Clear Sky Science · en
Simulation-based inference of cell migration dynamics in complex spatial environments
How immune cells find their way through crowded tissue
Our immune cells often have to squeeze through dense, maze‑like tissues to reach infection sites or lymph nodes. This study asks a deceptively simple question with big implications: how do these cells navigate such cluttered environments, and how can we reliably infer their behaviors from messy microscope data? By combining precisely engineered lab “mazes” with advanced computer simulations and modern machine‑learning tools, the authors show a new way to decode the rules that guide cell movement in complex surroundings.
Building a tiny maze for immune cells
To probe how environment shapes movement, the researchers focused on dendritic cells—immune sentinels that must travel from peripheral tissues to lymph nodes, guided by chemical attractants called chemokines. They built a microfabricated chip: a flat “pillar forest” of regularly spaced posts made from silicone rubber (PDMS), with narrow 10‑micrometer gaps that mimic the tight spaces in real tissues. One side of the chip was loaded with tens of thousands of dendritic cells; the opposite side contained a source of the chemokine CCL19, which creates a stable gradient across the pillars. Using time‑lapse microscopy, they tracked the nuclei of individual cells every 30 seconds as they tried to move toward the chemokine source. 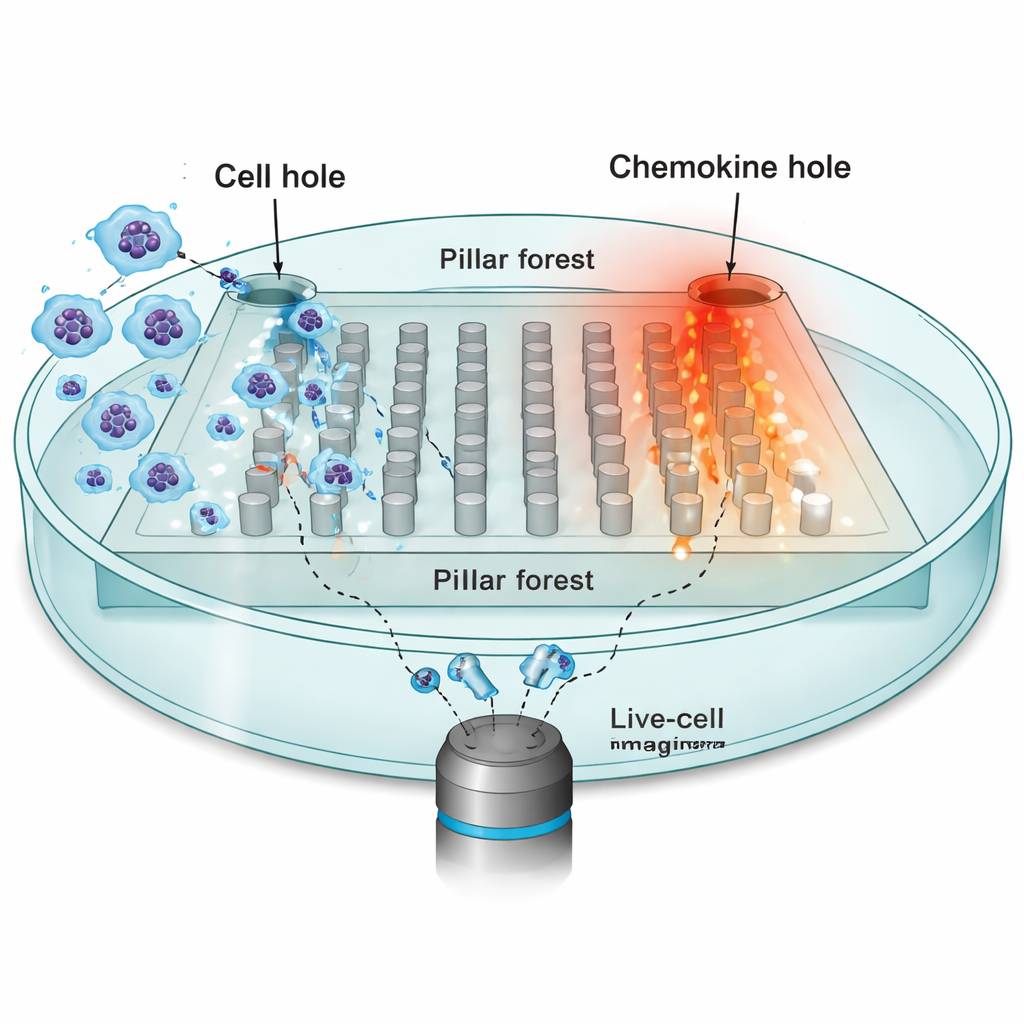
Turning biology into a virtual experiment
To make sense of this diversity, the team built a detailed computational model of cell migration using a framework known as the Cellular Potts model. Instead of treating a cell as a simple point, this approach represents each cell as an extended patch on a grid, allowing it to change shape, squeeze between pillars, and respond to chemical signals. The model includes four key ingredients: how strongly a cell is pulled by the chemokine gradient (mdir), how much it tends to persist in its current direction of motion (mrand), how often it reorients (captured by a rate λ), and its effective size (a). By adjusting these parameters and running simulations, the model produces synthetic tracks that can be compared directly to the recorded cell paths in the pillar forest.
Why hand‑picked measurements fall short
Traditionally, researchers summarize such movement data with a handful of familiar statistics—how far a cell travels (displacement), how fast it moves (velocity), and how its direction changes over time (turning angles). The authors first used these hand‑crafted measures within a technique called approximate Bayesian computation (ABC), which searches for parameter sets that make simulated tracks resemble the experimental ones. They found that while these summaries capture broad trends, they miss much of the fine‑grained structure in the data. As a result, some model parameters, especially those governing random persistence and reorientation timing, remained poorly constrained or even biased. Moreover, ABC required hundreds of thousands of simulations and many hours of computing time to reach acceptable fits.
