Clear Sky Science · en
Association of copy number alterations with the immune transcriptomic landscape in cancer
Why our genes matter for cancer immunotherapy
Cancer immunotherapy works by helping the immune system recognize and attack tumors, but only a minority of patients see lasting benefit. This study asks a basic question with big clinical consequences: how does the genetic chaos inside tumor cells – in particular, large gains and losses of DNA called copy number alterations – shape the immune reaction against cancer, and can we read those effects from gene activity patterns?
Taking a panoramic view of cancer data
To tackle this, the researchers assembled an enormous collection of 294,159 gene activity profiles from tumors and other tissues. These profiles, taken from several major public databases, capture which genes are turned on or off in thousands of samples spanning many cancer types and experimental conditions. Instead of looking at individual genes one by one, the team used a mathematical method to break each profile into underlying "components" – recurring patterns of genes that tend to rise and fall together, each reflecting a biological process such as an immune response or the effect of a DNA change.
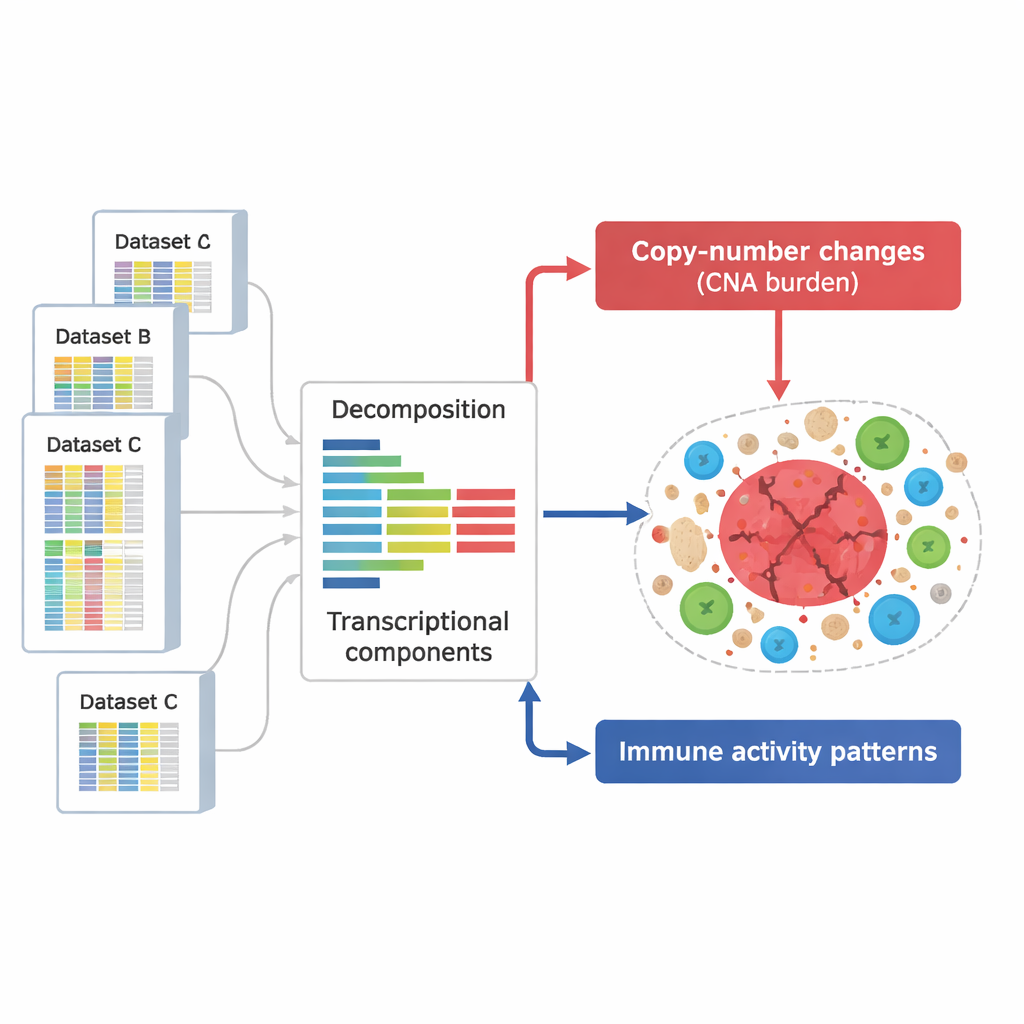
Separating DNA damage signals from immune signals
From these components, the scientists defined two key groups. One group captured the effects of copy number alterations – stretches of chromosomes that are repeatedly gained or lost in cancer cells. These patterns covered almost the entire genome, indicating that most regions affected by such alterations left a detectable imprint on gene activity. A second group of components was enriched for genes involved in immune functions, such as T cell activation, natural killer cell activity, and antigen presentation. Altogether, they identified 657 DNA-related components and 283 immune-related components, many of which could be reproducibly seen across independent datasets and technologies, suggesting that they represent robust, general features of tumor biology.
Linking patterns to treatment response
The team then asked whether these immune-related patterns could help predict who responds to immune checkpoint inhibitors, a major class of cancer immunotherapy drugs. Using data from 13 clinical studies covering 1,167 patients with seven types of cancer, they trained computer models to distinguish responders from non-responders based solely on the activity of the immune components in pre-treatment tumor samples. Some models performed strongly even when tested on completely independent patient groups; for example, a model trained on one breast cancer cohort accurately predicted responses in a separate breast cancer cohort and showed useful performance in several other cancers. A small set of immune patterns, including those linked to interferon responses, natural killer cells, and T cell activation, contributed most strongly to these predictions.
