Clear Sky Science · en
TONSOKU prevents the formation of large tandem duplications and restrains ATR–WEE1 checkpoint activation
When DNA Repair Goes Wrong in Plants
Our genomes, and those of crops and cancers, are constantly battered as cells copy their DNA. Usually, repair systems quietly fix problems before they cause trouble. This study looks at what happens when one such system, built around a protein called TONSOKU (TSK), is missing in a small weed plant often used in research. The result is a surprising surge of big DNA duplications and odd growth patterns—insights that could matter for both plant breeding and understanding human disease.
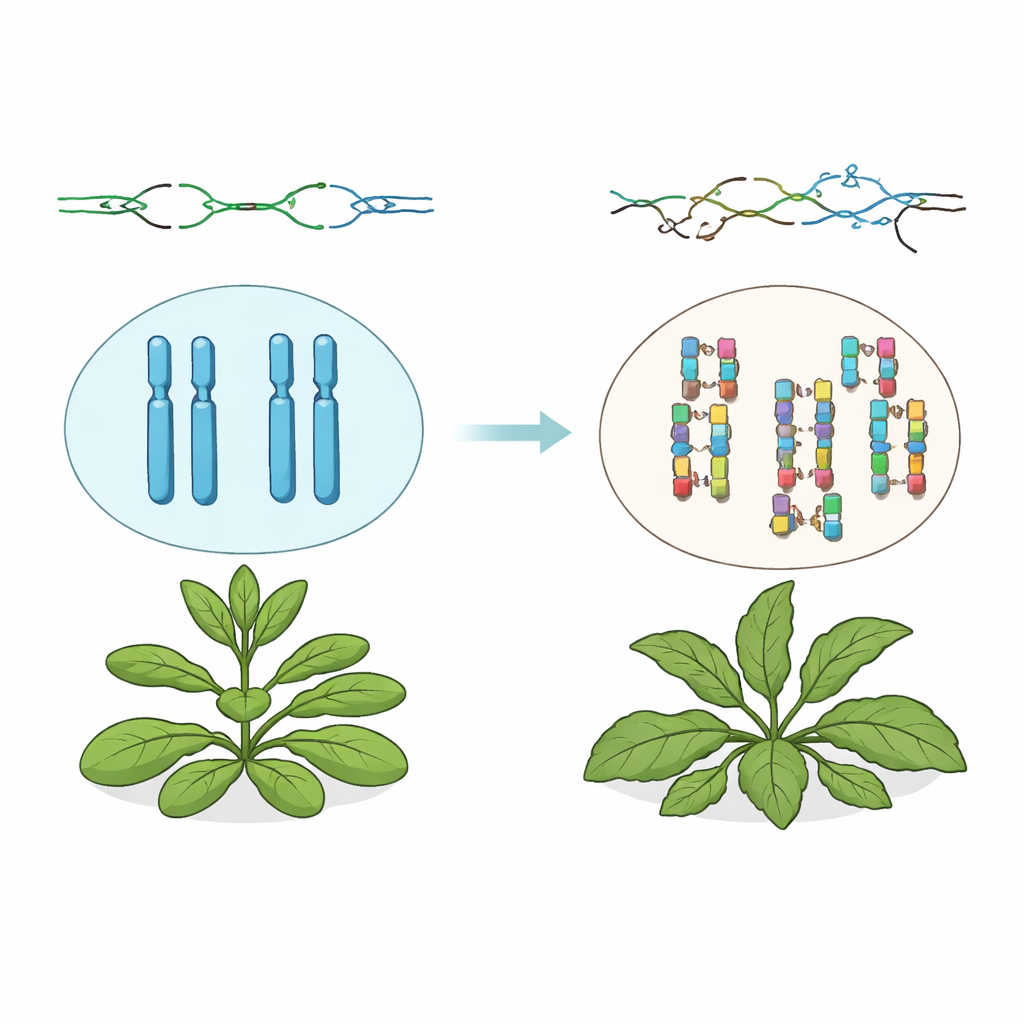
Extra Copies of DNA Pile Up
The authors worked with Arabidopsis thaliana plants that lack a working TSK gene. Although these plants are able to live and reproduce, their genomes tell a dramatic story. Using whole-genome DNA sequencing, the team found that, while small mutations stayed near normal levels, large stretches of DNA were frequently copied twice in a row—so-called tandem duplications. These added pieces ranged from under a thousand DNA letters to nearly one and a half million, often increasing total genome size by about 7 percent. Many of these duplications were unique to individual plants, showing that they arise repeatedly and independently as the plants grow and across generations.
Duplications Reshape Genes and Their Activity
These extra DNA segments are not just empty cargo. When the duplicated regions contain genes, those genes tend to be more active, simply because there are now more copies. By combining DNA sequencing with measurements of RNA (the readout of gene activity), the researchers showed that genes within duplicated regions often produced several times more RNA than their counterparts in normal plants. This boost can spill over, influencing genes outside the duplicated zones as well. Over time, such copy number changes can alter traits in major ways—something long exploited in crop domestication and often seen in human tumors.
A Backup Repair Pathway Creates the Damage
To find out how these duplications form, the team zoomed in on the ends of the duplicated segments. The patterns they saw matched the fingerprints of a fall-back DNA repair route called theta-mediated end joining, which relies on an enzyme known as Polymerase theta. When the usual high-precision repair system that depends on TSK fails, this backup pathway steps in to patch broken DNA. It does so by using tiny matching sequences as glue, which can easily lead to copying a chunk of DNA twice. Optical mapping of very long DNA molecules confirmed that the extra pieces sit right next to their original counterparts on the chromosomes, forming true tandem duplications rather than separate circles of DNA.
