Clear Sky Science · en
Synthetic aptamer mechanoreceptors enable cell-specific force sensing and temporal control via DNA circuits
Listening to Cells Through Gentle Pulls
Our bodies are full of cells that constantly push and pull on their surroundings, using tiny forces to decide when to grow, move, or change. This study describes a new way to eavesdrop on those forces with DNA-based devices that can be tuned to specific cell types and turned on and off over time. Such technology could one day help engineers guide tissue growth, understand how tumors behave, or design smart materials that react to living cells in real time.
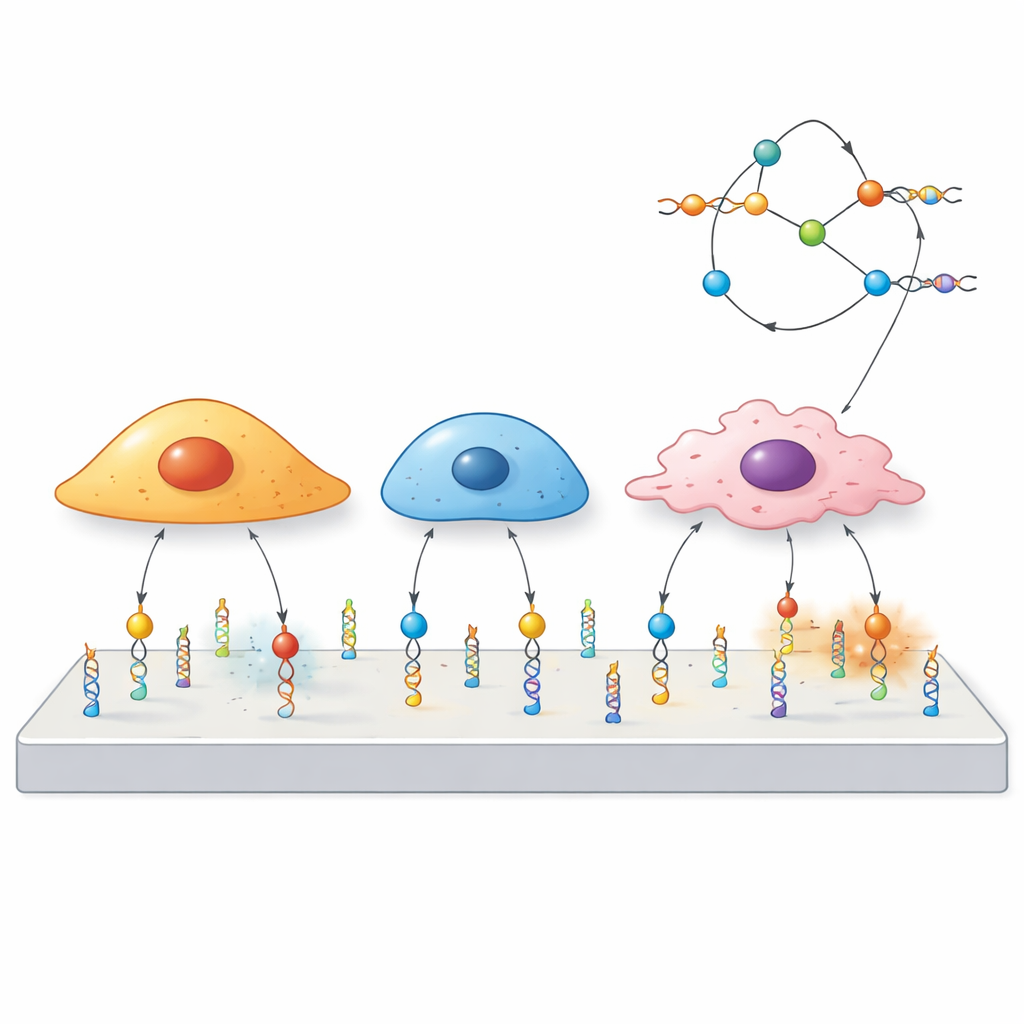
Using DNA as a Tiny Mechanical Switch
The researchers built “mechanoprobes” out of DNA, the same molecule that carries genetic information. Each probe has three parts: a short DNA sequence called an aptamer that sticks to a chosen protein on the cell surface, a paired DNA segment that acts like a spring tuned to break at a certain force, and a fluorescent dye that lights up when that spring snaps. When a cell pulls on the receptor bound by the aptamer strongly enough, the DNA duplex opens and the signal turns on. Because different aptamers latch onto different receptors, the team can decide in advance which cells or proteins will be able to trigger the probe.
