Clear Sky Science · en
Resolving DNA origami structural integrity and pharmacokinetics in vivo
Watching Tiny DNA Machines Inside the Body
DNA can be folded like paper into tiny three-dimensional shapes, a technique known as DNA origami. These nanostructures can be programmed to carry drugs, vaccines or signaling molecules with exquisite precision. But once they are injected into the body, do they stay intact long enough to work, or do they quickly fall apart? This article introduces a new way to watch these DNA machines in action inside living animals, revealing how long they survive, how they travel through the bloodstream and even which parts of each structure break first.
Why Folding DNA Could Change Medicine
DNA origami takes advantage of the same base-pairing rules that store our genetic information, but repurposes them to build nanoscale objects. Researchers can arrange proteins, drugs or immune-stimulating signals on these objects like studs on a Lego plate, creating highly controlled therapies for cancer, autoimmune disease and infections. For such precision treatments to be safe and effective, regulators and scientists must know how intact these structures remain in the body and how fast they are cleared—together known as their pharmacokinetics. Existing tracking methods usually attach fluorescent dyes or simply count DNA fragments, which can make disassembled debris look like fully intact nanostructures. Until now, there has been no practical way to tell, in living animals, whether the original DNA origami shapes themselves are still whole.
A Molecular “Integrity Test” for DNA Origami
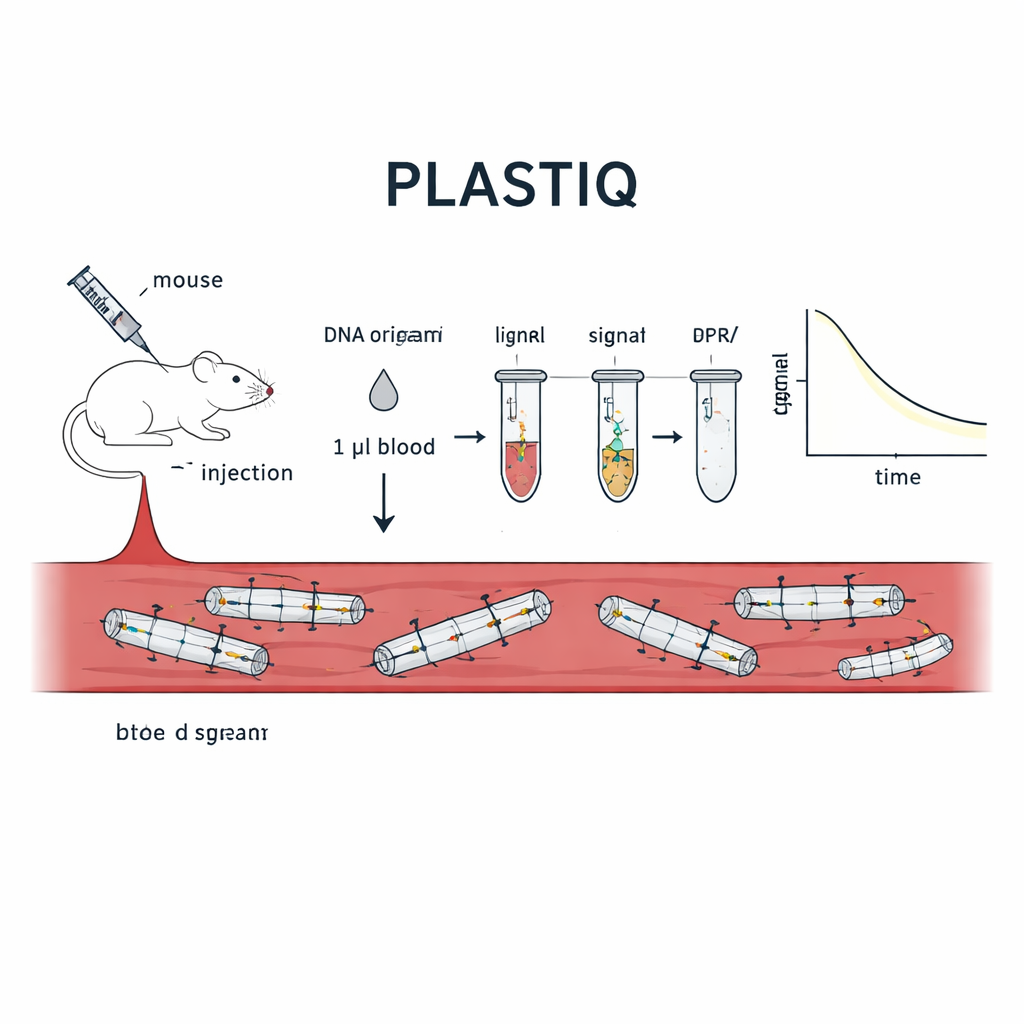
The team developed a method they call PLASTIQ, short for proximity ligation assay for structural tracking and integrity quantification. The key idea is to let the DNA origami report on its own condition. At specific positions in each structure, the researchers place pairs of short DNA staples that sit shoulder to shoulder along a helix when the origami is folded correctly. One of each pair carries a small chemical tag that allows them to be enzymatically joined, or ligated, but only if they are still held close together by an intact scaffold strand. If the structure melts, breaks or is chewed up by enzymes, those two pieces drift apart and can no longer be ligated. After ligation, the joined pairs are amplified using standard polymerase chain reaction and quantified by sensitive qPCR or sequencing, turning the presence of intact local segments into measurable signals from mere microliters of blood.