How genetic chaos reshapes tumor immunity
With this framework in place, the researchers systematically related overall copy number alteration burden – a measure of how extensively a tumor’s DNA is gained or lost – to the activity of each immune pattern across many cancer types. Most immune patterns showed an inverse relationship: tumors with high DNA alteration burden tended to have lower activity of components associated with beneficial immune functions, such as antigen presentation and infiltration by key immune cells. However, a notable minority of patterns moved in the opposite direction. Tumors with high alteration burden often had increased signals from immunosuppressive cell types, including regulatory T cells and certain macrophages, as well as inflammatory cells that can promote tumor growth rather than tumor destruction. Spatial analyses of tumor slices confirmed that regions with heavy DNA changes frequently coincided with low activity of helpful immune patterns and with “immune-excluded” zones where immune cells were confined to the tumor margins.
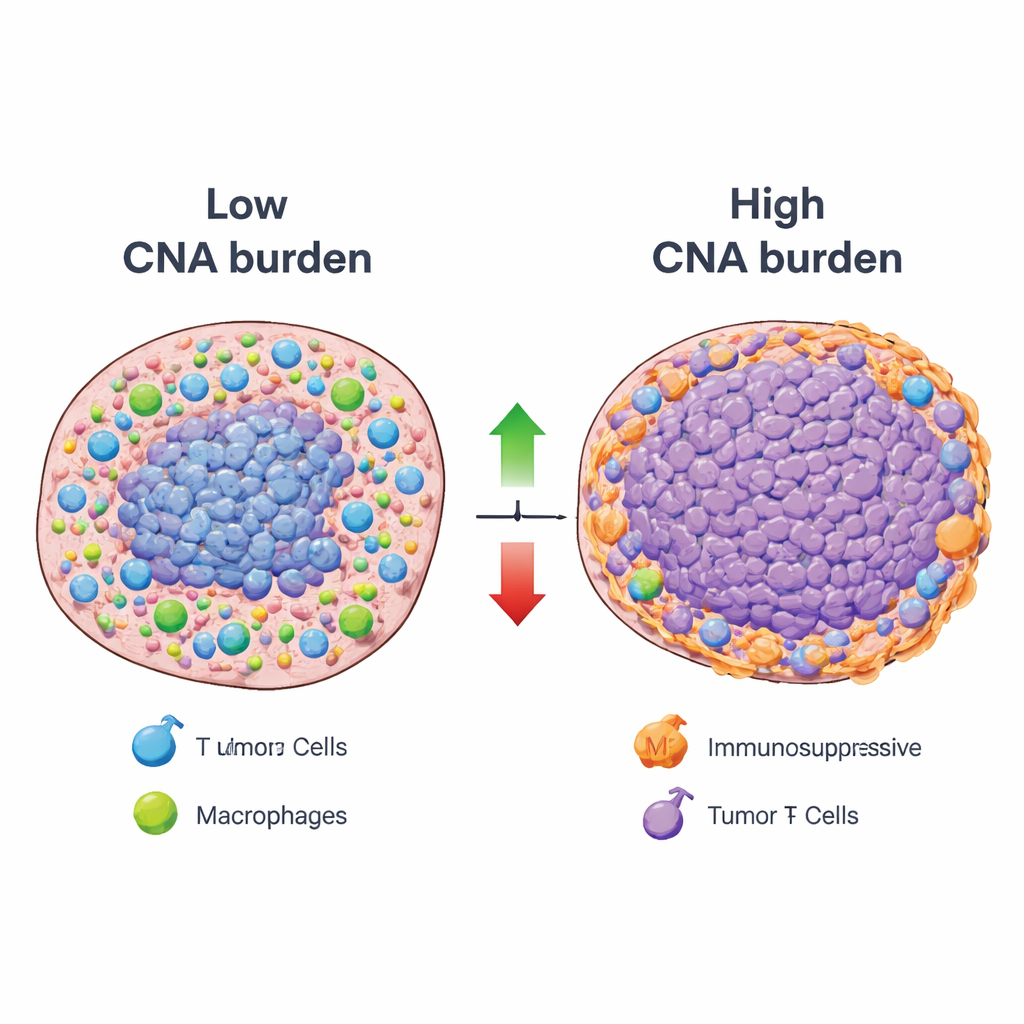
What this means for future cancer treatment
In plain terms, the study shows that tumors burdened by large-scale DNA gains and losses tend to both blunt helpful immune responses and foster suppressive or tumor-promoting immune environments. Yet, they are not immunologically silent; instead, they display specific, recurring immune states that may be vulnerable to targeted therapies, such as drugs blocking IL-17 or IL-23 signaling or strategies that reprogram certain macrophages. By mapping these DNA–immune relationships across cancers and making the resource publicly available, the work offers a detailed guide to why some genetically unstable tumors resist current immunotherapies and suggests new combination treatment strategies to help the immune system overcome that resistance.
Citation: Loipfinger, S., Bhattacharya, A., Urzúa-Traslaviña, C.G. et al. Association of copy number alterations with the immune transcriptomic landscape in cancer. npj Syst Biol Appl 12, 28 (2026). https://doi.org/10.1038/s41540-026-00649-8
Keywords: cancer immunotherapy, copy number alterations, tumor microenvironment, immune checkpoint inhibitors, transcriptomics