Clear Sky Science · en
Monovalent pseudo-natural products supercharge degradation of IDO1 by its native E3 KLHDC3
Turning Off a Cancer-Friendly Enzyme
Our immune system constantly patrols for cancer cells, but tumors often fight back by hijacking the body’s own chemistry. One key player in this tug-of-war is an enzyme called IDO1, which helps tumors disarm immune cells. This study describes a new class of small molecules that does more than merely block IDO1’s activity—it persuades cells to get rid of the enzyme altogether. By harnessing the cell’s natural trash-disposal machinery, these compounds may revive immune attacks on tumors in ways that older drugs could not.
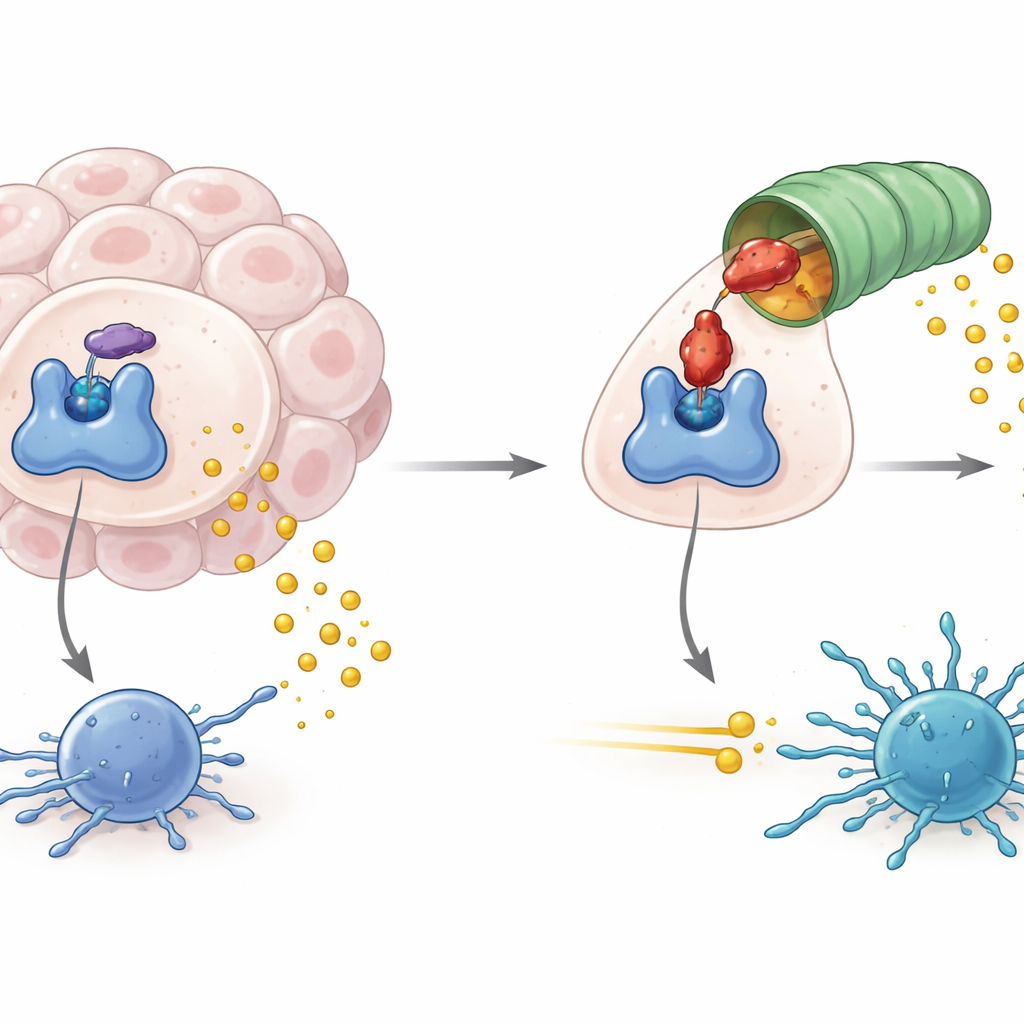
How Tumors Exploit a Metabolic Gatekeeper
IDO1 sits at the entry point of a metabolic pathway that breaks down the amino acid tryptophan into a product called kynurenine. When IDO1 is abundant and active, tryptophan levels fall and kynurenine rises, creating a chemical environment that dampens the activity of immune cells, including tumor-fighting T cells. High IDO1 activity has been linked not only to weaker anti-tumor immunity but also to virus-driven lymphomas and some forms of neurodegeneration. Drug developers have therefore focused on blocking IDO1’s enzyme activity, hoping to cut off this immunosuppressive signal and help existing cancer immunotherapies work better.
