Clear Sky Science · en
Stages of biomolecular condensate formation in pro-β-carboxysome assembly
Why tiny carbon factories matter
Earth’s food chains depend on photosynthesis, yet the key enzyme that pulls carbon dioxide (CO2) from the air, Rubisco, is slow and easily confused by oxygen. Cyanobacteria – microscopic, photosynthetic microbes – long ago solved this problem by building protein "micro-factories" called carboxysomes that trap CO2 around Rubisco. This study unravels how one crucial adaptor complex, made of proteins called ApN and CM, self‑assembles in stages to help build a working carboxysome. Understanding this choreography could guide efforts to install similar CO2‑boosting systems into crop plants and raise yields.
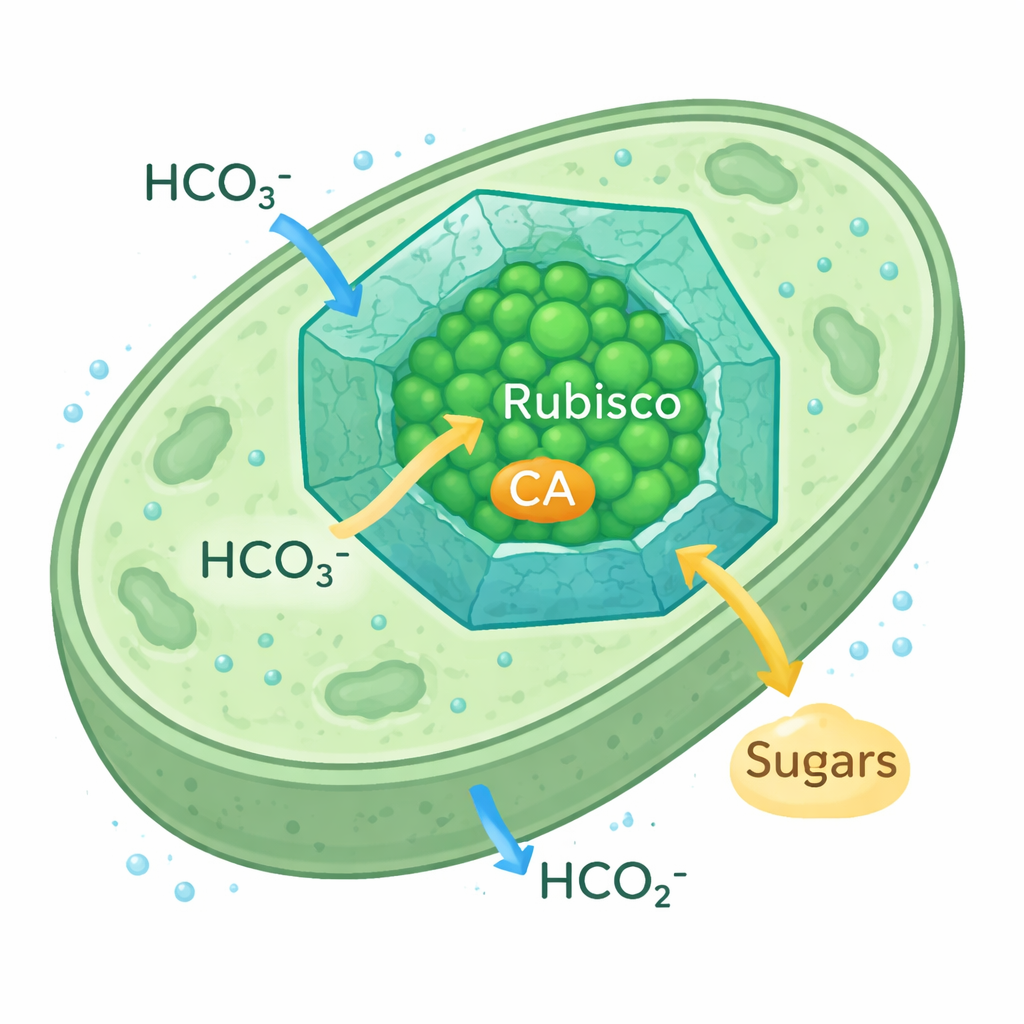
How microbes supercharge carbon capture
Cyanobacterial carboxysomes are tiny, icosahedral protein compartments that sit inside the cell. Their protein shell lets bicarbonate ions in while slowing the escape of CO2, creating a CO2‑rich bubble around Rubisco and its partner enzyme, carbonic anhydrase (CA), which converts bicarbonate to CO2. This CO2‑concentrating mechanism allows cyanobacteria to thrive even when CO2 in the environment is scarce. Because the Rubisco form used in these carboxysomes resembles plant Rubisco, researchers see them as a realistic model for re‑engineering plant chloroplasts to capture carbon more efficiently.
The puzzle piece that talks to the shell
Before the hard protein shell forms, cyanobacteria first build a soft, droplet‑like "pro‑carboxysome" condensate. Earlier work showed that a scaffolding protein called CM brings Rubisco and CA together into this droplet. The missing link was how the shell adaptor protein ApN joins this structure and later connects it to shell proteins. The authors found that when ApN is made alone it forms a four‑part bundle that ignores the other carboxysome components. In this state ApN cannot be pulled into droplets with Rubisco, CM and CA, suggesting that a different form of ApN must be used during real carboxysome assembly.
Co‑built adaptors and a redox switch
Looking at the genetic layout in cyanobacteria, the team noticed that apn sits directly behind cm in the same operon, so the two proteins are usually made together. Mimicking this in bacteria, they co‑expressed ApN and CM from a shared piece of DNA. Under these conditions ApN no longer formed a stand‑alone tetramer. Instead, three ApN subunits and one CM subunit assembled into a specific four‑part adaptor complex, denoted (ApN)3:CM. This hetero‑complex attached readily to Rubisco, CA and CM and could join condensates. Structural modeling and cryo‑electron microscopy showed that ApN and CM meet through a finely tuned "tongue‑and‑groove" interface; when key ApN residues in this interface were mutated, the adaptor would not form and cells failed to build proper carboxysomes.