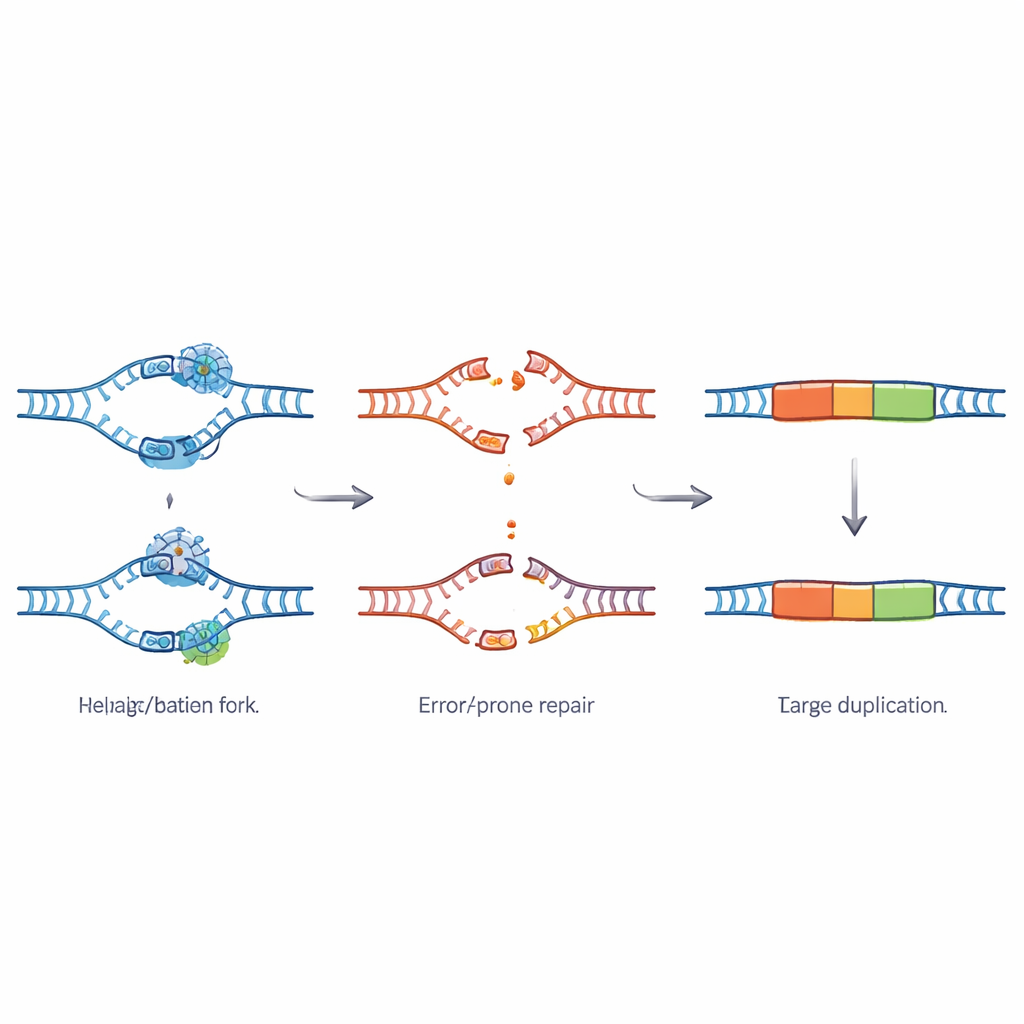
Stress Hotspots in the Genome
The duplicated regions are not scattered randomly. They tend to appear in parts of the genome that replicate late during cell division and that are packaged into tight, repetitive DNA known as heterochromatin. These areas are rich in special repeated motifs and in mobile DNA elements that are known to cause headaches for the DNA copying machinery. The borders around duplication sites also show signs of chronic stress: they contain more natural variation, longer stretches of repeats, more transposable elements, and more DNA replication start sites than typical regions. All of this points to these zones as hotspots where the copying of DNA frequently stalls or breaks down, especially when TSK is missing.
From Damaged DNA to Odd-Shaped Plants
Arabidopsis plants without TSK do not just carry scarred genomes; they also look strange. Many show twisted patterns of leaves and flowers, thick or split stems, and unusual branching. The researchers asked whether these visible quirks come from the duplications themselves or from the cell’s alarm system reacting to ongoing DNA damage. By disabling key players in the plant DNA damage response—proteins that sense stalled DNA replication and pause the cell cycle—they largely restored normal plant shape, even though the duplications kept forming. This shows that the developmental defects are mainly caused by chronic activation of the DNA damage response, not directly by the extra DNA copies.
Why This Matters Beyond One Weed
This work reveals a previously hidden route by which large DNA duplications can arise when a conserved repair pathway fails, linking replication stress, error-prone repair, and visible changes in an organism’s form. Similar TONSOKU-like proteins and repair systems exist in animals, including humans. In some cancers, genomes are peppered with large tandem duplications of a size strikingly similar to those seen in these plants and are strongly dependent on the same stress-sensing pathway the authors studied. Understanding how loss of TSK-like functions tips the balance from stable DNA maintenance to runaway duplication may therefore illuminate both how to engineer new plant traits and how certain tumors evolve and respond to therapy.
Citation: Thomson, G., Poulet, A., Huang, YC. et al. TONSOKU prevents the formation of large tandem duplications and restrains ATR–WEE1 checkpoint activation. Nat Commun 17, 2874 (2026). https://doi.org/10.1038/s41467-026-70906-1
Keywords: genome stability, tandem duplications, DNA replication stress, DNA repair pathways, plant development