Clear Sky Science · en
Structural dynamics of the midnolin-proteasome during ubiquitin-independent substrate turnover
How Cells Take Out the Trash in a Hurry
Inside every cell, proteins are constantly being made and destroyed. Many are tagged with small molecules called ubiquitin before they are fed into the cell’s main recycling machine, the proteasome. But some nuclear proteins must be removed so quickly that there’s no time for tagging. This study reveals how a little-known helper protein, midnolin, plugs directly into the proteasome to clear these urgent targets, offering clues to how cells keep gene activity under tight, rapid control.
A Shortcut to the Cellular Shredder
The 26S proteasome is a barrel-shaped complex that chews up proteins into small pieces. Traditionally, it recognizes targets by reading chains of ubiquitin attached to them. Recent work uncovered an alternative route: a “shortcut” pathway in which midnolin escorts certain nuclear proteins—especially immediate early gene products that rapidly turn genes on and off—straight to the proteasome without any ubiquitin tags. These proteins, such as transcription factors involved in stress responses and inflammation, must appear and disappear within minutes. The new paper combines high-resolution cryo-electron microscopy with biochemical experiments to map, nearly frame by frame, how midnolin latches onto the proteasome and positions its cargo for destruction.
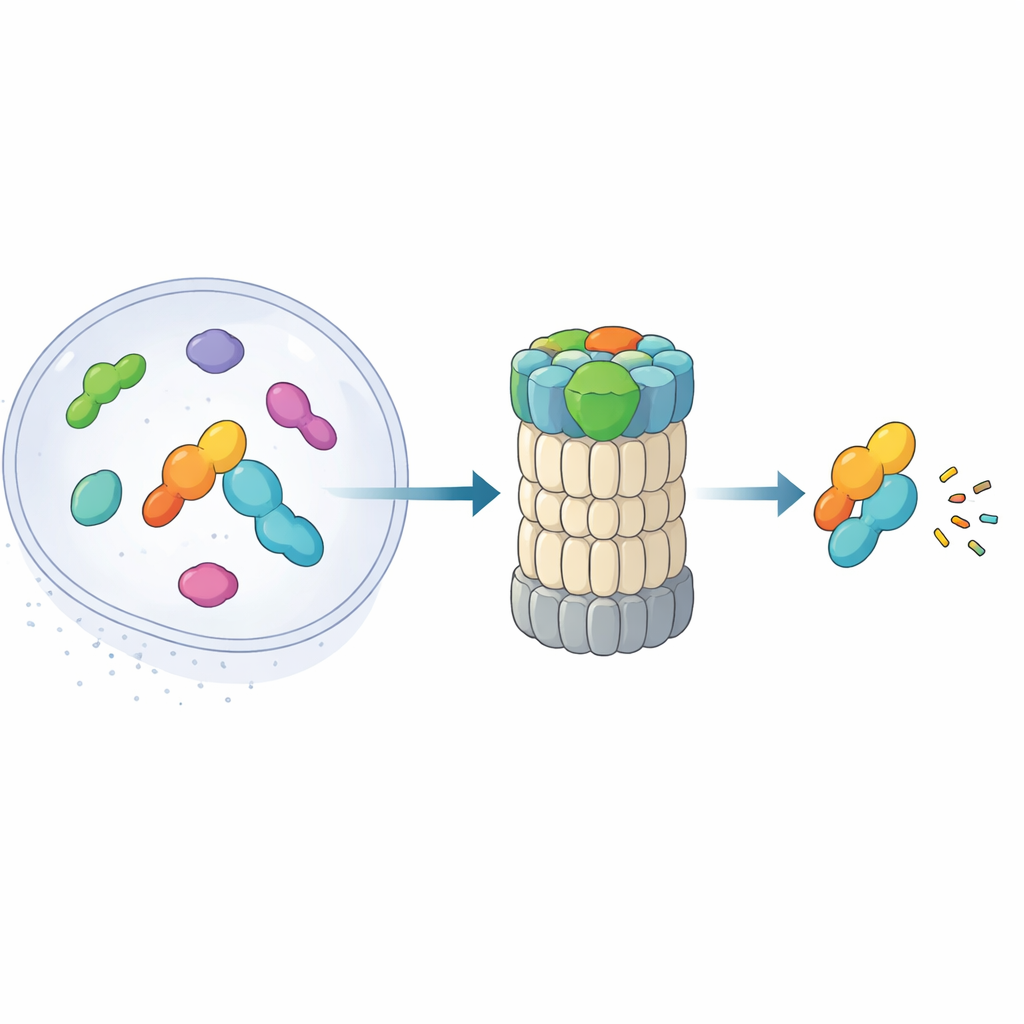
The Three-Part Helper That Grabs and Aims Targets
Midnolin works like a multi-tool with three key modules. One end, a C-terminal helix, plugs into a docking site on a proteasome component called RPN1, anchoring midnolin to the recycling machine. A central “Catch” region recognizes and grips a specific strand-like feature (a degron) in its target proteins. At the opposite end sits a ubiquitin-like domain. Surprisingly, this domain is not mainly for binding the proteasome, as similar domains do in other shuttle proteins. Instead, the authors show that it nestles against RPN11, an enzyme that normally trims ubiquitin from substrates. Here, RPN11 acts more like a scaffold: by embracing midnolin’s ubiquitin-like segment, it helps hold the Catch domain and its bound cargo directly above the narrow entry pore into the proteasome.
