Clear Sky Science · en
Metagenomics indicates an interplay of the microbiome and functional pathways in Parkinson’s disease
The Gut–Brain Story Behind a Familiar Disease
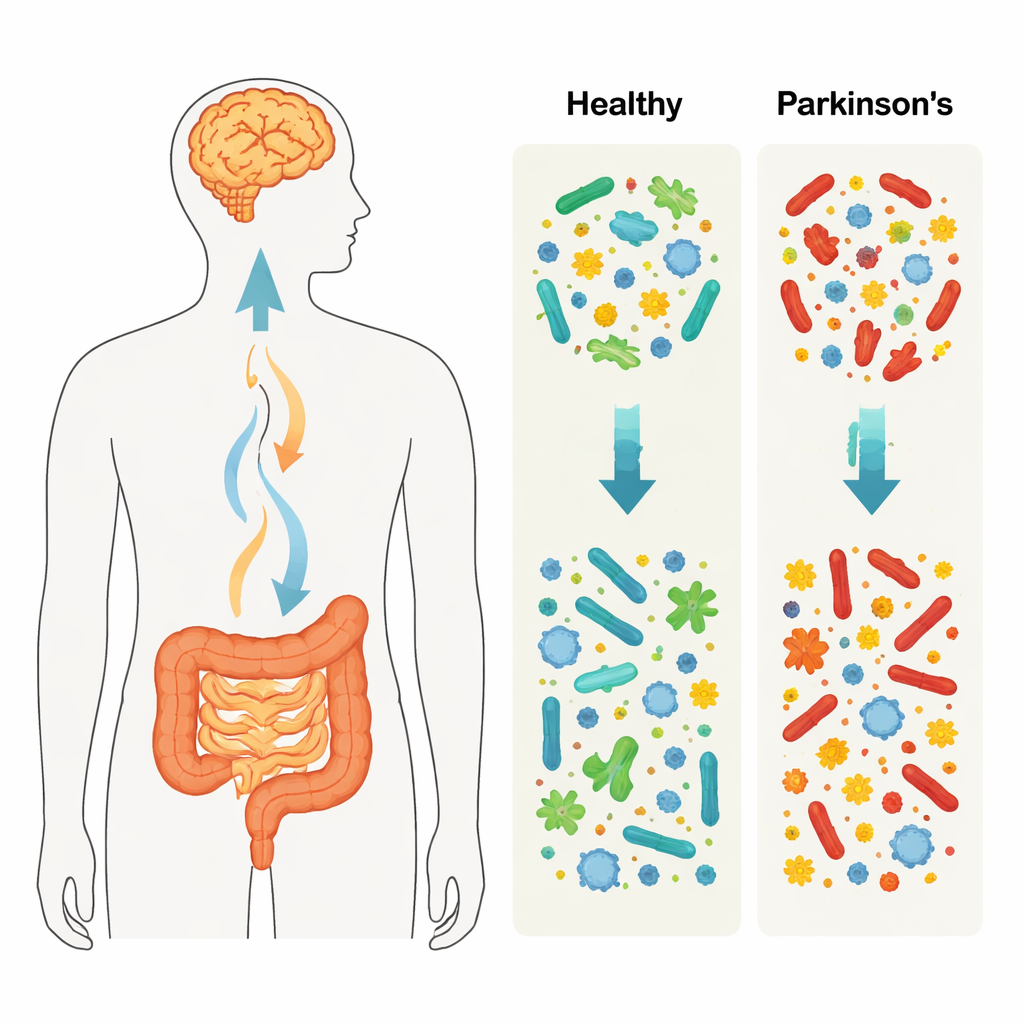
Parkinson’s disease is best known for its tremors and slowed movement, but the story may begin far from the brain—deep in the gut. Many people with Parkinson’s struggle with constipation and other digestive problems years before their first motor symptoms. This study explores whether the vast community of microbes living in our intestines, along with the chemical pathways they use, differs in people with Parkinson’s compared with healthy adults. By reading the genetic material in stool samples, the researchers looked for patterns that might one day help predict, explain, or even prevent this common brain disorder.
