Clear Sky Science · en
In silico modeling of anterior foregut endoderm differentiation towards lung epithelial progenitors
Turning Stem Cells into Lung-Building Cells
Scientists are learning how to coax a patient’s own stem cells into becoming replacement lung tissue, which could one day repair damage from diseases like COPD, fibrosis, or severe infections. This article explores how researchers used computer modeling to map and fine‑tune a crucial step in that journey: turning an intermediate cell type, called anterior foregut endoderm, into early lung epithelial progenitors—the starter cells that can eventually form the airways and air sacs of the lung.
Why Lung Starter Cells Matter
Human induced pluripotent stem cells (iPSCs) can be reprogrammed from adult tissues and then guided toward many different organs. To build lung tissue, these cells first pass through several developmental waypoints. One of these is the anterior foregut endoderm, a layer that normally gives rise to parts of the respiratory and digestive systems in the embryo. From there, with the right chemical cues, cells can become lung epithelial progenitors, which carry early lung markers and can later mature into specialized airway or alveolar cells. Because future cell therapies will require billions of these cells, researchers need ways to reliably boost yields and adapt protocols to different patient cell lines without endless trial‑and‑error in the lab.
Building a Virtual Version of Cell Differentiation
The team expanded an earlier mathematical framework to build what is, to their knowledge, the first population‑level model of this specific transition from foregut endoderm to lung progenitors. They considered two ways of representing the cells: a simple version that tracks only the total number of living cells, and a more detailed version that follows the anterior foregut cells and the lung progenitors separately. In both cases, the model also follows glucose and lactate in the culture medium, representing nutrients and waste. Using tools from systems biology, the researchers constructed many candidate equations for how cells grow, die, and differentiate, then used identifiability tests to discard models whose parameters could never be pinned down, even with perfect data.
Designing Smarter Experiments with the Model
Rather than just fitting whatever data happened to be available, the researchers let the model guide how new experiments should be run. They used simulated data to ask how often they would need to measure cell numbers and nutrient levels in order to accurately estimate model parameters, balancing statistical precision against the cost and labor of frequent sampling. This led them to a practical plan: daily measurements for glucose and lactate, and cell counts every one to two days, under four conditions that varied how hard the cultures were split on day 10 and whether the growth medium was refreshed daily. They then carried out these experiments, measuring both the total cell population and, via flow cytometry, the fractions that remained foregut cells or had become lung progenitors.
