Clear Sky Science · en
Real-time, high-resolution metabolic characterization of live bacteria using label-free optical metabolic imaging
Watching Germs React in Real Time
Bacteria are everywhere on and inside us, quietly shaping our health. They help digest food, protect our skin, and sometimes cause stubborn infections that shrug off antibiotics. This study introduces a way to watch living bacteria react to stress and drugs in real time, at the level of individual cells, without adding any dyes or labels. That ability could eventually help doctors choose the right treatment faster and help scientists understand why some infections are so hard to cure.
Light as a Window into Bacterial Life
Instead of adding stains or genetic tags, the researchers rely on the fact that bacteria naturally glow very faintly when certain molecules inside them are excited by laser light. Two of these molecules, known as NAD(P)H and FAD, are central players in how cells turn food into energy. By using rapid, high-resolution laser microscopy, the team can measure both the brightness and the tiny time delay of this glow, revealing how busy a cell’s metabolism is. Because the method is label‑free and non‑destructive, the same live bacteria can be followed over seconds and minutes as their environment changes. 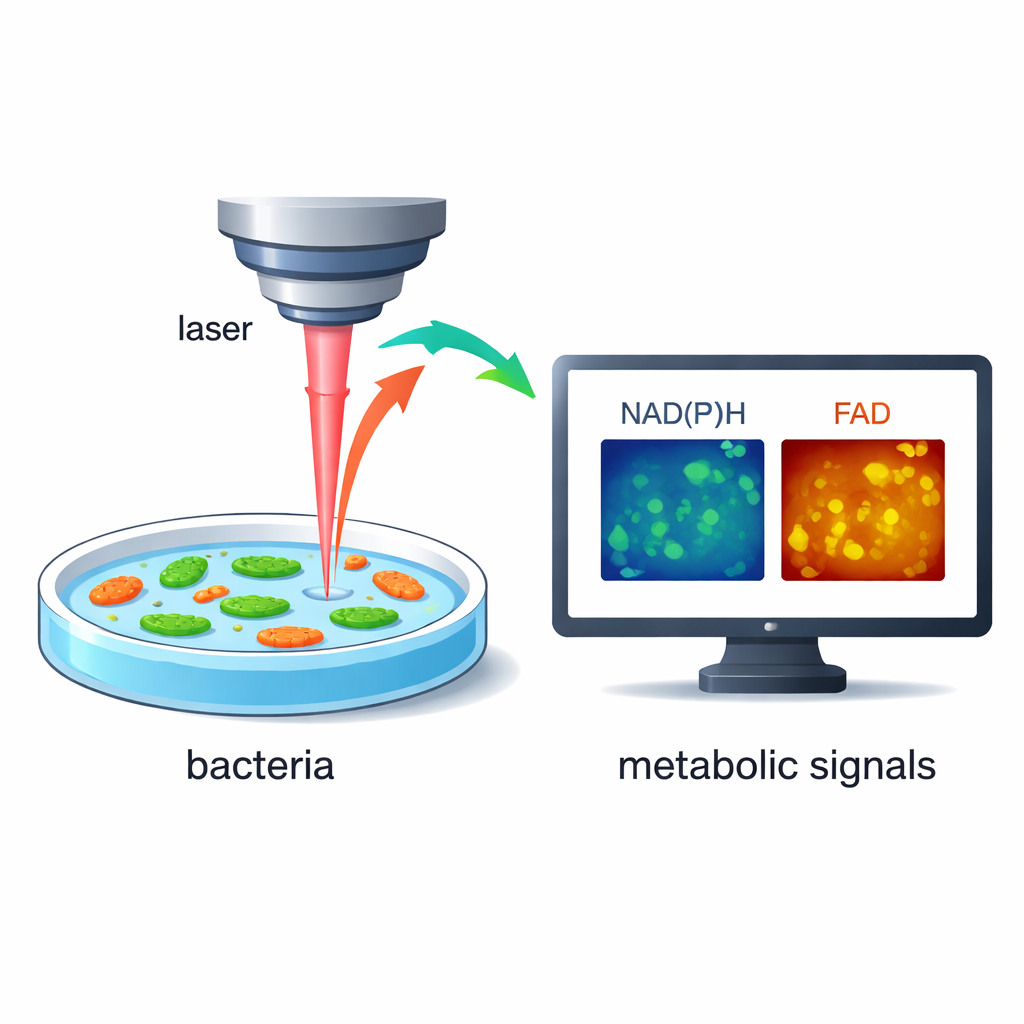
Seeing Differences Between Individual Bacteria
Most standard lab tests lump together millions of bacteria and report only an average response. Here, the authors show that their imaging method can pick out single Staphylococcus aureus cells in liquid culture and analyze each one separately, even at high cell densities. When they compared two colonies of S. aureus grown under the same conditions, one key signal (from FAD) differed between colonies, and there was a spread of values within each colony as well. This means that even genetically similar bacteria in the same flask can sit in very different metabolic states, a hidden diversity that standard bulk tests might miss but that may matter for how infections respond to treatment.
