Clear Sky Science · en
Biocompatible ligand balancing in transition metal coordination enables benign in-cell protein arylation
Turning Metals into Gentle Cell Tools
Many powerful chemical reactions rely on metals, but bring those same metals near living cells and you usually get trouble: damage, stress, and cell death. This study shows that with the right molecular "handle" around a nickel atom, it is possible to run a sophisticated reaction inside living cells without harming them. That breakthrough lets scientists mark thousands of specific spots on proteins and even follow the appearance of hard‑to‑track pathogens, opening new ways to map what is really happening inside cells in health and disease.
Why Metals Are Both Friends and Foes
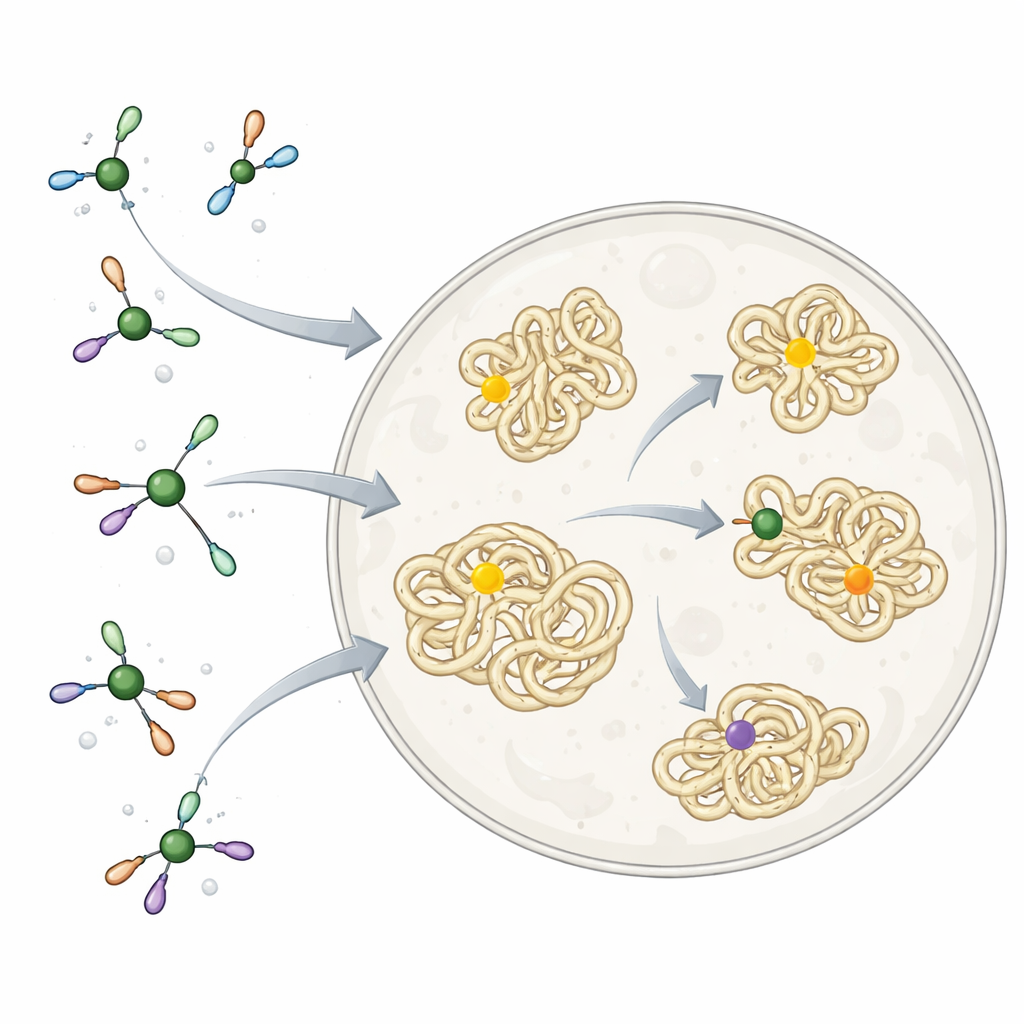
Nickel and other metals already work quietly inside our bodies as parts of natural enzymes, but they can also be toxic if they bind in the wrong place. Nature solves this by surrounding metals with carefully chosen small molecules and proteins, which steer them to the right targets and block unwanted reactions. Chemists, by contrast, often use metal reagents that are extremely reactive and not at all tuned for life. These have been excellent tools for building complex molecules in a flask, but far too harsh to use freely inside cells, especially when the goal is to join a small "tag" onto a specific amino acid in a protein without disturbing the rest of the cell.
