Clear Sky Science · en
Ligand-specific activation trajectories dictate GPCR signalling in cells
How One Receptor Can Behave Like Many Switches
Many of today’s medicines act on a huge family of cell-surface proteins called G-protein–coupled receptors, or GPCRs. These receptors influence heart rate, mood, breathing, and countless other body functions. For decades they were treated as simple on–off switches: a drug binds, the switch flips, and a signal flows inside the cell. This paper shows that reality is far richer. Using a new kind of fluorescent "spy" built directly into a receptor, the authors watch, in living cells, how different drugs send the same receptor along distinct activation paths—like choosing different routes through a city—to produce different patterns of signalling. Understanding these hidden routes could help design drugs that give the desired benefit while avoiding side effects.
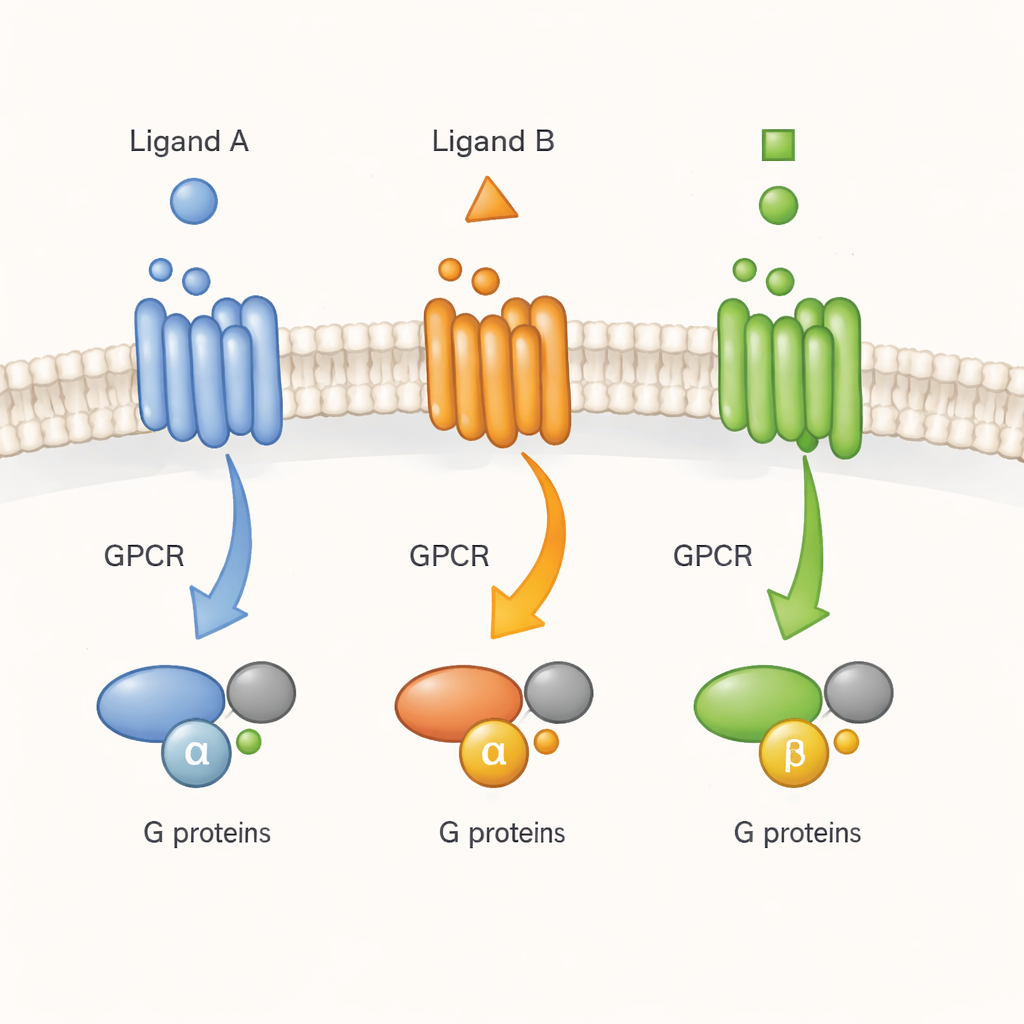
Watching a Cell’s Doorbell in Real Time
GPCRs are often described as molecular doorbells: a signal molecule (a ligand) rings from the outside, and the receptor passes the message to G proteins on the inside. Earlier work with purified receptors in detergents had suggested that GPCRs do not simply flip between one inactive and one active shape. Instead, they sample many shapes that drugs can stabilize to different degrees. But it was unclear whether the same complexity exists in the crowded, dynamic environment of a living cell. Here, the authors focus on one representative receptor, the M2 muscarinic acetylcholine receptor, an important regulator of heart and nerve activity, and ask whether different ligands drive it into distinct active forms that matter for real cellular signalling.
Building Tiny Light Reporters on the Receptor Surface
To follow the receptor’s movements without disturbing its normal function, the team used genetic code expansion, a technology that lets them insert a special designer amino acid at selected positions on the receptor’s outer surface. This chemical “anchor” can be clicked to a small fluorescent dye in living cells. By scanning 72 positions and keeping only those that still behaved like normal receptors, they built a panel of seven M2 receptor variants, each carrying a single dye at a different outer-loop site. When the natural messenger acetylcholine was applied, the dye brightness at these sites rose or fell in characteristic ways, revealing how each part of the receptor’s outer surface shifted as signalling began. Crucially, these labelled receptors could still activate G proteins and undergo normal internalization, showing that the reporters were faithful rather than disruptive.