Why Simple Enzyme Blockers Have Fallen Short
Despite strong results in animal studies, clinical trials of traditional IDO1 inhibitors have been disappointing. One emerging explanation is that IDO1 is more than a simple catalyst. Even when its chemistry is blocked, the protein itself can act as a signaling hub that still promotes tumor growth and spread. Some inhibitors even stabilize IDO1, increasing how much of the protein is present in cells. In other words, shutting down the catalytic function alone may leave its pro-tumor, non-enzymatic roles untouched—or worse, may actually strengthen them.
Designing Compounds That Trigger Self-Destruction
The authors searched a large library of “pseudo-natural” molecules—synthetic compounds built from fragments inspired by natural products but recombined in new ways. They found a family of molecules, dubbed iDegs, derived from a plant-like building block called (−)-myrtanol. These compounds bind inside IDO1’s usual pocket for its iron-containing cofactor, haem. By displacing haem, iDegs both weaken IDO1’s ability to make kynurenine and, crucially, reshape part of the protein’s tail region. High-resolution crystal structures reveal that when iDegs occupy the haem site, they loosen and partially disorder a C‑terminal helix that is normally held in a compact, protected conformation.
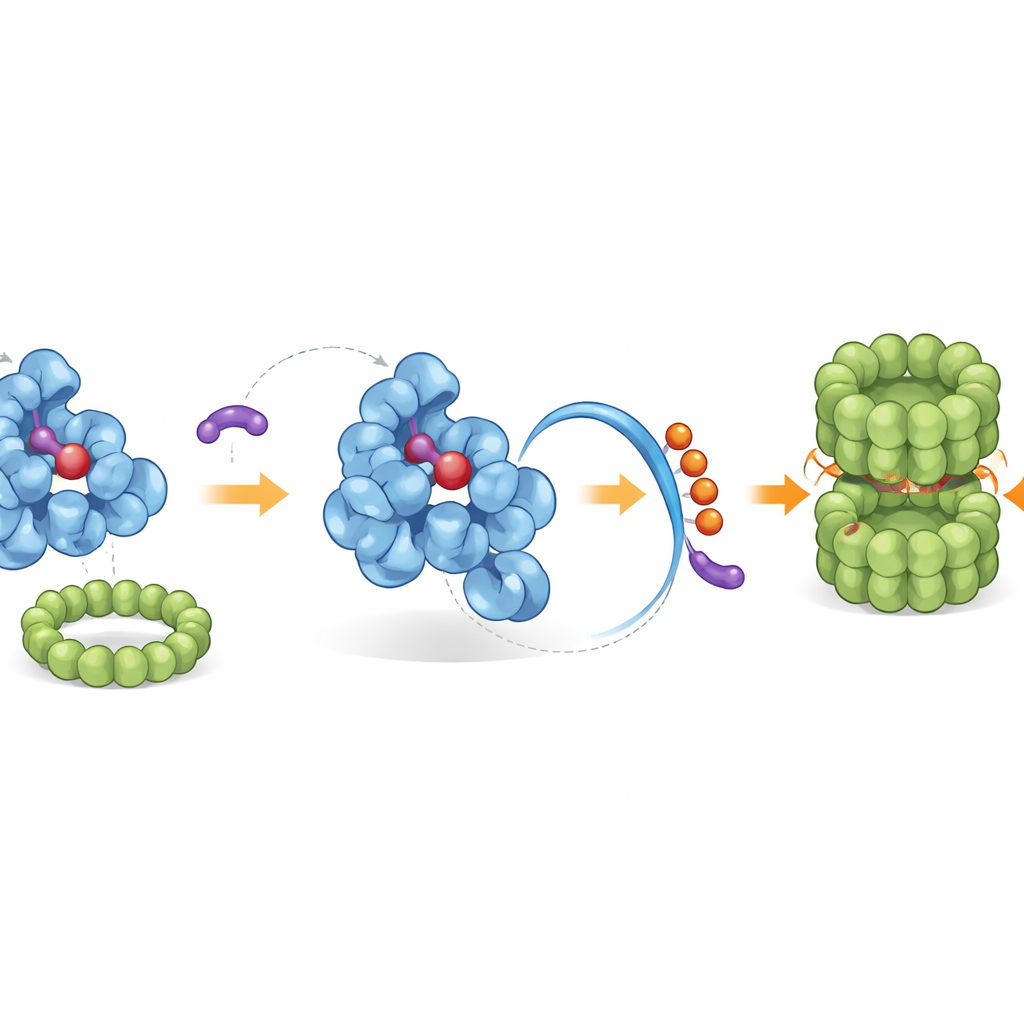
Recruiting the Cell’s Native Cleanup Crew
The reconfigured tail exposes a built-in “end tag” that the cell’s waste-disposal machinery recognizes. Using genetic screens and biochemical reconstitution, the researchers identified a specific ligase complex, called CRL2–KLHDC3, that naturally attaches small ubiquitin “flags” to IDO1’s exposed tail, marking it for destruction by the proteasome—the cell’s molecular shredder. Under normal conditions, only the haem-free form of IDO1 is efficiently tagged this way, while haem-bound IDO1 is protected. iDegs tip this balance by binding only the haem-free form, amplifying the natural degradation route. Potent members of the series not only block kynurenine production but also drive up ubiquitin tagging on IDO1 and sharply reduce its protein levels across several cancer cell types.
