Clear Sky Science · en
Transcriptional competence defines the heterochromatin nucleating potential of isolated MSR units
Hidden switches in our DNA
Our genomes are packed inside the cell nucleus in two main states: active regions that host genes, and deeply packed stretches long thought of as genetic “dark matter.” This study asks a deceptively simple question: what tells a patch of DNA to become this tightly packed, gene-silencing material—known as heterochromatin—in the first place? By dissecting a specific class of repetitive DNA in mice, the authors reveal that not all repeats are equal: only those able to support a special kind of transcription can flip the switch that builds and maintains these silent DNA neighborhoods.
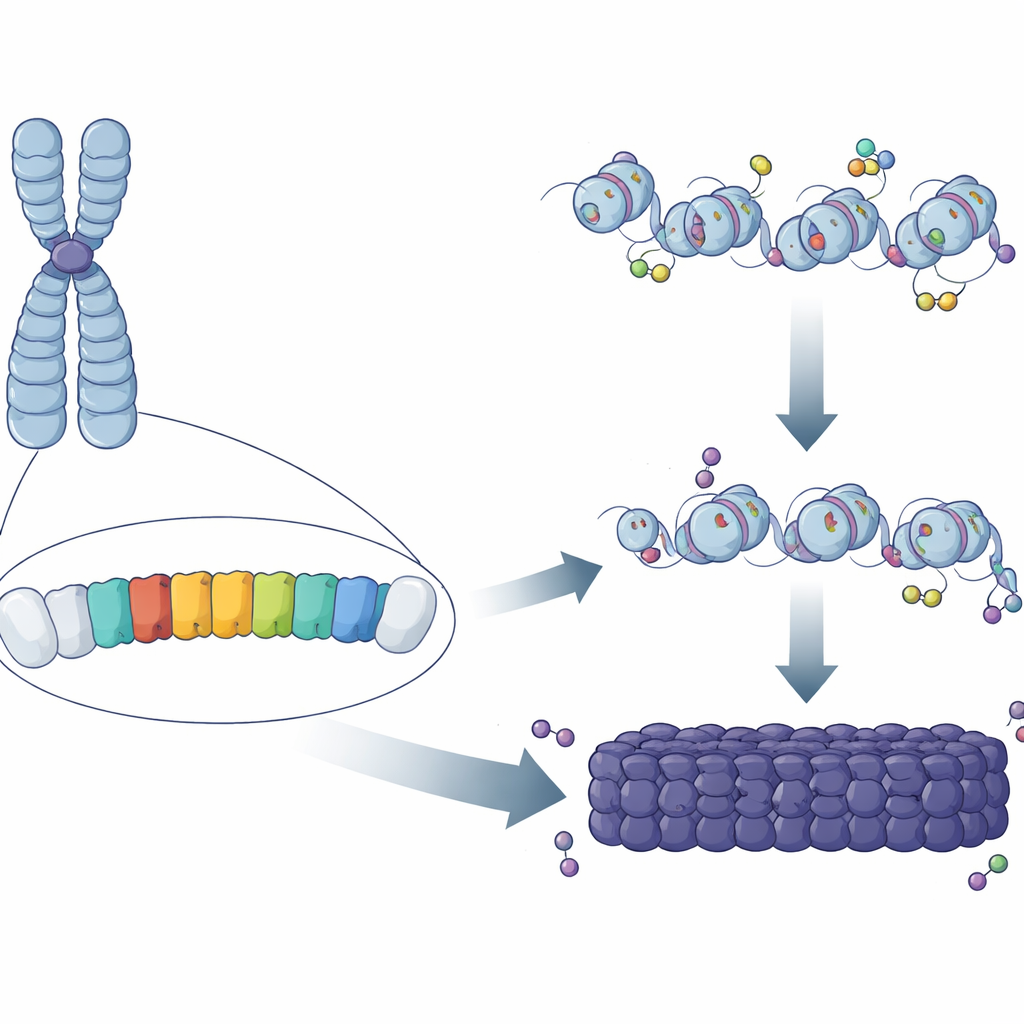
Repeating patterns in the genome
Nearly half of mammalian DNA is made of repeated sequences, many of them clustered in regions around the chromosome center. In mice, one major component of these regions is the “major satellite repeat” (MSR), a short A/T-rich DNA sequence copied hundreds of thousands of times. Classical work showed that these regions are coated with chemical marks and proteins that lock the DNA into a compact, protective state. But it remained mysterious why some MSR copies became fully heterochromatic while others, scattered elsewhere in the genome, did not. The authors reasoned that small differences in the sequence or behavior of individual MSR units might determine whether they can seed, or “nucleate,” a patch of heterochromatin.