Designing a Kinder Nickel Reagent
The researchers drew inspiration from how cells themselves handle nickel. They built a set of nickel complexes wrapped in a simple, biocompatible ligand called TMEDA. This small molecule acts like a soft clamp: tight enough to keep nickel from sticking to the wrong cellular components, but loose enough to let it perform a key reaction. The reaction joins an "aryl" fragment—a flat, ring‑shaped group commonly found in drugs—to the sulfur atom of the amino acid cysteine in proteins. On purified proteins in solution, these nickel complexes very rapidly and selectively attached aryl groups to single cysteine sites, and they worked on many different protein shapes and positions, showing that the chemistry was broadly compatible with real biological molecules.
Editing Proteins Inside Living Cells
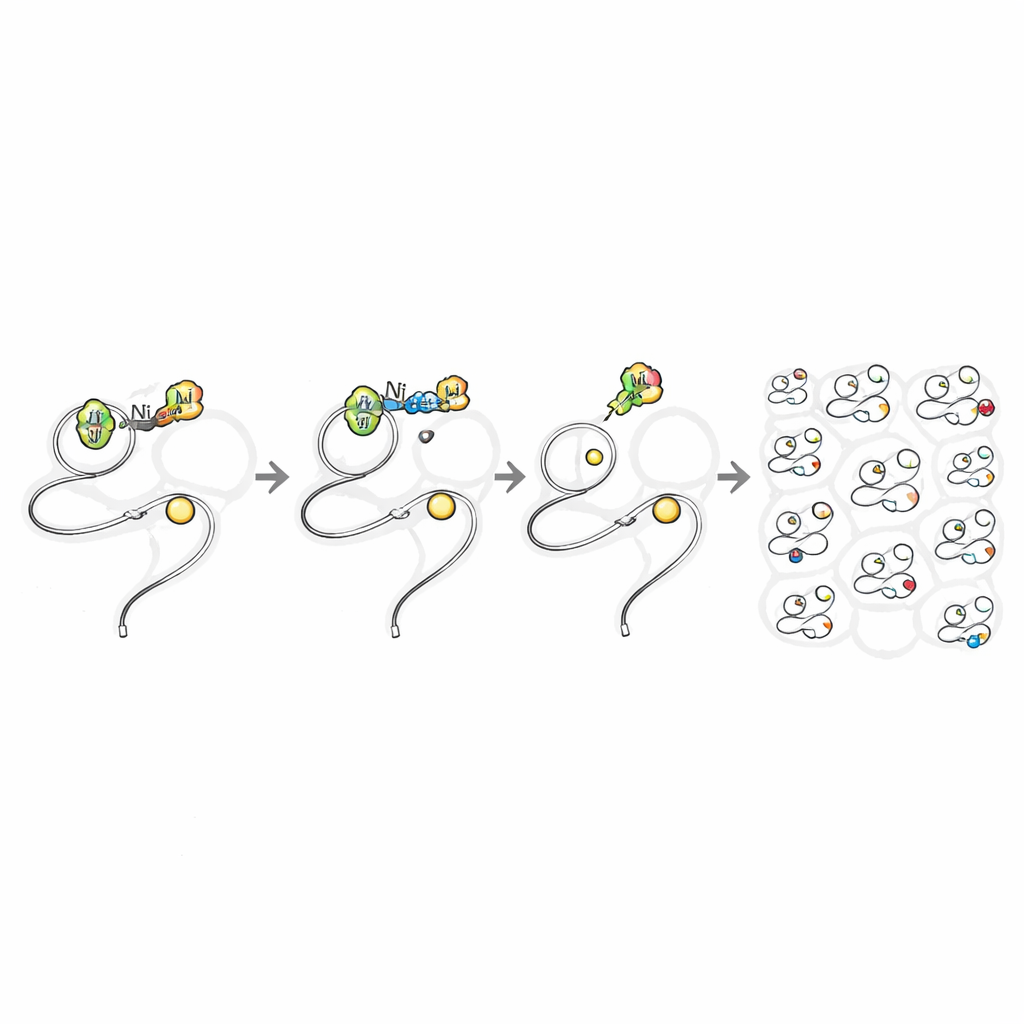
Next, the team asked whether these same reagents could work inside living cells without being poisonous. They compared simple nickel salts, which are known to be harmful, with the TMEDA‑bound nickel complexes. In mammalian cells, the plain nickel sources caused substantial cell death at relatively low doses, but the ligand‑balanced complexes remained well tolerated even at millimolar concentrations. That safety window allowed the researchers to bathe bacterial and mammalian cells in the nickel reagents long enough for them to slip inside and modify proteins. By building an azide "handle" into one version of the aryl group, they could click on fluorescent dyes or biotin tags after the reaction, revealing clear, dose‑dependent staining of proteins throughout the cytoplasm and nucleus of living cells.
Mapping Reactive Protein Sites Across the Proteome
With a safe and fast in‑cell reaction in hand, the authors turned it into a discovery tool. They treated live human cells with the azide‑bearing nickel reagent, then used a photocleavable biotin tag and advanced mass spectrometry to see exactly which cysteines had been modified. In a single experiment they detected nearly 11,000 cysteine sites across almost 5,000 proteins—about twice as many proteins as all previous live‑cell cysteine‑profiling studies combined. The labeling was highly selective for cysteine and showed little bias for particular protein types, locations, or known active sites. Remarkably, many of the targeted proteins were considered "non‑ligandable" by current drug‑discovery standards, including low‑abundance signaling proteins and redox‑sensitive switches that are difficult to study by genetics alone.