Putting the Test to Work in Living Mice
To prove that PLASTIQ really measures structural integrity, the authors first applied it to simple rod-shaped DNA origami designs in test tubes. Signals appeared only when the rods were intact and vanished after heat-induced unfolding, while more traditional DNA-based assays still registered DNA regardless of shape. They then injected these rods into mice through different routes—directly into the bloodstream, into the abdominal cavity, into muscle and under the skin—and drew just one microliter of blood at various times. PLASTIQ revealed how the concentration of intact structures spiked and then dropped, depending on the injection route, matching general expectations from drug pharmacology but with much finer structural detail. The method was sensitive down to 0.01 femtomolar, yet required minimal blood, allowing repeated sampling from the same animal over hours.
Testing Protective Coatings and Hidden Weak Spots
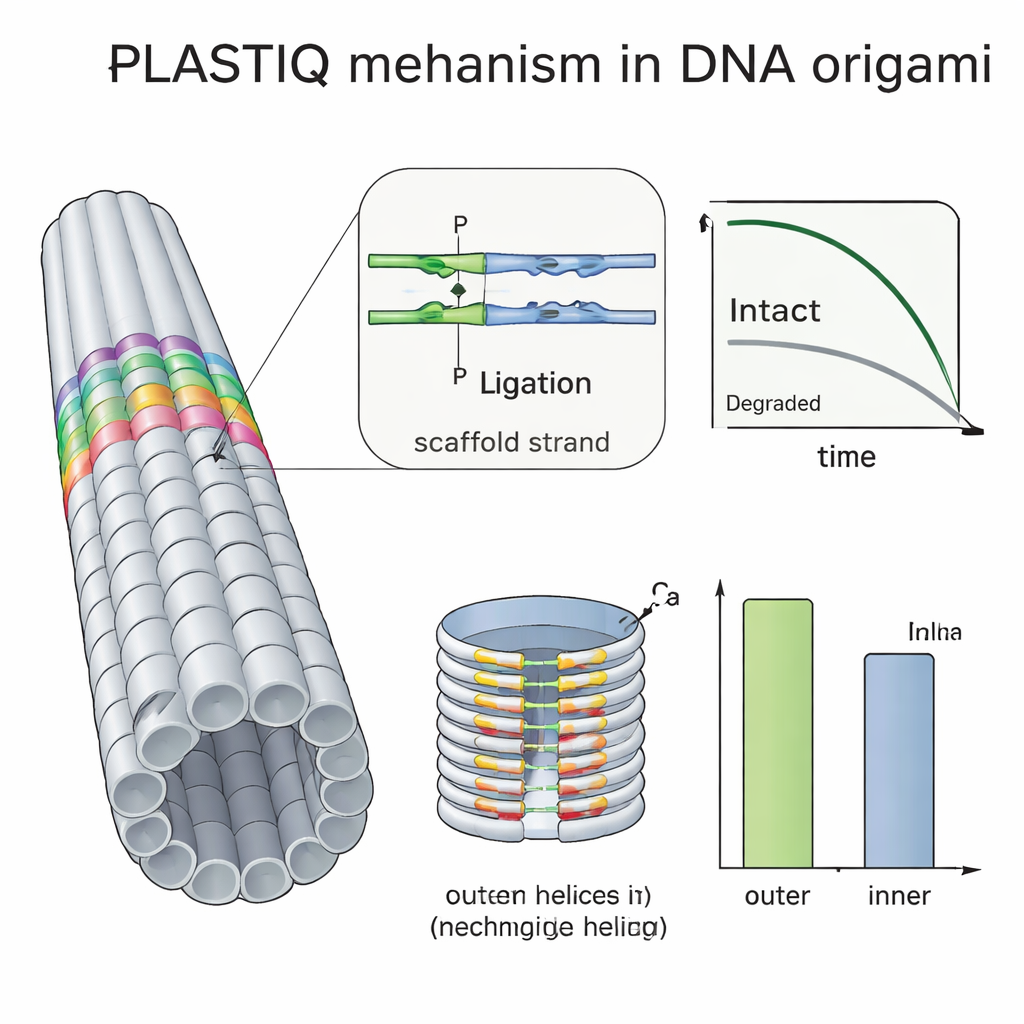
Many groups try to protect DNA origami by coating them with polymers such as PEG, crosslinking strands with ultraviolet light or adding chemical binders, but it has been hard to see how much these tricks actually help in vivo. Using PLASTIQ, the authors showed that PEG-coated rods in the bloodstream stay intact slightly longer than bare ones but are also cleared faster once they reach circulation, and that the coating does not prevent the PLASTIQ enzymes from doing their job. By combining PLASTIQ with simple mathematical models, they extracted absorption and elimination rates for different injection routes and coatings, and could even simulate repeated-dosing regimens. They then pushed the method further with a double-layered barrel-shaped origami, packed more densely on the inside than the outside. PLASTIQ measurements showed that staples on the exposed outer surface lost integrity faster than those facing the inner cavity, directly confirming that buried helices are shielded from degrading enzymes in blood.
What This Means for Future DNA-Based Therapies
To a non-specialist, PLASTIQ can be viewed as a structural checkup for nanoscale DNA devices traveling through the body. Instead of merely asking “is there DNA here?”, it asks “is this DNA still folded into the precise object we designed?”. By answering that question with high sensitivity from tiny blood samples, PLASTIQ provides drug developers with a way to optimize the design, coating and dosing of DNA origami-based medicines and vaccines. Over time, such detailed, quantitative insight into how these nanostructures behave in real biological environments could help turn DNA origami from an elegant laboratory concept into reliable, regulator-ready therapeutics.
Citation: Wang, Y., Rocamonde-Lago, I., Waldvogel, J. et al. Resolving DNA origami structural integrity and pharmacokinetics in vivo. Nat. Nanotechnol. 21, 268–276 (2026). https://doi.org/10.1038/s41565-025-02091-z
Keywords: DNA origami, nanomedicine, drug delivery, pharmacokinetics, nanotechnology