Guiding the Protein into the Grinding Core
Once midnolin has docked, the proteasome’s energy-hungry motor—an AAA+ ATPase ring—takes over. This ring sits just beneath the entry pore and pulls unfolded protein chains into the core, step by step. The cryo-EM snapshots reveal a series of “spiral staircase” positions adopted by the motor’s six subunits as they grip and hand off the substrate through the central channel. The authors captured four consecutive stages of this cycle, all with midnolin-bound material threaded through the pore, and even saw how the release of a magnesium ion in one subunit coincides with a downward shift of the entire ring. This suggests that coordinated ATP use and magnesium release help ratchet the protein deeper into the chamber where cutting sites reside.
Why Positioning and Co-Expression Matter
The team tested how each midnolin module contributes to efficiency. The anchoring helix alone can bind RPN1 but cannot significantly boost proteasome activity. Removing the Catch domain leaves many proteasomes stuck in an idle, substrate-free state, even though midnolin can still attach; replacing the Catch with a permanently fused substrate also fails to restore full activity. These findings indicate that Catch does more than just grab the target—it must also orient the protein precisely at the pore entrance. Biochemical and tumor transcriptome data further show that midnolin tends to be produced together with its substrates. When co-expressed, midnolin folds better and forms stable complexes, creating a ready-made “degrade me” package that can be quickly delivered to the proteasome when the cell needs to reset gene expression.
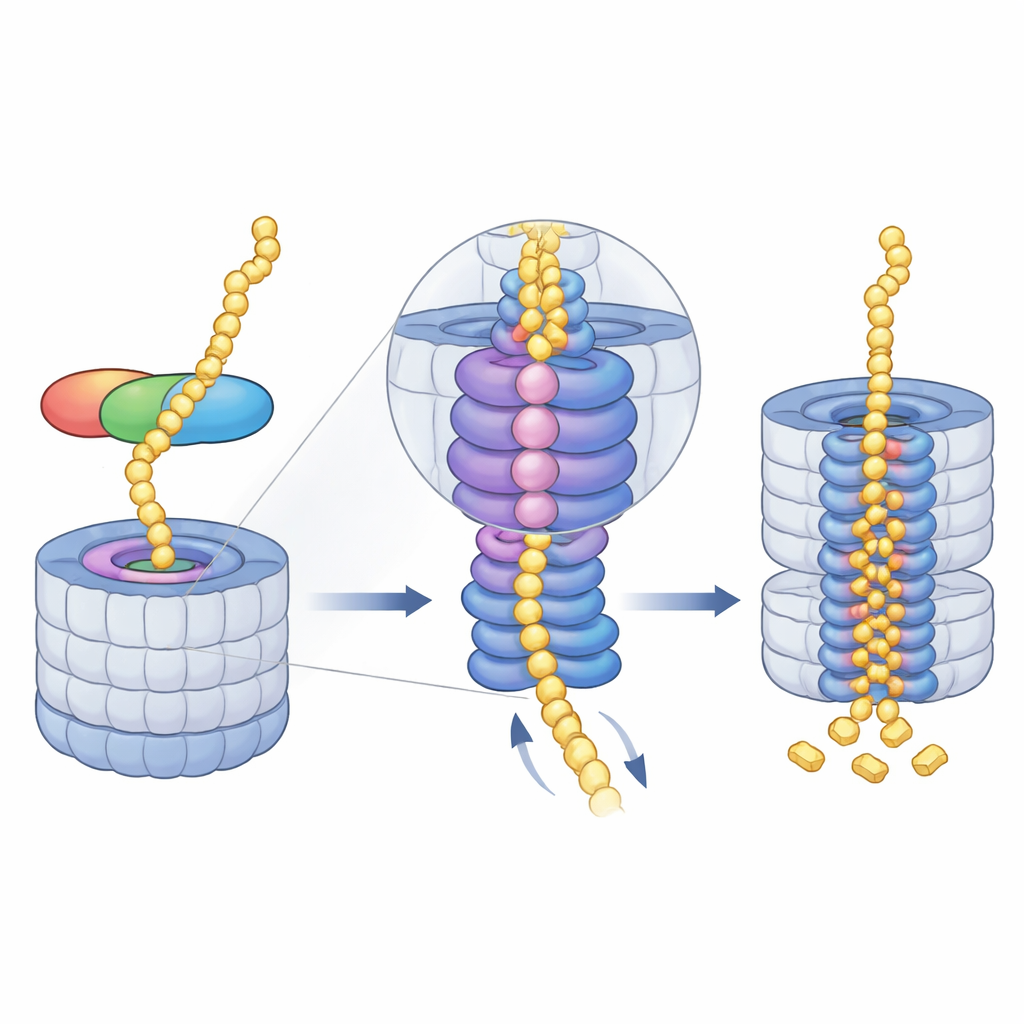
Self-Cleanup and Future Medical Possibilities
After the target protein has been pulled in and chopped up, midnolin itself is not spared. Its own unstructured regions can be threaded into the motor and fed into the same destruction chamber, ensuring that the helper does not accumulate unnecessarily. By illuminating this ubiquitin-free route to the proteasome, the study expands our understanding of how cells manage protein lifetimes on very tight schedules. It also hints at new therapeutic strategies: by engineering midnolin-based molecules that tether disease-causing nuclear proteins directly to the proteasome, researchers may one day be able to eliminate “undruggable” targets, including certain cancer drivers, more rapidly and selectively.
Citation: Zhu, C., Qin, L., Dai, Z. et al. Structural dynamics of the midnolin-proteasome during ubiquitin-independent substrate turnover. Nat Commun 17, 2800 (2026). https://doi.org/10.1038/s41467-026-71002-0
Keywords: proteasome, midnolin, protein degradation, ubiquitin-independent pathway, immediate early genes