Following Hidden Pathogens in Real Time
The same chemistry also proved sensitive enough to catch foreign proteins made during infection. In human cells carrying latent viral sequences, the method detected viral transcription factors present at extremely low levels, including alternative splice forms. The team then infected cells with two very different pathogens: the intracellular bacterium Chlamydia trachomatis and Sindbis virus, an RNA virus related to chikungunya. By pulsing infected cells with the nickel reagent at different times, they were able to trap cysteine sites on key bacterial ribosomal and regulatory proteins as the bacterium shifted between life‑cycle stages, and on critical viral non‑structural proteins that drive RNA replication. These marked sites now stand out as potential weak points for future antiviral or antibacterial strategies.
What This Means for Future Cellular Chemistry
By carefully balancing the ligand shell around nickel, this work shows that a traditionally risky metal can carry out a precise, covalent protein‑editing reaction deep inside living cells with minimal harm. That makes it possible to paint a detailed, functional map of reactive cysteine sites across the proteome, including proteins that are scarce, transient, or hard to drug. It also offers a way to track and probe pathogens within their host cells at the level of individual amino acids. More broadly, the study suggests that many other "forbidden" metal chemistries might be tamed in similar fashion, opening a new era in which the powerful tools of synthetic chemistry operate safely inside living systems.
Citation: Fu, X., Liu, W., Demyanenko, Y. et al. Biocompatible ligand balancing in transition metal coordination enables benign in-cell protein arylation. Nat. Chem. 18, 457–472 (2026). https://doi.org/10.1038/s41557-025-02017-1
Keywords: cysteine profiling, nickel bioconjugation, live-cell protein labeling, chemoproteomics, pathogen proteome mapping