Clear Sky Science · en
A metrological foundation for absolute transcriptomics using International System of Units-anchored calibrators
Why turning RNA signals into real numbers matters
Modern gene tests can read out which genes are turned on or off in our cells, but they stumble on a basic question: how many molecules are actually there? Today’s RNA sequencing technologies mostly compare relative changes between samples rather than giving hard, trustworthy counts. That is a problem if you want to set universal disease thresholds, compare results across hospitals, or build precise models of how cells work. This study introduces a new way to anchor RNA sequencing to the same international units used in chemistry and physics, transforming fuzzy relative signals into absolute, comparable numbers.
The trouble with comparing gene activity
RNA sequencing works by breaking RNA molecules into fragments and counting how many times each gene is represented. But two types of distortion creep in. First, systemic differences between experiments – such as different laboratories, machines, or sample preparation methods – create “batch effects” that make the same sample look different when run twice. Second, sequence-dependent effects – where genes with certain lengths or base compositions are more or less likely to be captured – mean that even within a single sample, some genes are consistently overcounted and others undercounted. As a result, scientists are largely forced to talk about fold-changes between conditions rather than true molecule counts, and these fold-changes themselves can be misleading from batch to batch.
A new set of yardsticks for RNA measurements
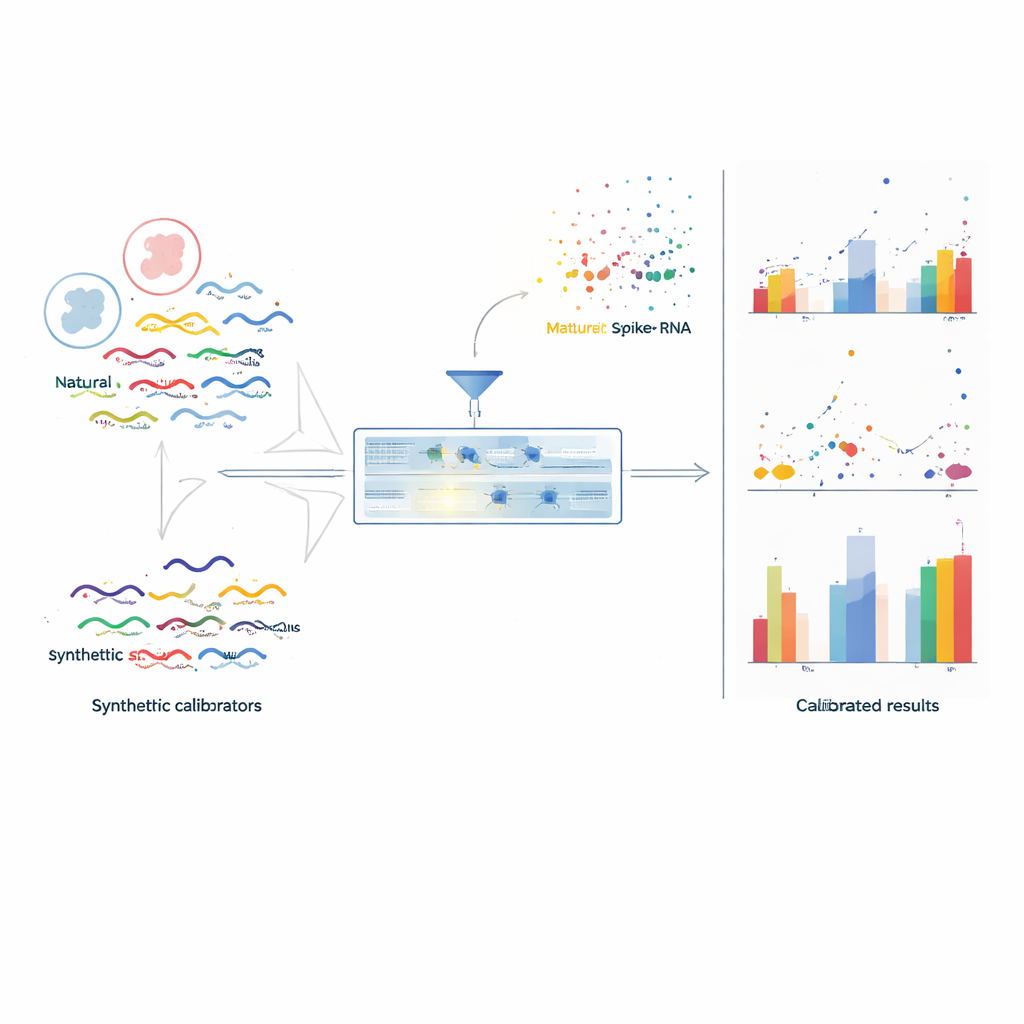
To fix this, the authors created TranScale, a panel of 100 synthetic RNA molecules designed to behave like real human transcripts while remaining computationally distinct. These standards span a wide range of lengths, sequence features, and clinically relevant variants such as splice forms and gene fusions, closely echoing the diversity of actual cellular RNA. Crucially, each TranScale molecule is assigned an exact concentration using a primary measurement technique called isotope dilution mass spectrometry, which is traceable to the International System of Units (SI). By mixing a known, tiny amount of TranScale into every RNA sample before sequencing, the experiment gains an internal ruler that experiences the same laboratory steps and distortions as the natural RNAs.
Turning noisy reads into absolute counts
With TranScale present in each library, the team can compare the number of sequencing reads for each spike-in molecule to its certified concentration. For each batch, they select well-behaved spike-ins and fit a straight-line calibration curve linking read-based units to true molecule counts. This simple model simultaneously captures both batch-wide and sequence-related biases. The same curve is then applied to all genes in the sample, converting their relative readouts into absolute copy numbers per unit of RNA. In a large multi-lab, multi-platform study deliberately designed to produce strong batch effects, this calibration cut the median variation of absolute measurements across centers from above 85% to below 15–25%, and restored the correct clustering of biological samples that had been obscured by technical noise.
Seeing hidden errors and fixing them
The TranScale standards also act as diagnostic probes of data quality. By comparing measured values with their certified truths, the authors separated two kinds of error: how wrong each gene’s absolute level is, and how wrong the ratios between conditions are. They found surprising examples where relative differences looked consistent but absolute numbers were badly distorted, and vice versa. This means conventional checks that focus only on fold-changes can miss serious problems. After calibration, both absolute levels and ratios of spike-ins and thousands of real human genes matched closely with independent digital PCR measurements and an external reference dataset. The corrected data revealed a much clearer quantitative landscape, making it possible to compare housekeeping genes to cancer drivers on the same absolute scale and to link DNA changes, such as co-amplified cancer genes, directly to their RNA outputs.
From relative trends to clinical thresholds
Finally, the researchers showed how absolute scaling can sharpen medical decisions. Using an oncogene often measured in breast cancer, they defined a fixed cutoff based on digital PCR and asked whether RNA sequencing could reliably call samples as normal or tumor across many batches. Uncorrected data gave inconsistent answers because of batch effects. After TranScale calibration, every library agreed with the true classification. By tying RNA sequencing to SI units through biomimetic standards, this work lays a metrological foundation for transcriptomics. It opens the door to universal diagnostic cutoffs, robust data sharing between centers, and more precise, system-level models of how genes are expressed in health and disease.
Citation: Zhang, Y., Yang, B., Yu, Y. et al. A metrological foundation for absolute transcriptomics using International System of Units-anchored calibrators. Nat Commun 17, 2747 (2026). https://doi.org/10.1038/s41467-026-70582-1
Keywords: RNA sequencing, absolute quantification, metrology, gene expression calibration, biomolecular standards