Clear Sky Science · en
Spatial FBA reveals heterogeneous Warburg niches in renal tumors and lactate consumption in colorectal cancer
Why tumor metabolism in space matters
Cancer cells do not all behave the same, even within a single tumor. Some live close to blood vessels, others deep in poorly fed regions, and these locations shape how they obtain and use fuel. This study introduces a new way to read that hidden “metabolic map” from cutting-edge spatial gene expression data. By doing so, the authors uncover surprising patterns in how different tumors handle a key molecule, lactate, challenging the classic view that cancers only dump lactate out as waste.
Turning gene maps into metabolic activity
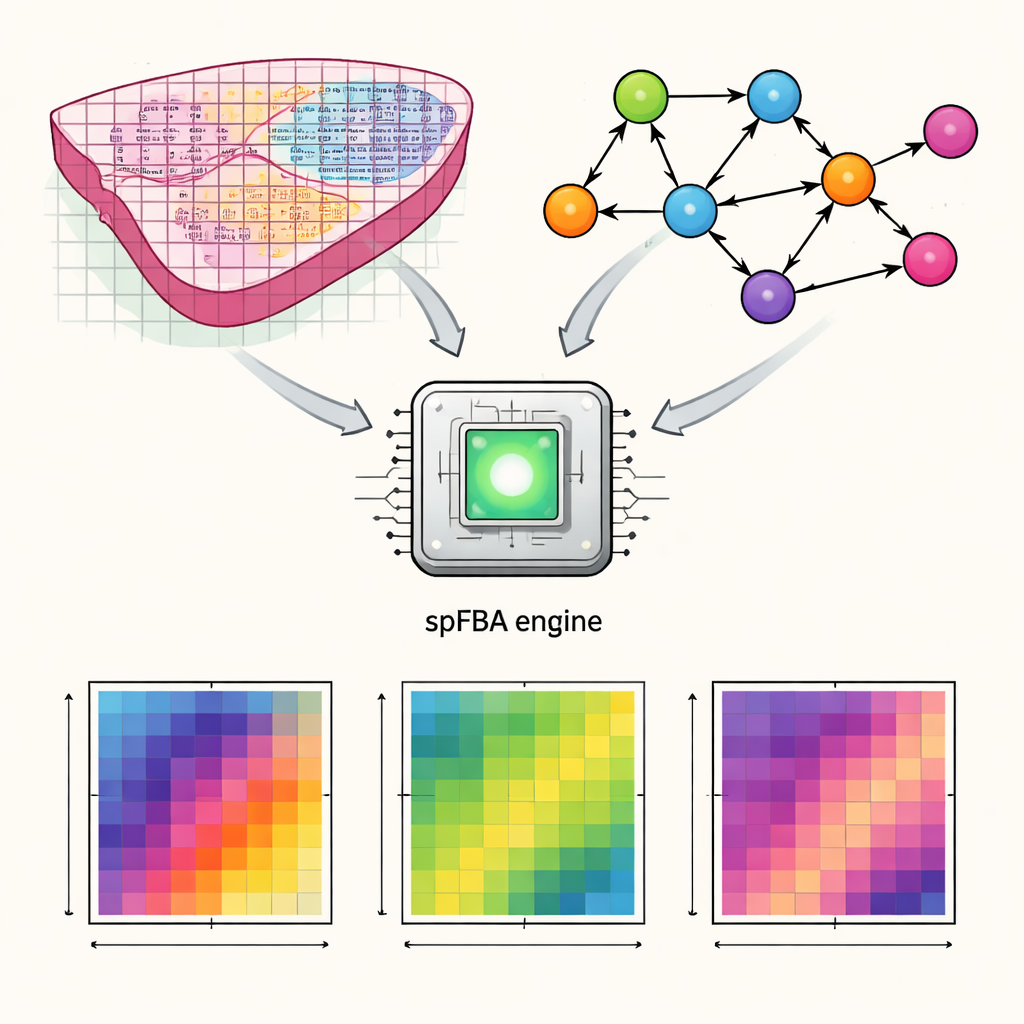
The authors developed a computational framework called spatial Flux Balance Analysis, or spFBA. Spatial transcriptomics technologies measure which genes are active at thousands of tiny points across a thin slice of tissue. spFBA combines these spatial gene activity maps with detailed models of human metabolism, which describe how nutrients like glucose, oxygen, and amino acids are transformed inside cells. Instead of assuming that every region of a tissue shares the same metabolic goal, spFBA treats each spot independently, exploring which reaction patterns are possible there under the laws of chemistry and mass balance. The output is a set of “flux enrichment scores” that indicate, for each location, how strongly different metabolic reactions are likely to flow and in which direction.
