Clear Sky Science · en
Time-resolved functional genomics using deep learning reveals global hierarchical control of autophagy
How Cells Recycle Themselves—and Know When to Stop
Our cells survive lean times by recycling their own parts through a process called autophagy—literally, “self‑eating.” Too little recycling and damaged components pile up; too much and the cell cannibalizes itself. This study asks a deceptively simple question with big implications for aging and disease: how does a living cell turn autophagy on and off at the right moments as nutrients rise and fall?
Following Thousands of Cells Through a Feast‑to‑Famine Cycle
To tackle this, the researchers used baker’s yeast as a model and watched nearly six thousand different mutant strains as they went through starvation and recovery. Each yeast strain carried fluorescent tags marking structures involved in autophagy, allowing a high‑powered microscope to capture images every hour as nutrients were removed and then restored. Instead of manually sorting these images, the team trained deep‑learning algorithms to recognize when a cell was in an autophagic state, based entirely on image features rather than human judgment. This produced precise time‑courses of autophagy activity for almost the entire yeast genome, revealing how quickly and how strongly each mutant responded to nutrient shifts. 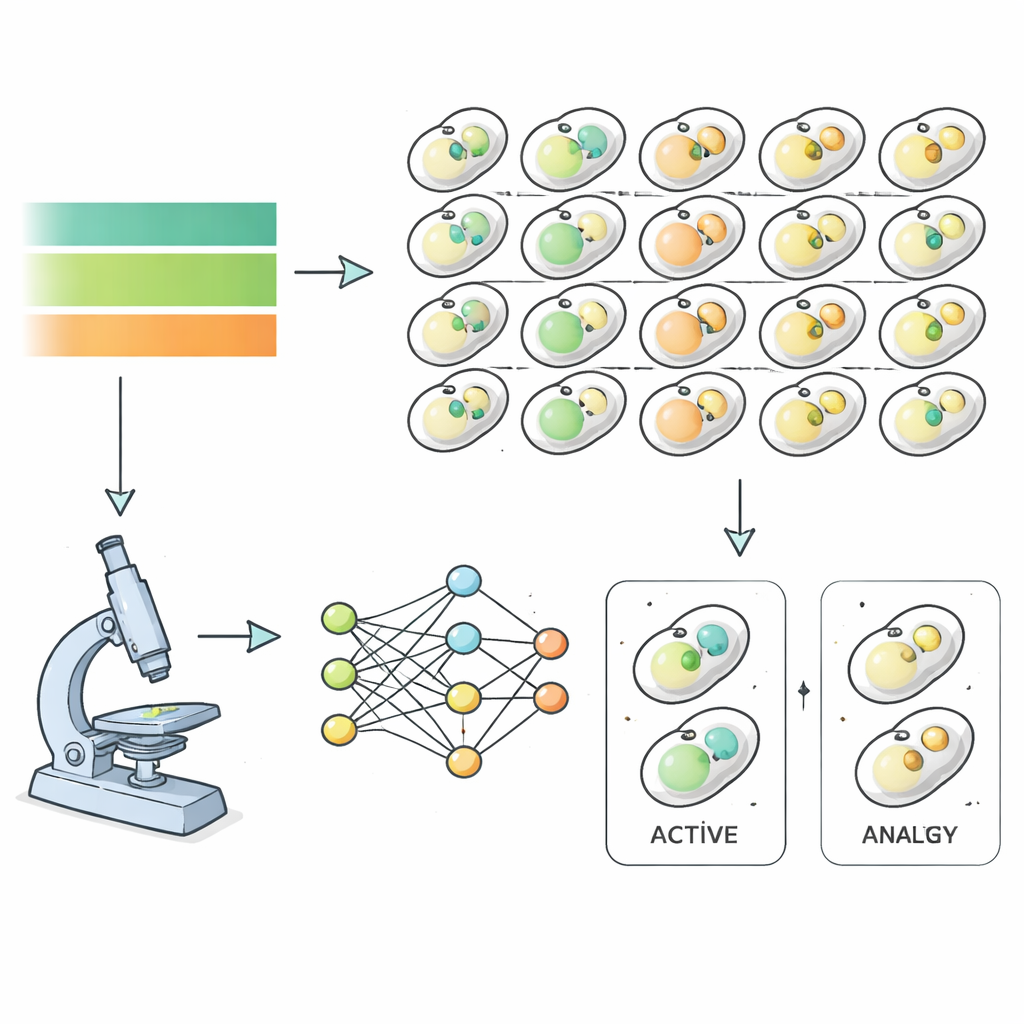
Six Distinct Ways Cells Mis‑Handle Recycling
By mathematically fitting these time‑courses, the authors extracted detailed “kinetic fingerprints” for each mutant—how fast autophagy turned on, how long it stayed high, and how sharply it shut down. Grouping these signatures uncovered six broad behavior types. Some mutants were “ultrasensitive,” responding very quickly to starvation and to nutrient return. Others were “hyposensitive,” reacting sluggishly. A “hyperactive” group showed abnormally high baseline recycling, while several groups had varying degrees of failure to mount a proper response at all. Importantly, known core autophagy genes fell into the most severe classes, validating that the method captures genuine biology rather than measurement noise.