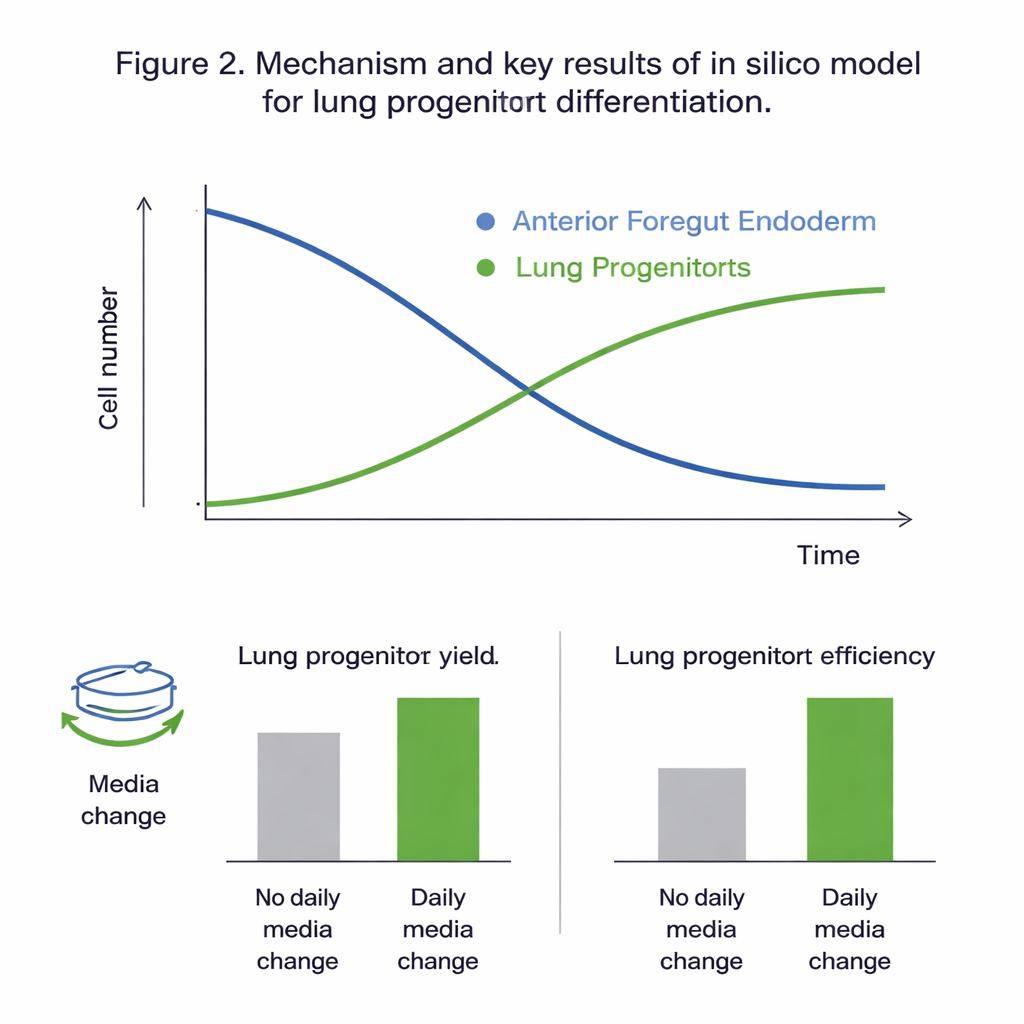
What the Virtual Experiments Revealed
Fitting all candidate models to the experimental data showed that the two‑population model, which tracks foregut and lung progenitor cells separately, could be calibrated reliably and captured the observed behavior best. Statistical checks indicated that, over the time window studied (days 11 to 15 of the protocol), the dynamics were driven mainly by proliferation and differentiation of the foregut cells, while proliferation of the lung progenitors themselves contributed little. Global sensitivity analysis reinforced this picture, highlighting foregut cell growth, death, and differentiation rates—and the influence of glucose—as the main levers controlling the outcome. The calibrated model reproduced unseen data with errors comparable to the natural variability of the experiments, suggesting it was accurate enough to explore “what‑if” scenarios in silico.
Optimizing Media Changes and Cell Splitting
With a trustworthy virtual system in hand, the team tested how two practical protocol choices affect results: how much the culture is diluted (the split ratio) on day 10, and whether the growth medium is replaced every day. The simulations predicted that daily media changes nearly double the number of lung progenitor cells and the yield per starting cell, mainly by preventing nutrient depletion and buildup of waste and unstable signaling molecules. Experiments agreed closely with these predictions. The model also suggested that using higher split ratios—spreading cells more thinly on day 10—improves the “yield per input cell” by around a quarter, even though it reduces the absolute number of cells. In both cases, these changes had little effect on the final proportion of lung progenitors within the culture, primarily shifting how many cells could be produced efficiently.
What This Means for Future Lung Therapies
For a non‑specialist, the key message is that the authors have built a kind of flight simulator for a critical step in growing lung cells from stem cells. By combining carefully designed experiments with rigorous mathematical modeling, they show how simple protocol choices—like how often to change the culture medium and how densely to seed cells—can dramatically influence how many lung‑building cells are produced, without changing their quality. This type of in silico modeling can help streamline future protocols, reduce experimental guesswork, and ultimately support more reliable and scalable production of lung progenitors for research, disease modeling, and, eventually, regenerative therapies.
Citation: Mostofinejad, A., Romero, D.A., Brinson, D. et al. In silico modeling of anterior foregut endoderm differentiation towards lung epithelial progenitors. npj Syst Biol Appl 12, 29 (2026). https://doi.org/10.1038/s41540-026-00650-1
Keywords: lung progenitor cells, induced pluripotent stem cells, in silico modeling, cell differentiation, regenerative medicine