Building a test site in the genome
To test this idea cleanly, the team engineered mouse embryonic stem cells to contain an artificial landing pad in a quiet stretch of chromosome 2—a region with no nearby genes or repeats and no detectable activity. Into this neutral site they inserted different DNA fragments: intact MSR units, heavily scrambled MSR variants, and control elements such as viral promoters or sections of mobile elements. This allowed them to ask, unit by unit, which sequences are able to attract the hallmark heterochromatin features: a specific chemical tag on histone proteins (H3K9me3), binding of HP1 proteins, and incorporation of the linker histone H1, all of which together thicken and stabilize the local chromatin.
Only transcription-ready repeats seed silent chromatin
The results were strikingly selective. A single intact MSR unit inserted at the test site was not enough to change the chromatin. However, three or more tandem copies of the intact MSR sequence converted the surrounding region into a heterochromatin “island,” with strong H3K9me3, HP1, and histone H1. By contrast, equally long stretches of scrambled MSR sequence, or of another repeat type (LINE-1 5' untranslated region), failed to do so, even though they could drive strong transcription. The key difference was that multi-copy intact MSR units supported a modest, bi‑directional transcription that produced short, non‑standard RNA molecules tightly associated with chromatin. This pattern, rather than high, gene-like transcription, correlated with the ability to nucleate heterochromatin.
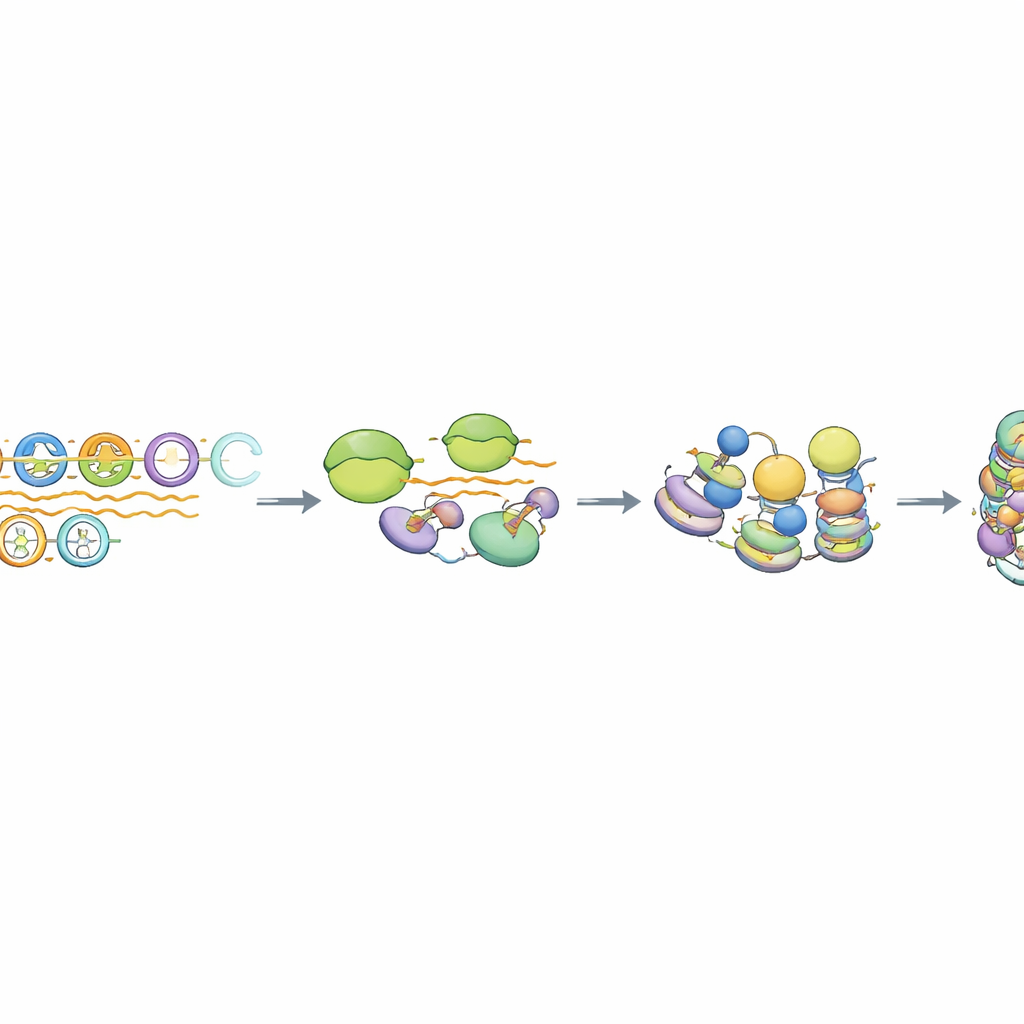
A special kind of transcription and RNA processing
Digging deeper, the authors found that RNA polymerase II, the enzyme that normally makes messenger RNA, briefly engages the MSR arrays but does not proceed efficiently into long transcripts. The resulting RNAs lack typical mRNA hallmarks such as a protective 5' cap and long poly‑A tails, and they remain near the DNA that produced them. A protein machine called the Integrator complex, known for trimming and terminating certain noncoding RNAs, was specifically enriched at intact MSR units. When Integrator’s