Mapping a Hierarchy of Control Over Autophagy
Autophagy does not happen in isolation; it is wired into the cell’s wider regulatory networks. The team overlaid their dynamic profiles onto large maps of genetic and protein interactions in yeast. Genes whose loss caused strong autophagy defects tended to sit close to the known autophagy “core machinery” in these networks, while subtler mutants were farther away, consistent with more indirect control. By examining which cellular processes were enriched in each behavior class, they found that starvation‑triggered activation is strongly shaped by pathways handling RNA, protein production, and energy metabolism, whereas shutting autophagy down after nutrients return depends more on membrane traffic and fusion steps that clear recycling vesicles.
Zooming In on How Vesicles Form and Clear
Autophagy proceeds through two main stages: building cargo‑filled vesicles and then fusing them with the cell’s recycling compartment for breakdown. To dissect these stages, the researchers mined the “latent space” inside their neural network—the compressed representation it uses to distinguish images. By projecting this internal representation onto a two‑dimensional map, they could see cell populations move from a region with no vesicles, through one with many free vesicles, and finally to a region where vesicles had been cleared. Using statistical comparisons against reference mutants locked at different stages, they quantified for each gene whether it mainly affected vesicle formation, vesicle clearance, or both. This analysis showed that the control of vesicle formation is highly sensitive to nitrogen levels over time, whereas the clearance machinery behaves more steadily and is often the main limiter of overall recycling flux. 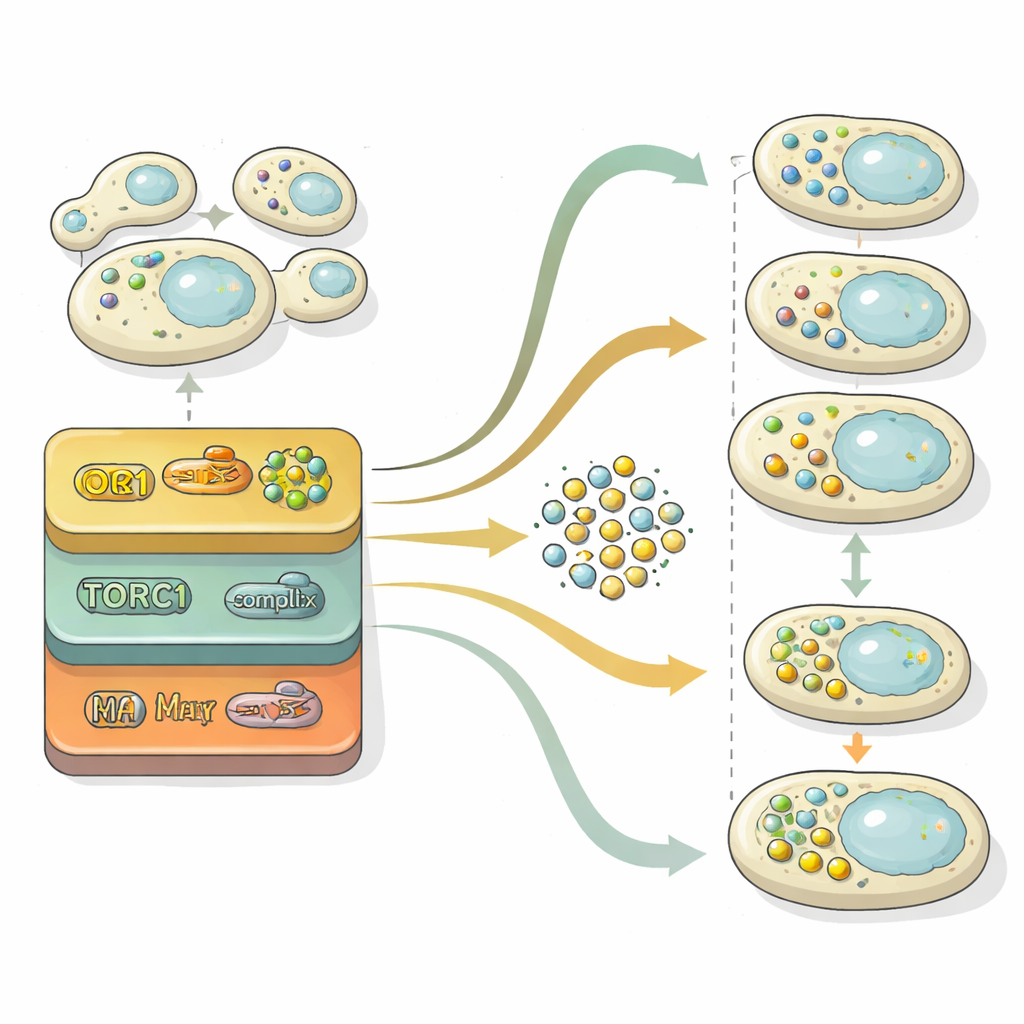