Letting neural networks learn what matters
To overcome these limits, the study turned to a newer family of methods called neural posterior estimation (NPE). Here, a neural network is trained directly on many pairs of simulated data and underlying parameters. One part of the network automatically learns its own compact set of “summary features” from entire collections of cell tracks; another part learns how those features map back to likely parameter values. Crucially, these learned features are optimized explicitly for accurate parameter inference, not for human interpretability. The authors then re‑used the learned summaries inside an ABC framework, creating a hybrid pipeline that combines the robustness of ABC with the flexibility of neural networks. 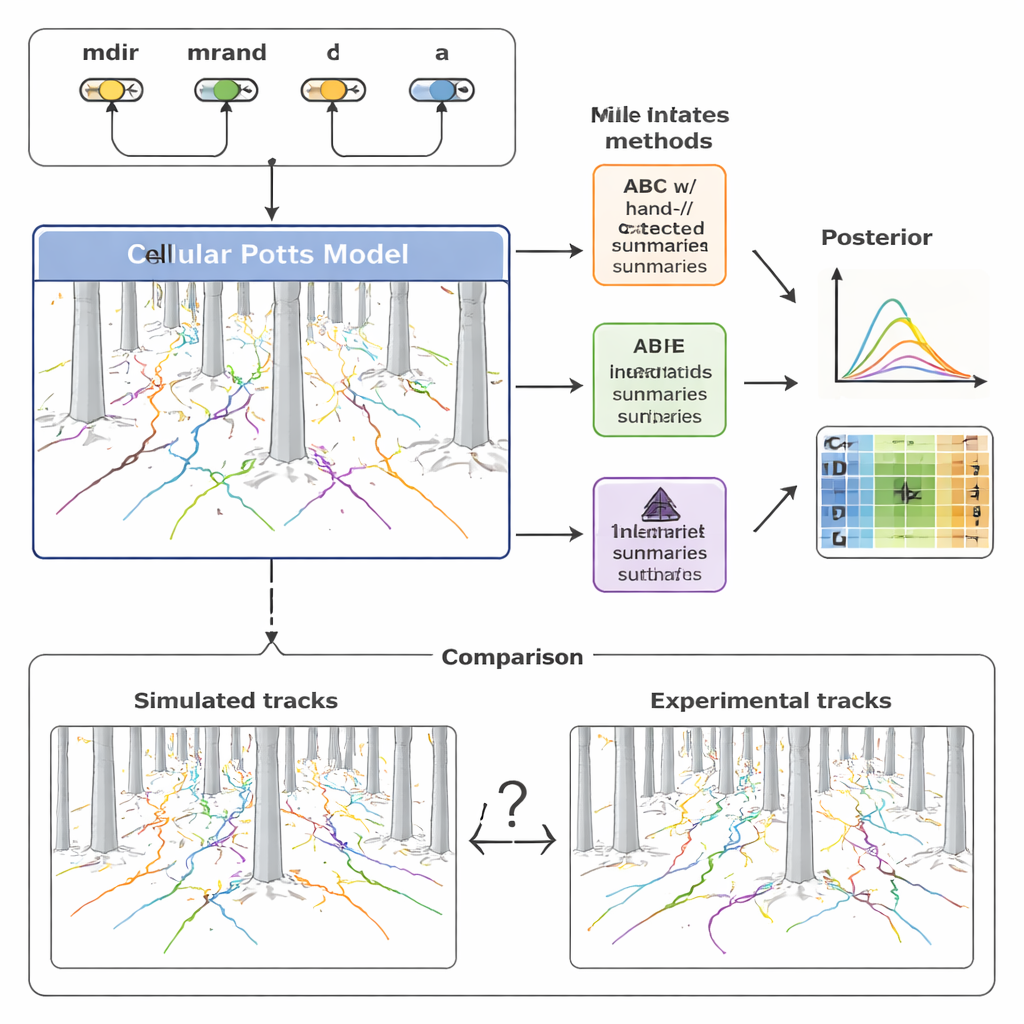
What the new approach reveals about cell navigation
Armed with this calibrated model, the researchers explored how chemokine signals and physical obstacles jointly shape migration. The inferred cell size suggested that dendritic cells effectively shrink and deform to fit through the 10‑micrometer gaps, consistent with their known flexibility. Simulations indicated that persistent random motion is a major driver of how widely cells spread, even without chemokine guidance, and that the pillar forest can trap cells when strong directional cues and persistence work together. Surprisingly, the model predicts that a chemokine signal that is briefly active early on can, in some cases, help more cells reach the target than a constant signal, because prolonged attraction may keep cells circling within obstacles rather than escaping them.
Why this matters for biology and modeling
For non‑specialists, the key message is twofold. First, immune cell migration in tissues is not just about following a chemical trail; it emerges from a subtle interplay between guidance cues, cells’ own movement tendencies, and the physical layout of their surroundings. Second, extracting these rules from imaging data requires letting the computer learn which patterns in the data are most informative, rather than relying solely on simple, human‑designed measurements. By integrating microengineered experiments, shape‑resolving simulations, and neural network–based inference, this work provides a powerful template for studying how cells of many types move through complex environments, with potential applications ranging from understanding immune surveillance to designing better cancer therapies.
Citation: Arruda, J., Alamoudi, E., Mueller, R. et al. Simulation-based inference of cell migration dynamics in complex spatial environments. npj Syst Biol Appl 12, 20 (2026). https://doi.org/10.1038/s41540-026-00648-9
Keywords: cell migration, dendritic cells, chemokine gradients, simulation-based inference, microfluidic pillar forests