cutting activity was reduced, MSR-derived RNA levels rose substantially, but the key repressive histone mark persisted while HP1 binding changed subtly. Genome‑wide analysis showed that only the most intact ~10–15% of MSR copies behave this way, highlighting a subset of “competent” repeats wired for this transcription‑coupled silencing pathway.
Unwound DNA as a promoter mimic
The team also explored how MSR DNA itself encourages this unusual transcription. Multi-copy MSR arrays, but not single or double copies, showed clear signs of locally unwound DNA and RNA:DNA hybrids, structural features often seen near active promoters and at pause sites. These configurations were enhanced when topoisomerase enzymes were inhibited, and they coincided with greater MSR transcription and stronger heterochromatin features. The authors propose that three or more tandem MSR units create a physical DNA topology that mimics a promoter, inviting polymerase and transcription factors to engage just enough to generate short RNAs that, together with specific proteins, reinforce a compact chromatin architecture.
Why this matters for genome health
To a lay observer, this work reveals that parts of our “junk” DNA act as carefully tuned switches, using a blend of DNA shape, low‑level transcription, and RNA processing to build the genome’s protective shell. Only MSR units that can support this controlled, non‑messenger transcription can spark new heterochromatin, while scrambled or overly active elements cannot. This DNA/RNA-based logic helps explain how cells distinguish between regions to be kept silent and those allowed to host genes, and why mis‑regulated satellite RNAs are linked to cancer and developmental problems. In essence, the study shows that the genome’s repetitive “background” is not passive filler but an active engineer of nuclear architecture and stability.
Citation: Lo, YH., Shukeir, N., Erikson, G. et al. Transcriptional competence defines the heterochromatin nucleating potential of isolated MSR units. Nat Commun 17, 2653 (2026). https://doi.org/10.1038/s41467-026-70991-2
Keywords: heterochromatin, satellite DNA, noncoding RNA, chromatin structure, genome stability