Finding Hidden Force Pathways on Cell Surfaces
Most existing force sensors focus on integrins, well-known proteins that help cells grip their surroundings but are found in many cell types, limiting selectivity. Here, the team instead targeted “noncanonical” receptors that are not usually thought of as mechanical. They showed that an aptamer called AS1411, which binds a protein named nucleolin common on cancer cells, only produced signal when it could specifically recognize its target. By designing probes that required either weaker or stronger forces to open, they discovered that nucleolin transmits modest forces compared to integrins and does not by itself help cells spread on a surface. In contrast, another aptamer, Sgc8, which recognizes the receptor PTK7, could report forces even when cells barely adhered, forming ring-like patterns that hinted at a different source of motion.
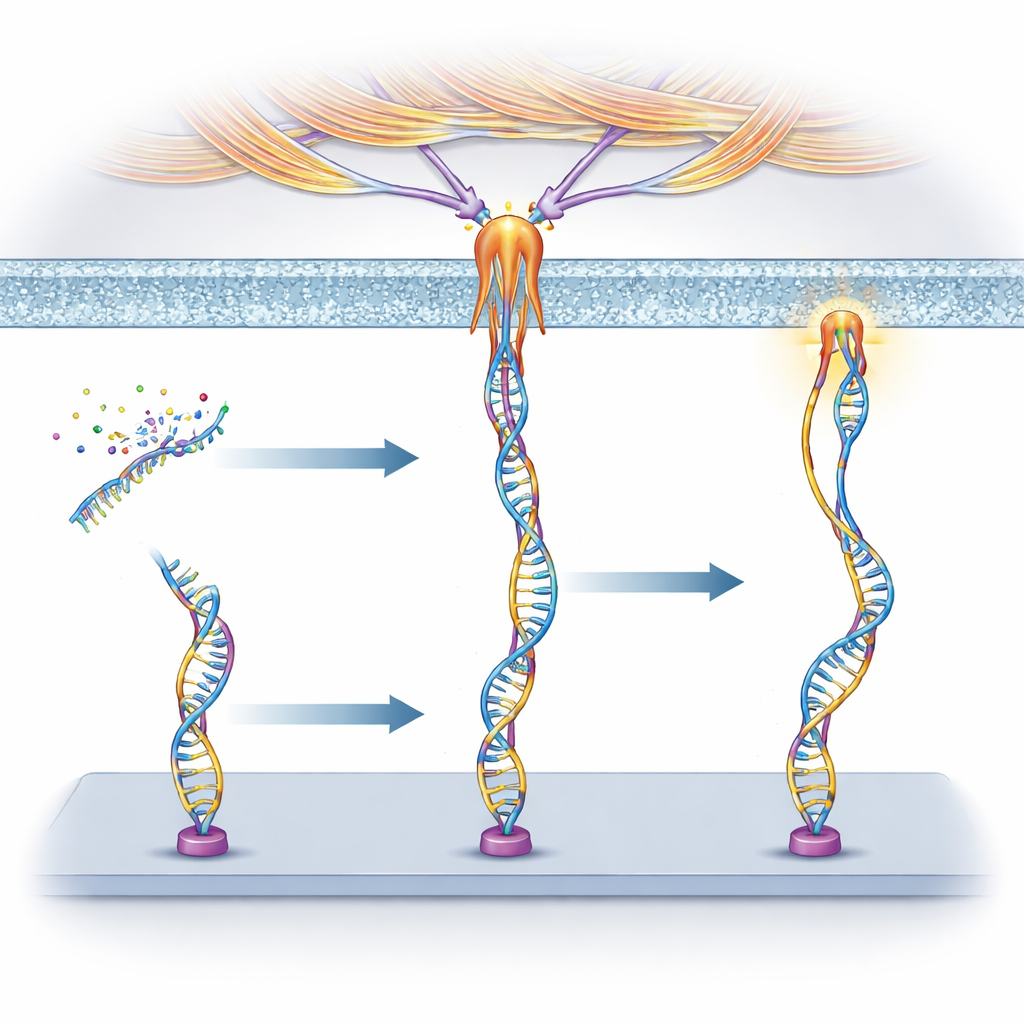
Tracing Where the Forces Come From
To understand what was pulling on these DNA devices, the researchers used drugs that selectively block different cell processes. For nucleolin, they found that its forces depended strongly on myosin, the same motor protein that helps muscles contract, and appeared at sites where the cell forms focal adhesions—special contact points used for gripping and sensing stiffness. This suggests that nucleolin is indirectly linked to the cell’s internal actin–myosin machinery and can share its contractile forces. For PTK7, however, blocking a process called macropinocytosis—where cells ruffle their membranes and gulp in fluid—greatly reduced the signals. The forces arose mainly from early membrane ruffling driven by actin growth, not from later steps in vesicle sealing, revealing a distinct mechanical pathway.
Dialing in Cell Selectivity with Molecular Precision
Because each aptamer recognizes a particular surface receptor, the same DNA scaffold can be reprogrammed simply by swapping one aptamer for another. The team demonstrated this with probes for three different targets—PTK7, mucin-1, and EpCAM—across a panel of cancer cell lines with high or low levels of each protein. Cells rich in a given receptor