Clear Sky Science · en
Polyamine-dependent metabolic shielding regulates alternative splicing
How Tiny Molecules Help Cells Read Their Genes
Inside every cell, the same DNA script can be read in many ways, allowing a limited number of genes to produce a huge variety of proteins. This flexibility depends on a process called alternative splicing, which edits RNA messages before they are turned into proteins. This paper reveals that small, positively charged molecules called polyamines quietly steer this editing step, acting as a kind of protective shield that helps cells decide which RNA versions to make. Because alternative splicing is crucial in cancer, brain function, and stem cells, uncovering this hidden layer of control has wide-reaching implications for health and disease.
Small Charged Helpers in Every Cell
Polyamines are tiny, flexible molecules that carry multiple positive charges. Cells make them from basic nutrients, and they have long been known as helpers for growth and survival, especially in fast-dividing cells like tumors. The authors asked whether polyamines also serve as signals, not just as fuel. By partially blocking polyamine production in prostate cancer cells and in mice, they measured how proteins and RNA changed over time. They found that long before total polyamine levels dropped dramatically, there was a strong wave of changes in protein phosphorylation—chemical tags that turn protein switches on or off—especially on components of the cellular RNA editing machinery, the spliceosome.
Rewiring the Cell’s RNA Editor
Looking specifically at RNA, the team used deep sequencing to track alternative splicing when polyamine synthesis was inhibited. Hundreds of RNA segments were skipped or included differently, in cancer cell lines, normal cell types, and mouse tissues. These changes were not simply a side effect of slower cell division or of another known polyamine-dependent process called hypusination. Instead, when the researchers re-supplied cells with outside polyamines, many splicing shifts snapped back toward normal. Both drugs and genetic tools that lowered polyamines produced similar splicing patterns, and further lowering polyamines with a drug combination made the changes even stronger, confirming that the effect was tightly linked to polyamine availability.