Drugs Leave Distinct “Conformational Fingerprints”
The researchers then compared several drugs that all turn on the M2 receptor but with different strengths: the body’s own acetylcholine, a super-strong synthetic agonist called iperoxo, and two weaker partial agonists, arecoline and pilocarpine. Each drug produced a unique pattern of fluorescence changes across the seven reporter sites—a conformational fingerprint. At most positions, the size of the movement tracked how strongly the drug activated the receptor. At two sites, however, the relationship was inverted: the weaker drugs produced the largest changes, and the strongest drug produced almost none. Such behaviour cannot be explained by a single active state. Instead, it indicates that the same receptor, in living cells, can adopt several distinct active shapes, some favoured by strong drugs, others by weak ones.
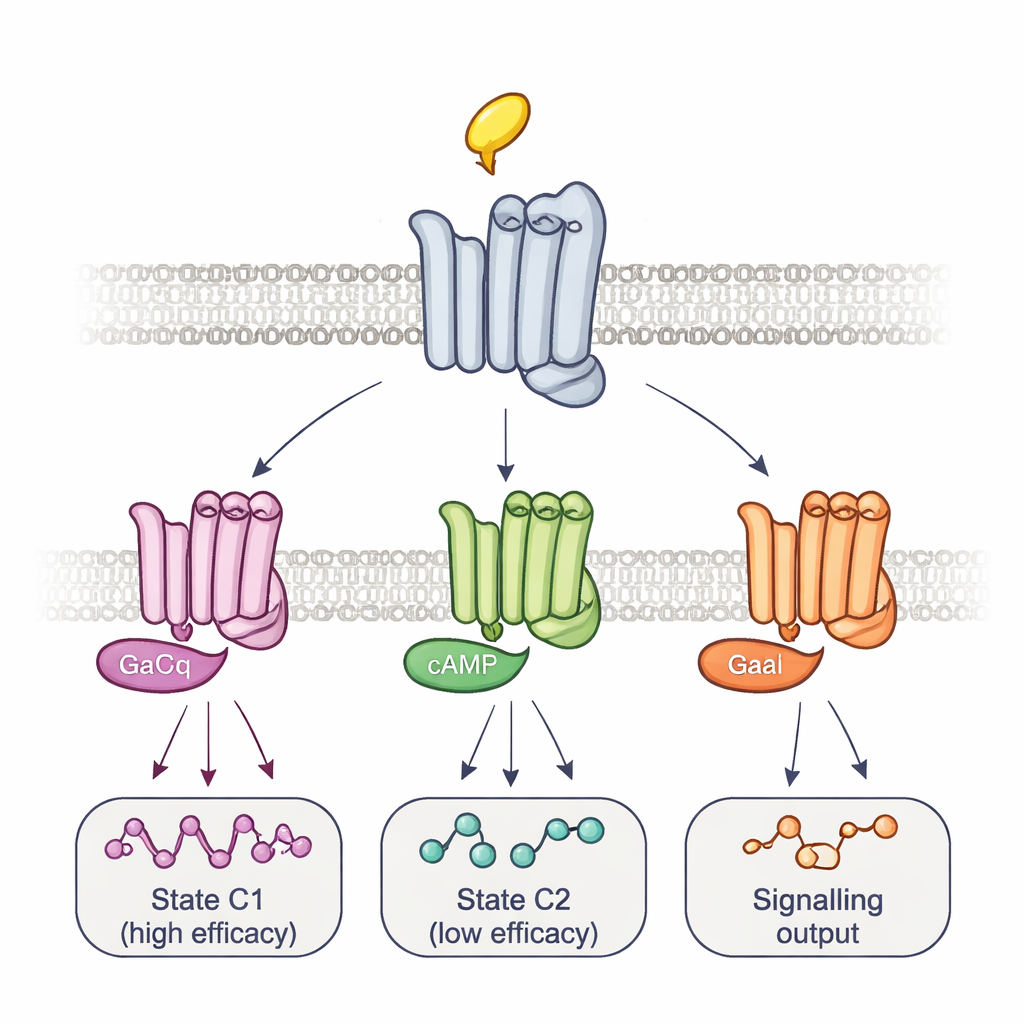
Multiple Complexes and Timed Routes to Signalling
To connect these shapes to actual signalling, the team altered the G proteins themselves. Overexpressing a mutant G protein that forms very tight, long-lived complexes with receptors selectively erased the signal from some reporter sites while enhancing others. This pattern, together with the timing of the fluorescence changes, revealed at least two main receptor–G-protein complexes: a fast-forming, high-efficacy complex and a slower, low-efficacy one. Different drugs shifted the balance between these complexes and even used different intermediate steps to get there, tracing drug-specific activation trajectories. Using a separate bioluminescence assay that monitors a panel of 14 G-protein subtypes, the authors showed that these equilibria help determine not just how strongly a drug activates signalling overall, but which exact G proteins are turned on. For example, arecoline preferentially activated certain Go proteins, whereas pilocarpine strongly favoured the low-efficacy complex.
Why This Matters for Better Medicines
For non-specialists, the key message is that a single receptor is not just one switch but a cluster of related switches, each reachable by different paths and each wired to slightly different downstream effects. This study provides a direct view of those paths and states in intact cells, rather than in simplified test-tube systems. By mapping how particular drugs bias the receptor toward specific complexes and G-protein partners, researchers gain a blueprint for designing “smarter” medicines—compounds that nudge receptors into states that drive helpful signals while avoiding those linked to side effects. The fluorescent biosensor strategy developed here should be adaptable to many other receptors, opening a window onto the real-time choreography of drug action in living cells.
Citation: Thomas, R., Jacoby, P.S., De Faveri, C. et al. Ligand-specific activation trajectories dictate GPCR signalling in cells. Nature 650, 1053–1062 (2026). https://doi.org/10.1038/s41586-025-09963-3
Keywords: GPCR signalling, ligand efficacy, G proteins, conformational biosensors, drug discovery