Clear Sky Science · en
Semi-automated genomic newborn screening highlights complexities in reporting
Why Tiny Blood Spots Could Hold Big Answers
Within days of birth, most babies already undergo a quiet life-saving test: a heel prick that spots blood on a card. This study explores what happens if we go a step further and read much more of a baby’s genetic code from that same blood. The researchers in South Australia set out to see whether adding large-scale DNA analysis to routine newborn checks could safely find more treatable conditions early—without overwhelming families and doctors with confusing or unhelpful information.
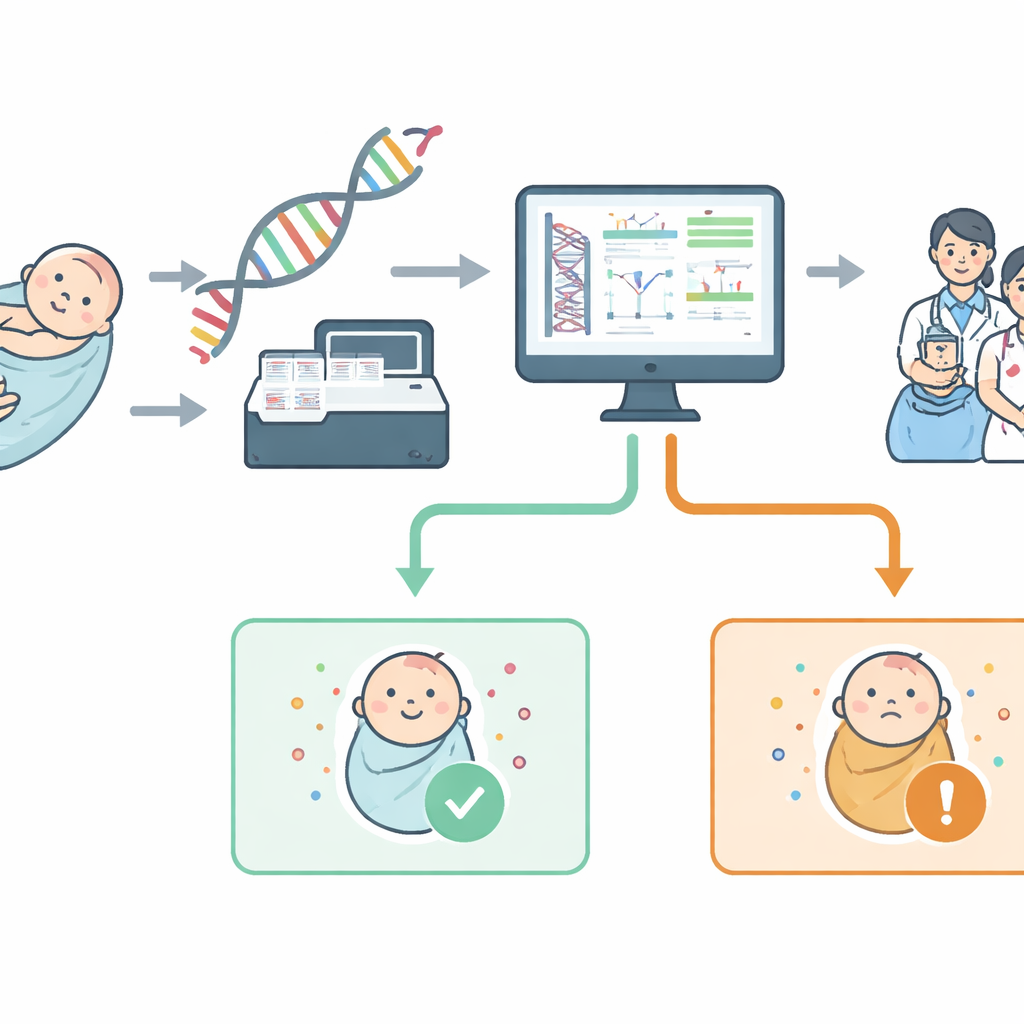
From Heel Prick to Genome Scan
Traditional newborn screening looks for a few dozen conditions by measuring chemicals in the blood. In contrast, genomic newborn screening reads stretches of DNA to look directly for disease-causing changes in specific genes. The NewbornsInSA project built a test based on whole-genome sequencing, but deliberately limited the search to a "virtual" panel of 613 genes. These genes were chosen with local doctors using simple rules: the condition should start in childhood, cause serious health problems, have a meaningful treatment or prevention strategy, and be reliably detectable in the lab. Public and patient groups then helped group these conditions into familiar body-system categories so parents could better understand what the test covers.
Putting the New Test Through Its Paces
Before offering this DNA-based check to new families, the team had to show it worked. They pulled 46 stored blood cards from children whose genetic diagnoses were already known but hidden from the analysts. Using whole-genome sequencing and two separate software systems, they asked whether their pipeline would correctly flag babies who truly had one of the target conditions. The result was encouraging: the process found 97% of the true cases and produced no false alarms after expert review. The one missed case highlighted a key limitation of any gene-based screen—when the scientific evidence behind a specific DNA change is incomplete, computers and even experts may be forced to label it “uncertain” and treat it as a non-result.
Letting Computers Do the Heavy Lifting
Reading an entire genome generates thousands of variants for each baby, far too many for humans to inspect one by one. To cope with this, the researchers built a semi-automated workflow. Custom scripts, plugged into a commercial analysis platform, scan the annotated variant list and automatically set aside babies who clearly have no worrying findings, labeling them as “low chance.” Only cases with changes that look potentially harmful—such as known disease-causing variants or likely damaging disruptions of a gene—are passed to expert analysts for detailed review. In the first 100 real newborns tested, this automation immediately cut the number of babies needing manual review by more than half, a crucial step if such screening is to be scaled to entire populations.
Real Babies, Real Decisions
When the team applied the workflow to the first 100 enrolled newborns, five babies received results suggesting a high chance of a specific condition. These included heart rhythm disorders that can cause dangerous irregular beats, a metabolism problem already suspected by standard screening, a gene change that makes certain antibiotics risky for hearing, and a mild form of inherited high blood sugar. In each case, the result triggered tailored follow-up: heart checks and ongoing monitoring, electronic alerts to avoid particular drugs, or planning for future pregnancy care. At the same time, many other findings were deliberately not reported because they were tied to mild symptoms, adult-onset cancers, uncertain risk, or complex patterns that current knowledge cannot confidently interpret in a healthy newborn. Those tricky calls required hours of reading the literature and discussion among geneticists and specialists, underlining that expert judgment still matters.
Balancing Early Help and Future Burden
This work shows that it is technically possible to use whole-genome data from a tiny blood spot to spot more treatable childhood conditions than standard tests, with high accuracy and support from clever software. But it also lays bare the ethical and practical tightrope: every extra gene checked raises the chance of uncovering information that may not help the baby now, may worry parents, or may only become relevant decades later. The NewbornsInSA team’s semi-automated, carefully curated approach—where computers rule out clear negatives and humans focus on nuanced cases—offers a blueprint for how genomic newborn screening might be rolled out responsibly, while ongoing follow-up studies track its real-world benefits and risks for families.
Citation: Chowdhury, A., Marri, S., Anastasi, L. et al. Semi-automated genomic newborn screening highlights complexities in reporting. npj Genom. Med. 11, 13 (2026). https://doi.org/10.1038/s41525-026-00553-4
Keywords: genomic newborn screening, whole genome sequencing, dried blood spots, genetic variant interpretation, early detection of rare diseases