A Hidden Brake on Recycling: The Retrograde Pathway
One of the most striking discoveries concerned the so‑called retrograde (RTG) pathway, which conveys signals from mitochondria to the nucleus. Yeast cells lacking RTG components produced too many autophagosomes even when nutrients were plentiful and struggled to clear them once nutrients returned. Detailed follow‑up experiments showed that this pathway controls expression of key autophagy genes, especially the master initiator ATG1, largely independent of the well‑known TORC1 nutrient sensor targeted by drugs like rapamycin. In effect, RTG signaling acts as a time‑varying brake that prevents runaway autophagy in rich conditions and helps terminate the response when conditions improve.
Why This Matters for Health and Therapy
Altogether, this work delivers a genome‑wide, time‑resolved map of how cells tune autophagy across changing nutrient conditions. It reveals that activation and shutdown are not mirror images: many more genes influence how recycling starts than how it stops, and long‑term balance depends heavily on vesicle clearance and transcriptional brakes such as the RTG pathway. For a general reader, the key takeaway is that cellular self‑cannibalism is under layered, hierarchical control, and that we can now chart these layers with high precision. This kind of systems‑level view could eventually guide more nuanced therapies that adjust autophagy in specific phases—boosting beneficial recycling without triggering the harmful effects of chronic, uncontrolled self‑digestion.
Citation: Chica, N., Andersen, A.N., Orellana-Muñoz, S. et al. Time-resolved functional genomics using deep learning reveals global hierarchical control of autophagy. Nat Cell Biol 28, 465–479 (2026). https://doi.org/10.1038/s41556-025-01837-0
Keywords: autophagy, deep learning, yeast genetics, nutrient sensing, cellular recycling