produced strong signals, while those with little receptor barely responded, even when mixed together in the same dish. Strikingly, cells that shared the same receptor could still show different spatial patterns of force, underlining that force flow through a receptor depends not only on its presence but also on each cell type’s internal wiring.
Programming When Cells Are Allowed to Pull
Taking advantage of the fact that aptamers are made of DNA, the researchers connected their probes to DNA reaction networks that control whether the aptamer can bind at all. Complementary “blocker” strands can temporarily hide the aptamer, and “activator” strands can later remove the blockers through a strand-exchange reaction, restoring force sensitivity. They also designed an RNA-based blocker that can be cut by an enzyme called RNase H, and further slowed this cutting with decoy molecules that compete for the enzyme. In this way, they turned mechano-sensing into a timed program: cells could only generate signals after a built-in delay or within chosen time windows.
Why This Matters for Future Living Materials
In everyday terms, this work turns DNA into a smart mechanical lock that opens only for certain cells, only under certain forces, and only at chosen times. By revealing that less obvious receptors like nucleolin and PTK7 can carry mechanical information through very different routes inside cells, it broadens our picture of how cells “feel” their environment. Because the entire system is built from nucleic acids, it can plug directly into the fast-growing toolbox of DNA circuits and nanodevices. This creates a foundation for materials and engineered tissues that not only sense when cells push or pull, but also respond with carefully timed biochemical actions, potentially guiding healing, growth, or cancer treatment in a highly programmable way.
Citation: Xu, T., Sethi, S., Drees, C. et al. Synthetic aptamer mechanoreceptors enable cell-specific force sensing and temporal control via DNA circuits. Nat Commun 17, 2492 (2026). https://doi.org/10.1038/s41467-026-70765-w
Keywords: mechanotransduction, DNA aptamers, cell mechanics, synthetic biology, DNA reaction networks