Who Was Studied and What Was Measured
The researchers compared stool samples from 55 North American adults with Parkinson’s disease to those from 42 healthy adults drawn from an existing public study. Instead of focusing on a few chosen microbes, they used whole genome sequencing, which captures DNA from bacteria, viruses, fungi, single-celled eukaryotes, and the genes that drive their metabolism. This allowed them to ask two basic questions: which organisms are present, and what are they genetically capable of doing? Alongside the microbial census, the team used standard ecological measures to describe how diverse and evenly balanced each person’s gut community was.
Shifts in the Gut’s Bacterial Residents
At the broadest level, the guts of people with Parkinson’s contained a different balance of major bacterial groups than those of healthy controls. Two large groups, Firmicutes and Actinobacteria, were relatively more abundant in Parkinson’s, while Bacteroidetes and some unclassified bacteria were less common. People with Parkinson’s had bacterial communities that were more even—no single type dominated—but they actually hosted fewer distinct kinds overall. When the researchers examined finer taxonomic levels, they identified dozens of specific bacterial lineages that differed between groups, painting a picture of widespread but nuanced reshaping of the gut ecosystem in Parkinson’s disease.
Viruses, Fungi, and Other Hidden Players
The team also looked beyond bacteria to the viruses that infect them (phages), DNA viruses more generally, and to fungi and protists. The collection of phages in Parkinson’s patients showed clear differences in family composition and diversity compared with healthy adults, suggesting that the viral predators of gut bacteria are also rearranged. Other DNA viruses and fungi likewise showed distinct overall community patterns between groups, although their absolute levels were low and varied a great deal from person to person. Protists were sparse and broadly similar, with one common gut inhabitant, Blastocystis, appearing more often in Parkinson’s samples but without strong statistical support. Together, these findings hint that Parkinson’s may be associated with multi-kingdom changes in the gut, not just shifts in bacteria alone.
Changes in Microbial Activity Pathways
Beyond who is there, the study asked what the gut microbes are poised to do. By linking genes in the sequencing data to known biochemical pathways, the researchers found that people with Parkinson’s had a broader and more evenly distributed set of microbial functions. Several pathways stood out as especially enriched. These included routes involved in energy production, fat burning, and recycling components of bacterial cell walls, as well as pathways that generate molecules tied to antioxidant defenses and purine metabolism. Others were related to processing transfer RNA, a type of molecule that can give rise to small fragments being explored as biomarkers in neurodegenerative diseases. While the study did not measure the actual chemicals produced, the genetic potential for these activities was clearly different in Parkinson’s guts.
What This Could Mean, and What It Does Not Yet Prove
The authors emphasize that their work is exploratory and comes with caveats. The healthy comparison group was drawn from another study that used different laboratory methods, and detailed lifestyle and diet information was not available, so some differences could reflect technical or environmental factors rather than Parkinson’s itself. Sparse detection of certain organisms and the absence of direct measurements of microbial byproducts limit how far the results can be linked to symptoms or disease progression. The study’s participants with Parkinson’s were also mostly white, highly educated, and relatively affluent, which may not reflect the broader patient population.
Why These Findings Matter Going Forward
Despite these limitations, the study strengthens the idea that Parkinson’s disease is entangled with changes in the gut ecosystem and its chemical output. The distinct patterns of bacteria, phages, fungi, and metabolic pathways suggest that the intestinal environment in Parkinson’s is remodeled in complex ways that may influence inflammation, oxidative stress, and the handling of molecules connected to brain health. For a layperson, the takeaway is that Parkinson’s might not be only a brain disease but part of a body-wide disturbance that begins, in part, in the gut. Future, more detailed studies that combine genetics, chemical measurements, and clinical data across diverse groups will be needed to turn these microbial fingerprints into reliable tools for earlier detection or new treatment strategies.
Citation: Park, S.J., Özdinç, B.E., Coker, K.G. et al. Metagenomics indicates an interplay of the microbiome and functional pathways in Parkinson’s disease. npj Parkinsons Dis. 12, 60 (2026). https://doi.org/10.1038/s41531-026-01271-5
Keywords: Parkinson’s disease, gut microbiome, metagenomics, gut–brain axis, microbial metabolism