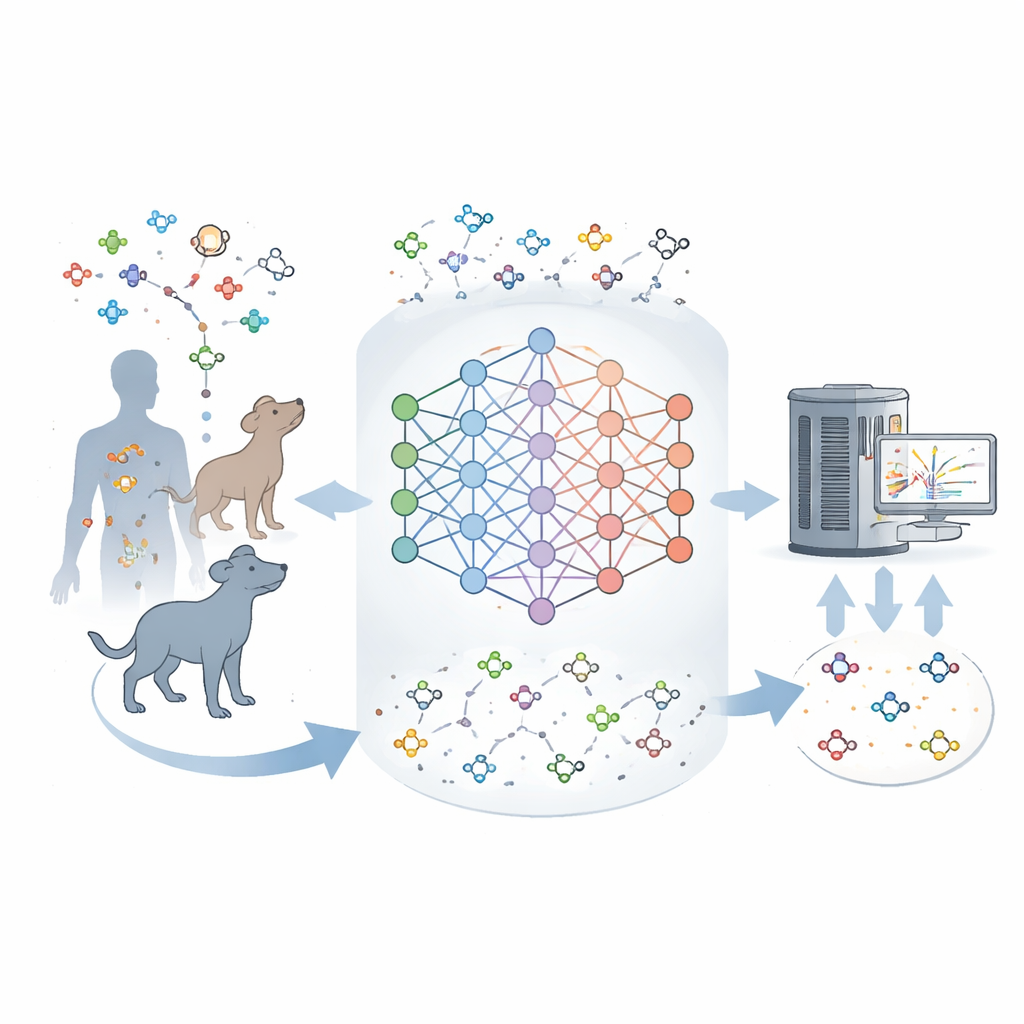
Why So Many Molecules Remain Unknown
Modern instruments can weigh and partially fingerprint thousands of molecules in a tissue sample at once. But turning these fingerprints into exact structures is hard. Existing databases list many known metabolites, yet most of the signals seen in real samples do not match anything in those catalogs. This gap suggests that current maps of metabolism are incomplete and that many natural molecules in mammals have never been described. The authors set out to build a tool that could learn from known metabolites and then imagine the most plausible missing ones, in much the same way that language models predict likely words in a sentence.
Teaching a Machine the Grammar of Metabolism
The team trained a neural network called DeepMet on about 2,000 well-established human metabolites, encoding each as a short string that describes its structure. After initial training on drug-like molecules to learn general chemical rules, DeepMet was fine-tuned on this metabolite set. When asked to generate new structures, the model produced molecules that occupied the same regions of chemical space as real metabolites and even reproduced many known types of enzyme reactions, despite never being told those rules explicitly. In other words, DeepMet appeared to internalize the unwritten grammar that links basic building blocks such as sugars and amino acids into biologically realistic small molecules.
Predicting Which New Molecules Probably Exist
The researchers then sampled one billion candidate molecules from DeepMet and counted how often each unique structure appeared. Frequently repeated structures tended to look more like known metabolites, share common chemical cores with them, and match plausible enzyme transformations. To test whether these high-frequency candidates correspond to real molecules, the team compared DeepMet’s predictions against metabolites that were added to the Human Metabolome Database after the model’s training data had closed. DeepMet had already generated most of these later discoveries and ranked many of them among its most likely candidates. From the thousands of top-ranked, database-absent structures, the authors purchased or synthesized 80 and checked real human samples by mass spectrometry. They confirmed the presence of several previously unrecognized metabolites, some of which had been overlooked even though they appear in existing literature.
