Clear Sky Science · en
TONSL suppresses polymerase theta-dependent tandem duplications through chromatin-guided repair
When DNA Makes Dangerous Extra Copies
Every cell must copy its DNA with astonishing accuracy, yet our genomes still face damage that can rearrange chromosomes. One especially important change is the “tandem duplication,” where a stretch of DNA is copied and pasted right next to its original location. Such extra copies can help evolution by providing new gene material, but in cancer they often drive uncontrolled growth. This study uncovers how a protein called TONSL helps cells avoid these risky duplications, revealing a hidden safeguard that appears to be shared across animals and plants. 
Extra Copies: Helpful for Evolution, Risky for Health
Large back-to-back DNA repeats, known as tandem duplications, are among the most common structural changes in complex genomes. They can double entire genes, sometimes enabling new functions to evolve over long time scales. Yet when scientists look at healthy tissues, they find very few fresh tandem duplications, suggesting cells actively prevent them. In many cancers, however, these duplications are abundant, reshaping the genome on a grand scale. The mystery has been how normal cells usually repair DNA breaks without accidentally creating such large copied segments.
Finding a Hidden Genome Guardian
To hunt for natural protectors against these duplication events, the researchers turned to the tiny roundworm Caenorhabditis elegans, which can be bred and sequenced in large numbers. By reanalyzing a collection of about 2,000 mutagenized worm strains, they identified a handful that had unexpectedly high numbers of tandem duplications. A common thread among these strains was damage to a single gene, called tnsl-1, which encodes the protein TONSL. When the team deliberately knocked out this gene and propagated worms for many generations, their genomes accumulated tandem duplications at a steady pace, roughly one per generation, scattered across all chromosomes. Surprisingly, the animals remained largely healthy, revealing that a genome can quietly amass many large duplications without immediately collapsing.
Fast Cells Make Small Duplications, Slow Cells Make Big Ones
Closer inspection showed that the new duplications came in two main size ranges: one cluster around tens of thousands of DNA letters, and another around a few hundred thousand. The team suspected that the tempo of cell division might shape how big a duplication becomes. Worm embryos divide very quickly, with cell cycles lasting about 20 minutes, while germline cells that give rise to eggs and sperm divide much more slowly. Using carefully designed genetic crosses and whole-genome sequencing of offspring, the researchers could time when duplications arose. They found that small duplications formed during rapid early embryonic divisions, whereas large ones originated later in the slowly cycling germline. This pointed to a model in which the length of time available for DNA repair and copying determines how far a break-driven copying process can run before it is sealed off. 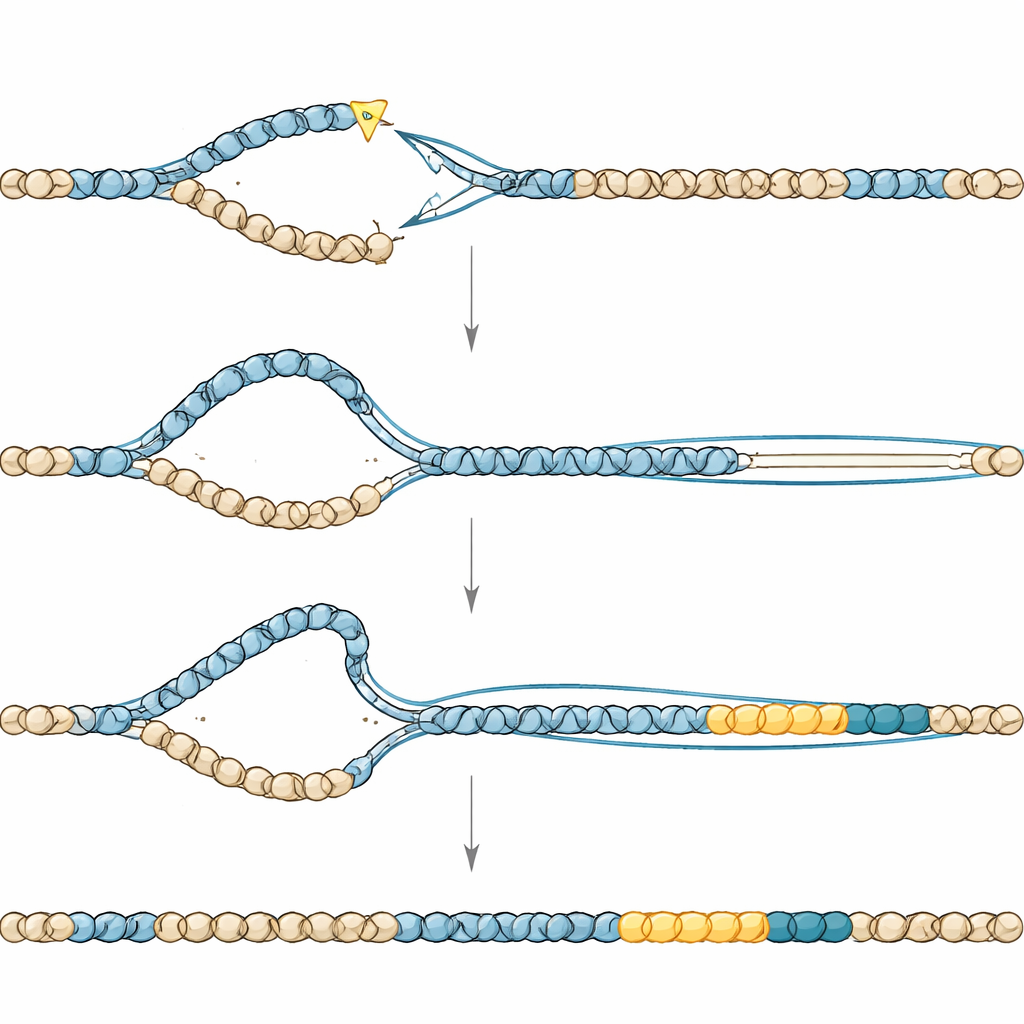
How Faulty Repair Machinery Builds Duplications