Testing the method in kidney tumors
To see if spFBA produces biologically sensible results, the team first turned to clear cell renal cell carcinoma, a kidney cancer known for relying heavily on sugar breakdown (glycolysis) and lactate release, a phenomenon called the Warburg effect. Using published spatial data from ten kidney tumor samples, they asked whether the predicted metabolic patterns matched the tissue’s known structure. They found that spots grouped by their inferred metabolic fluxes lined up well with the histological organization seen under the microscope and with clusters based on gene expression alone. Importantly, spFBA recovered the expected metabolic contrast between tumor and normal kidney tissue: tumor regions showed higher glucose use, stronger biomass production (a proxy for cell growth), and robust lactate secretion. At the same time, oxygen use remained substantial, revealing that different parts of the same tumor can mix fermentation and respiration depending on local blood supply.
Colorectal cancers reveal a different lactate story
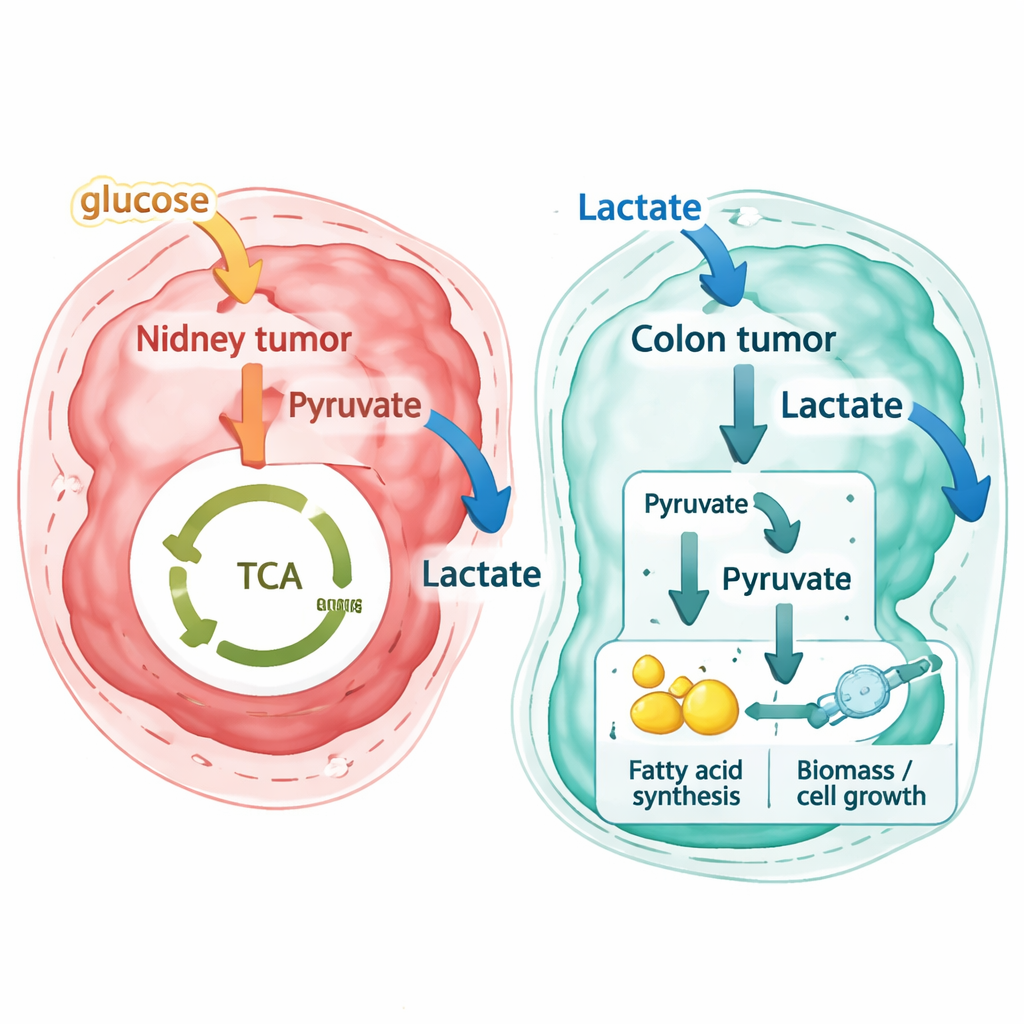
The researchers then applied the same pipeline to new high-resolution spatial datasets from a patient with primary colorectal cancer and matched liver metastases, as well as to an independent public colorectal dataset generated with another technology. Here, the results were strikingly different. Instead of exporting lactate as waste, most colorectal cancer regions—both in the original colon tumor and in liver metastases—were predicted to import lactate from their surroundings. Stromal cells in the nearby supporting tissue tended to release small amounts of lactate, while tumor cells acted as strong consumers. By tracing reaction-level flux patterns, the authors show that these metastatic cells do not simply burn lactate in the usual energy-producing cycle inside mitochondria. Rather, they convert lactate into building blocks that feed into pathways for making fats and other components needed for cell growth, a “pseudo–reverse Warburg effect” in which lactate becomes raw material for biosynthesis.