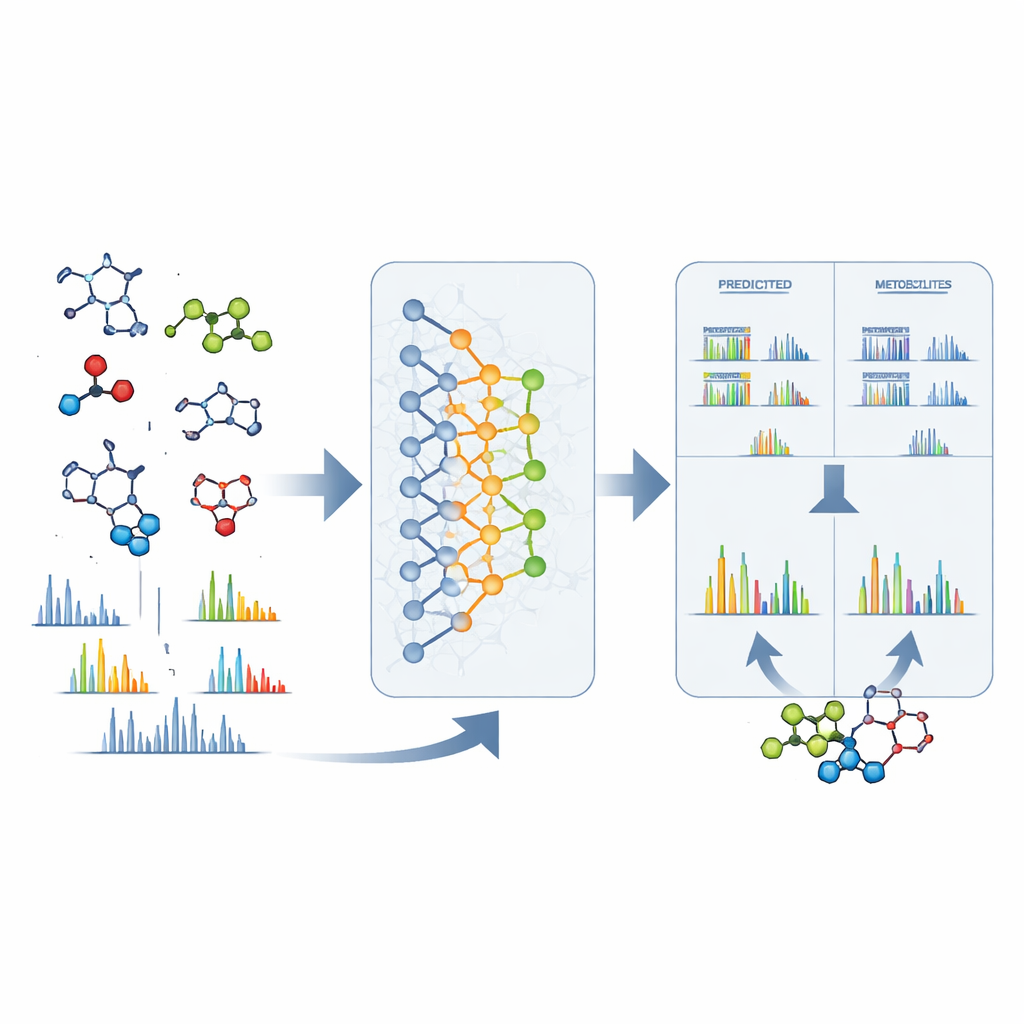
From Raw Signals to Concrete Structures
DeepMet is also useful once an unknown peak has been seen in a mass spectrometer. Given only the exact mass of a mystery molecule, the model can list many structures that would weigh the same and rank them by how metabolite-like they seem. In nearly a third of test cases, the correct structure came out on top; in many more, it appeared among only a handful of high-ranking candidates and was usually very similar in shape to the model’s favorite. To further narrow things down, the authors combined DeepMet with separate software that predicts how each candidate would break apart in a mass spectrometer. Matching these predicted patterns to real experimental spectra roughly doubled identification accuracy. Searching large public datasets with this combined approach yielded tentative structures for many previously anonymous signals and pointed to metabolites that differ across diseases, diets and microbiome states.
Illuminating the Chemical Dark Matter of Life
By blending chemical intuition learned from data with powerful pattern-matching against mass spectra, DeepMet provides a roadmap for discovering new metabolites in a targeted, practical way. It cannot yet reveal every unknown molecule—some structures lie too far from those it has seen, and certain isomers remain indistinguishable without specialized methods. But the study shows that language-model style tools can not only invent realistic molecules, they can also anticipate real compounds that biologists will later confirm in animals and humans. For a layperson, the takeaway is that AI can now help chemists systematically uncover hidden chemistry in our bodies, potentially revealing new biomarkers, tracing diet–microbe–host links, and gradually turning today’s metabolic dark matter into tomorrow’s well-charted biology.
Citation: Qiang, H., Wang, F., Lu, W. et al. Language model-guided anticipation and discovery of mammalian metabolites. Nature 651, 211–220 (2026). https://doi.org/10.1038/s41586-025-09969-x
Keywords: metabolomics, chemical language models, DeepMet, mass spectrometry, metabolic dark matter