A Hidden Target: The SF3 Splicing Module
To pinpoint where in the splicing machinery polyamines act, the authors compared the splicing signature caused by lowering polyamines with a large reference map generated by individually silencing more than 300 known splicing factors. The closest match pointed to a specific part of the spliceosome called the SF3 subcomplex, which helps recognize key signals in RNA. Computational analysis of public protein–RNA binding data supported this link: RNAs whose splicing was altered by polyamine loss were especially likely to be bound by SF3 proteins. When the team partially disabled SF3 components genetically or with a drug, the splicing effects of polyamine depletion were largely erased, showing that intact SF3 activity is required for this new regulatory pathway. 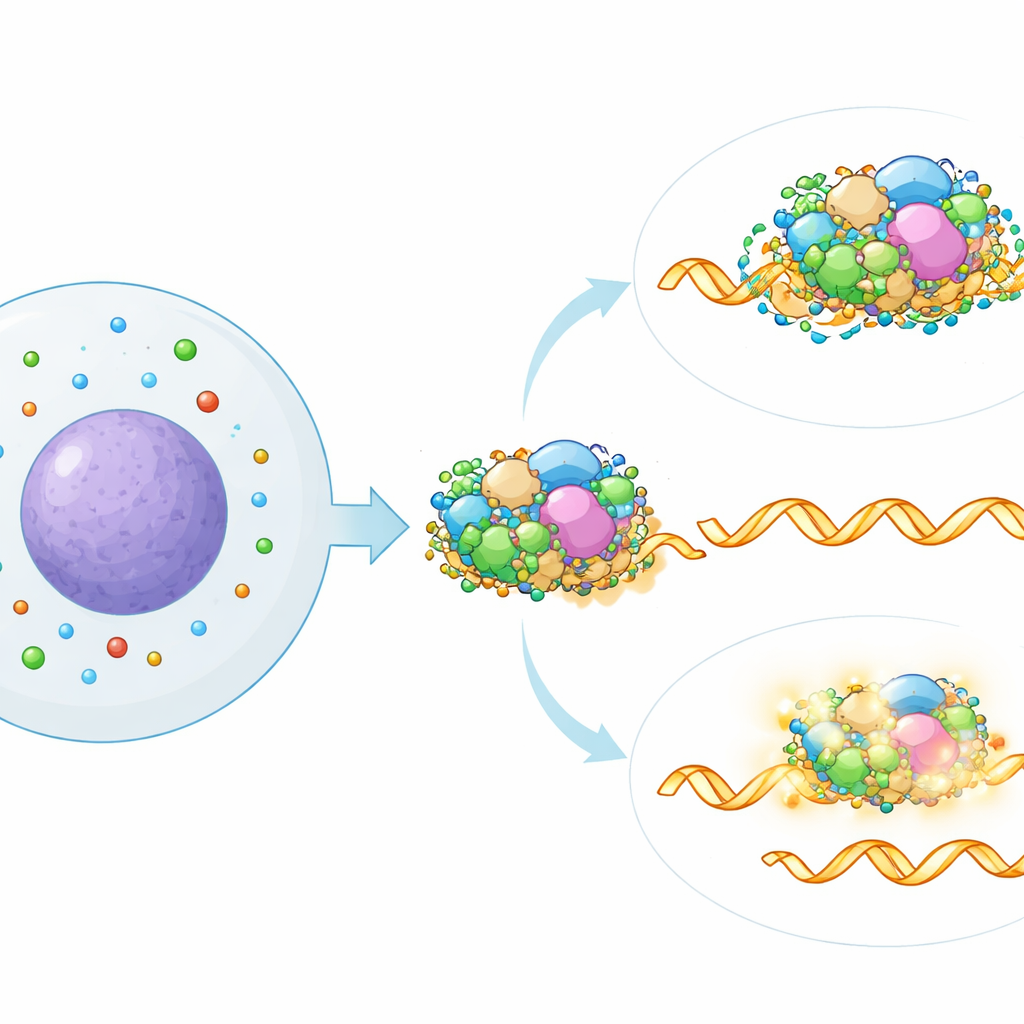
Metabolic Shielding: How Polyamines Block Protein Tagging
Diving deeper, the researchers noticed that the most strongly affected phosphorylation sites on SF3 proteins clustered within short stretches rich in negatively charged amino acids. Molecular modeling and nuclear magnetic resonance experiments showed that polyamines nestle into these acidic patches, forming multiple electrostatic contacts and partially covering nearby serine residues where phosphate groups are usually added. This physical "hug" reduces how accessible these spots are to protein kinases—the enzymes that attach phosphates. In test-tube experiments, adding polyamines directly blocked a kinase called CK1 from phosphorylating SF3 proteins. In cells, inhibiting CK1 and its close relative CK2 dampened the splicing changes caused by polyamine loss, and specially engineered mice whose SF3A3 protein lacked three key phosphorylation sites became largely insensitive to polyamine-dependent splicing shifts. 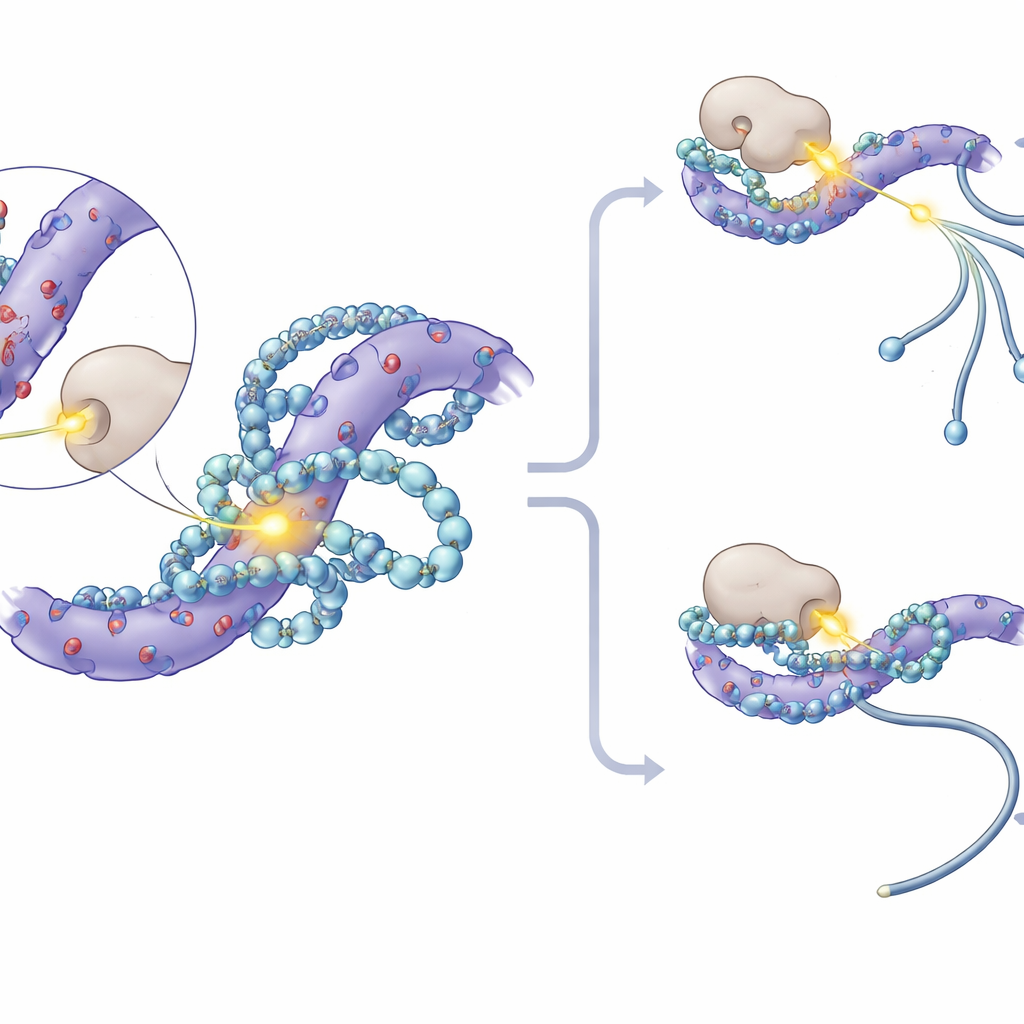