Shifting the Fate of IDO1—and Immune Responses
This dual-action behavior has important functional consequences. In migration assays with ovarian cancer cells, a clinically explored IDO1 inhibitor that stabilizes the enzyme actually sped up cell movement, in line with IDO1’s non-enzymatic, pro-tumor role. By contrast, an iDeg compound slowed migration, consistent with both enzymatic inhibition and loss of the protein itself. The study therefore uncovers a new way to manipulate IDO1: not by forcibly dragging it to an unrelated ligase with a bulky designer molecule, but by “supercharging” a native degradation pathway that already exists in cells.
What This Could Mean for Future Cancer Therapies
For non-specialists, the key message is that the quality—not just the quantity—of enzyme inhibition matters. These new small molecules act like precision switches: they lock IDO1 into a vulnerable shape that the cell is primed to destroy, cutting off both its metabolic and signaling contributions to tumor survival. Because iDegs are compact, drug-like, and rely on the cell’s own ligase for removal of IDO1, they may offer a more subtle and broadly applicable approach than earlier “protein-degrader” drugs. If similar strategies can be applied to other problematic proteins, this work hints at a new generation of therapies that coax diseased cells to dismantle their own molecular weapons.
Citation: Hennes, E., Lucas, B., Scholes, N.S. et al. Monovalent pseudo-natural products supercharge degradation of IDO1 by its native E3 KLHDC3. Nat. Chem. 18, 585–596 (2026). https://doi.org/10.1038/s41557-025-02021-5
Keywords: IDO1, cancer immunotherapy, targeted protein degradation, ubiquitin ligase, small-molecule degraders