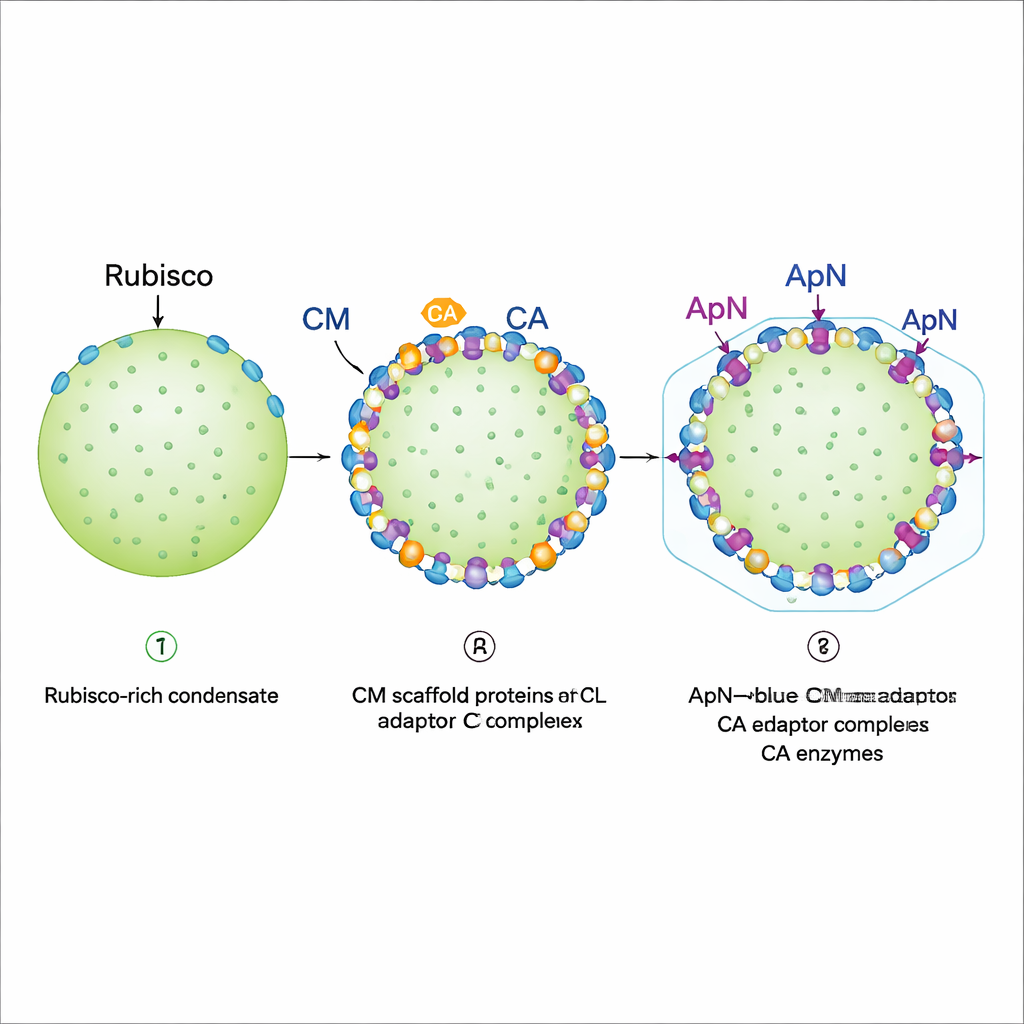
Positioning at the edge and preparing for the shell
Fluorescence imaging of reconstituted droplets revealed a stepwise assembly program. First, Rubisco, full‑length CM and a shorter CM variant form a dense condensate that behaves like a gel: Rubisco is essentially locked in place. Next, CA and the (ApN)3:CM complex are added. Both preferentially settle at the outer rim of the existing droplet, provided CA arrives before or together with (ApN)3:CM. This edge localization is crucial because the tail of ApN carries an "encapsulation peptide" that binds shell proteins. The authors also discovered a built‑in redox switch: as the interior of the maturing carboxysome becomes more oxidizing, two conserved cysteine residues in ApN can be oxidized, nudging the adaptor from a (ApN)3:CM tetramer toward a (ApN)2:CM trimer. In living cyanobacterial cells, altering these cysteines slowed growth and produced oversized but still functional carboxysomes, underscoring their role in fine‑tuning maturation.
Lessons for building better crops
To a non‑specialist, these molecular details may sound arcane, but they boil down to a simple message: timing and partnership matter. The work shows that ApN must be co‑produced with CM to avoid forming a useless self‑cluster and instead become an adaptor that parks at the surface of the pro‑carboxysome, ready to recruit the shell. A redox‑sensitive step then helps remodel this adaptor as the compartment closes and its internal chemistry changes. For plant engineers hoping to transplant cyanobacterial CO2‑concentrating machinery into crops, the study highlights that it is not enough to move individual parts; one must also respect the order, ratios and chemical triggers that guide carboxysome assembly in nature.
Citation: Zang, K., Hong, X., Nguyen, N.D. et al. Stages of biomolecular condensate formation in pro-β-carboxysome assembly. Nat. Plants 12, 447–464 (2026). https://doi.org/10.1038/s41477-026-02227-6
Keywords: carboxysome assembly, cyanobacterial CO2-concentrating mechanism, Rubisco condensates, biomolecular condensates, synthetic photosynthesis