A Designer Molecule That Separates Roles
The study also introduces a polyamine-like drug, BENSpm, that is more strongly positively charged but simultaneously shuts down the cell’s own polyamine production. BENSpm bound tightly to SF3 acidic patches and blocked kinase action, just like natural polyamines, yet it did not rescue cell growth when real polyamines were scarce. This allowed the authors to separate classic polyamine functions (supporting growth and metabolism) from the newly defined shielding role. In mouse embryonic stem cells, depleting polyamines pushed cells toward losing their "stemness" marker Nanog and altered their splicing landscape. BENSpm restored both the splicing profile and Nanog expression despite ongoing suppression of normal polyamine synthesis, indicating that metabolic shielding is a key requirement for maintaining stem cell identity.
Why This Discovery Matters
In everyday terms, this work shows that polyamines act as tiny protective gloves around sensitive parts of key splicing proteins. When the gloves are on, kinases cannot easily grab and tag those sites, and the cell’s RNA editing stays on course. When polyamines are low, the gloves come off, phosphorylation surges, and splicing patterns shift, with consequences for cancer cells and stem cells alike. By defining this "metabolic shielding" mechanism and by providing a tool molecule that selectively mimics it, the study opens the door to new ways of tuning gene reading without changing the DNA itself, potentially inspiring future therapies that target splicing in cancer and regenerative medicine.
Citation: Zabala-Letona, A., Pujana-Vaquerizo, M., Martinez-Laosa, B. et al. Polyamine-dependent metabolic shielding regulates alternative splicing. Nature 651, 819–828 (2026). https://doi.org/10.1038/s41586-025-09965-1
Keywords: polyamines, alternative splicing, RNA processing, metabolic signaling, stem cell regulation