Watching Antibiotics Strike Within Seconds
The team then followed individual S. aureus cells as they were exposed to several antibacterial treatments, including common antibiotics and household bleach. Using their fast imaging setup, they captured changes from the moment the drug solution touched the cells through the next 30 minutes. Strong treatments caused NAD(P)H signals to drop and FAD signals to rise, often within seconds, reflecting stalled energy production and a shift toward a more oxidized state inside the cell. The exact timing and pattern of these changes depended on the drug type and dose: for example, protein‑blocking and cell‑wall‑targeting antibiotics produced a brief dip and partial recovery, while bleach drove a sharp, sustained shift. These distinct “light signatures” suggest that the technique can sense not only whether cells are stressed or dying, but also how they are being damaged. 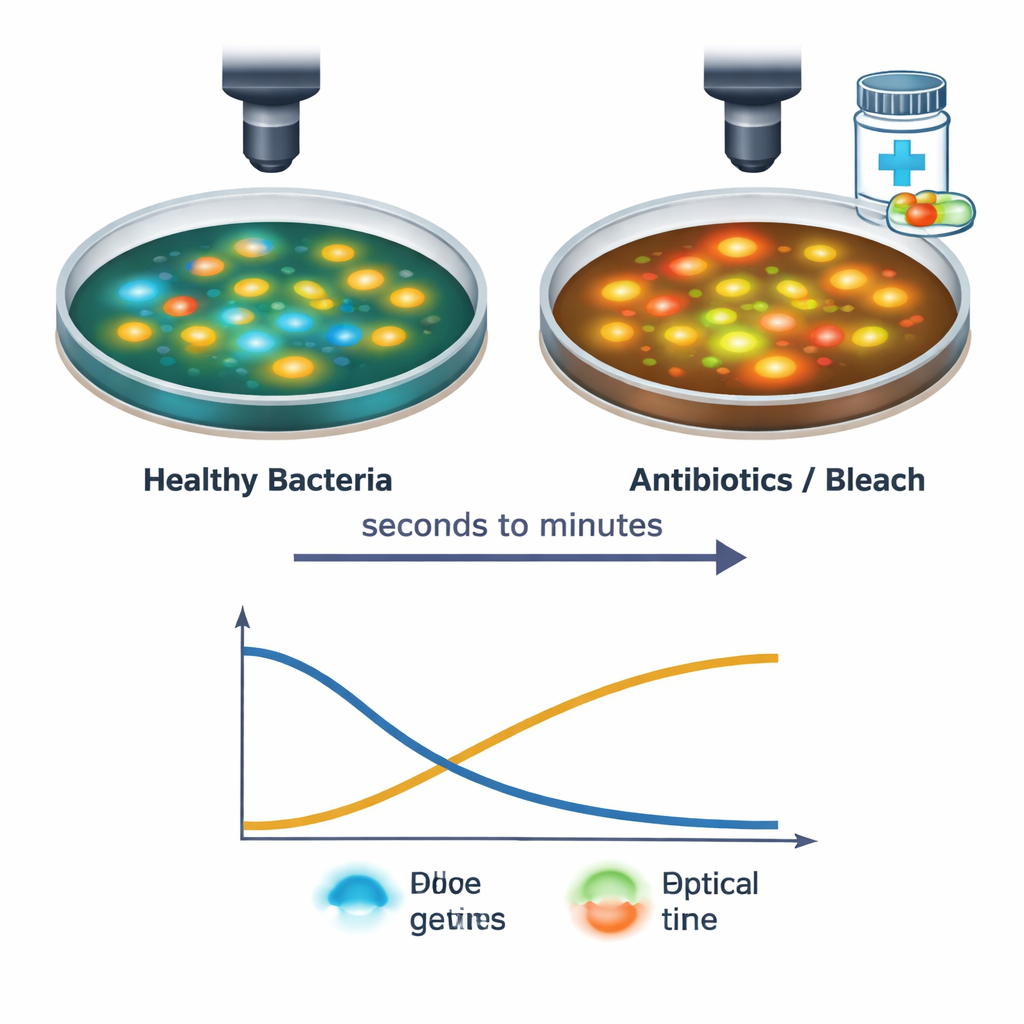
Peering into the Hidden World of Biofilms
Bacteria rarely live alone in nature; instead, they build slimy, protective communities called biofilms on surfaces ranging from medical implants to teeth. In these dense layers, some cells may be highly active while others lie dormant, a mix that helps the community survive antibiotics. Using large stitched images, the researchers mapped the metabolic landscape within S. aureus biofilms and compared it with free‑swimming cells. They found bright, metabolically active pockets surrounded by less active regions and showed that, across several species, biofilm bacteria tended to have a more oxidized metabolic profile than their free‑living counterparts. Additional optical signals suggested extra lipids and proteins, consistent with the glue‑like material that holds biofilms together.
Why This Matters for Future Medicine
Together, these results show that label‑free optical metabolic imaging can provide rapid, detailed, and non‑invasive snapshots of how bacteria live, adapt, and die. For a lay reader, the key message is that scientists can now “listen in” on bacterial energy use in real time, without disturbing the cells, using only carefully tuned light. In the long run, this kind of technology could help identify hard‑to‑culture bacteria, spot pockets of drug‑tolerant cells inside biofilms, or quickly test how a patient’s infection responds to different antibiotics. That could support faster, more tailored treatments and deepen our understanding of the invisible microbial worlds that affect our health every day.
Citation: Sorrells, J.E., Yang, L., Iyer, R.R. et al. Real-time, high-resolution metabolic characterization of live bacteria using label-free optical metabolic imaging. npj Biofilms Microbiomes 12, 55 (2026). https://doi.org/10.1038/s41522-026-00920-0
Keywords: bacterial metabolism, optical imaging, biofilms, antibiotic response, microbiome