The junctions where duplicated segments joined the original DNA bore the fingerprints of a specialized, error-prone repair pathway called polymerase-theta–mediated end joining. When this pathway’s key enzyme was disabled, duplications almost vanished, but the worms paid a heavy price in fertility, implying that this same pathway is also needed to rescue broken chromosomes when TONSL is absent. Other experiments pointed to a second player, a process resembling “break-induced replication,” in which a broken DNA end uses an intact template to copy large stretches of sequence. Removing a helicase that promotes such long-distance copying shrank the size of the duplications without stopping them, bolstering the idea that extended, poorly controlled DNA synthesis underlies these events.
Chromatin as the Quiet Director of Repair
TONSL is known to bind specific chemical marks on newly replicated DNA packaging proteins, suggesting it helps rebuild chromatin—the structure that wraps and organizes DNA—right after copying. The authors engineered a subtle change in the worm protein that weakens this chromatin binding. Worms carrying this single amino-acid swap still formed duplications, but fewer and smaller ones, indicating that TONSL’s grip on fresh chromatin directly restrains runaway DNA synthesis at repair sites. In its absence, repair intermediates can extend too far before finally being stitched together by the error-prone joining pathway, leaving behind a duplicated block instead of a clean fix.
A Shared Defense From Worms to Plants
To test whether this protective role is conserved in other species, the team examined a mutant of the model plant Arabidopsis thaliana lacking the related protein TONSOKU. After only a few generations, these plants accumulated a striking number of large tandem duplications, adding up to roughly a one-percent increase in genome size per generation—a massive change on evolutionary scales. The DNA junctions again carried the characteristic short overlaps and insertions associated with the same error-prone repair pathway, suggesting that worms and plants use a deeply conserved chromatin-guided system to steer replication-associated breaks away from duplication-prone outcomes.
What This Means for Cancer and Genome Stability
In plain terms, this work shows that TONSL and its plant cousin act as quality controllers that sit at the crossroads of DNA copying and repair. By helping to rebuild chromatin properly after a break, they keep repair processes short and tidy, preventing long stretches of DNA from being copied twice in a row. Without this control, cells still repair breaks, but they do so by overextending and then crudely rejoining the DNA, leaving behind tandem duplications. Because similar duplication patterns are seen in several human cancers, including tumors with defects in replication-associated factors, understanding TONSL’s role may eventually help explain why certain cancers acquire such heavily reworked genomes and suggest new ways to predict or influence how their DNA will evolve under therapy.
Citation: van Schendel, R., Romeijn, R., Kralemann, L.E.M. et al. TONSL suppresses polymerase theta-dependent tandem duplications through chromatin-guided repair. Nat Commun 17, 2875 (2026). https://doi.org/10.1038/s41467-026-70905-2
Keywords: genome stability, tandem duplications, DNA repair, chromatin, TONSL