Metabolic niches and the tumor-stroma frontier
Because spFBA keeps the spatial layout, it can pinpoint where particular metabolic exchanges occur. In kidney tumors, the method revealed coexisting metabolic “neighborhoods”: better-oxygenated, blood-rich interfaces where cancer cells both respire and ferment, and deeper, poorly vascularized cores that rely more strongly on fermentation. In colorectal liver metastases, spFBA highlighted sharp contrasts at the border between tumor and stroma, where glutamate and lactate handling changed direction, suggesting intense exchange at the invasive front. Across all datasets, predicted growth-related fluxes correlated with independent gene-based measures of proliferation, supporting the biological realism of the model. Crucially, the same algorithm predicted lactate secretion in kidney cancer but lactate uptake in colorectal cancer, indicating that the observed differences arise from the data, not from built-in model bias.
What this means for understanding and treating cancer
For non-specialists, the key message is that cancer metabolism is not only abnormal—it is also highly local. The same tumor can host multiple metabolic niches, and similar-looking tumors in different organs may use the same molecules in opposite ways. This work shows that by layering spatial gene expression over detailed metabolic models, researchers can infer where tumors are likely to be hungry for particular nutrients such as lactate. In colorectal cancer, the discovery that tumor cells extensively consume lactate and channel it into growth-related processes raises new questions about how diet, gut microbes, and the liver’s own metabolism shape disease progression, and whether blocking lactate use could make these tumors more vulnerable to therapy.
Citation: Maspero, D., Marteletto, G., Lapi, F. et al. Spatial FBA reveals heterogeneous Warburg niches in renal tumors and lactate consumption in colorectal cancer. npj Syst Biol Appl 12, 32 (2026). https://doi.org/10.1038/s41540-026-00654-x
Keywords: cancer metabolism, spatial transcriptomics, lactate, Warburg effect, colorectal cancer