Clear Sky Science · en
Transient molecular chimerism for exploiting xenogeneic organelles
Borrowed Solar Panels in a Single Cell
We usually think of solar-powered life in terms of plants and algae, but some single-celled predators take a shortcut: they steal the "solar panels"—chloroplasts—from their prey. This study explores how one such microbe, Rapaza viridis, keeps these stolen parts running for weeks, even though they come from a completely different species. By uncovering how the host’s own proteins slip into and support these foreign chloroplasts, the work sheds light on how complex cell parts like chloroplasts may have first evolved long ago.
How a Microbe Steals and Uses Green Power
Rapaza viridis is a tiny flagellate that eats a specific green alga called Tetraselmis. Instead of digesting everything, Rapaza keeps the captured chloroplasts, now called “kleptoplasts,” and throws away the rest of the algal cell, including its nucleus. These kleptoplasts are sliced into pieces and passed on to daughter Rapaza cells. For about two weeks, the host can live almost entirely on the energy and carbon made by these borrowed chloroplasts, even though they no longer receive instructions or replacement parts from their original algal genome. This unusual lifestyle offers a live window into what early steps toward permanent chloroplasts might have looked like.
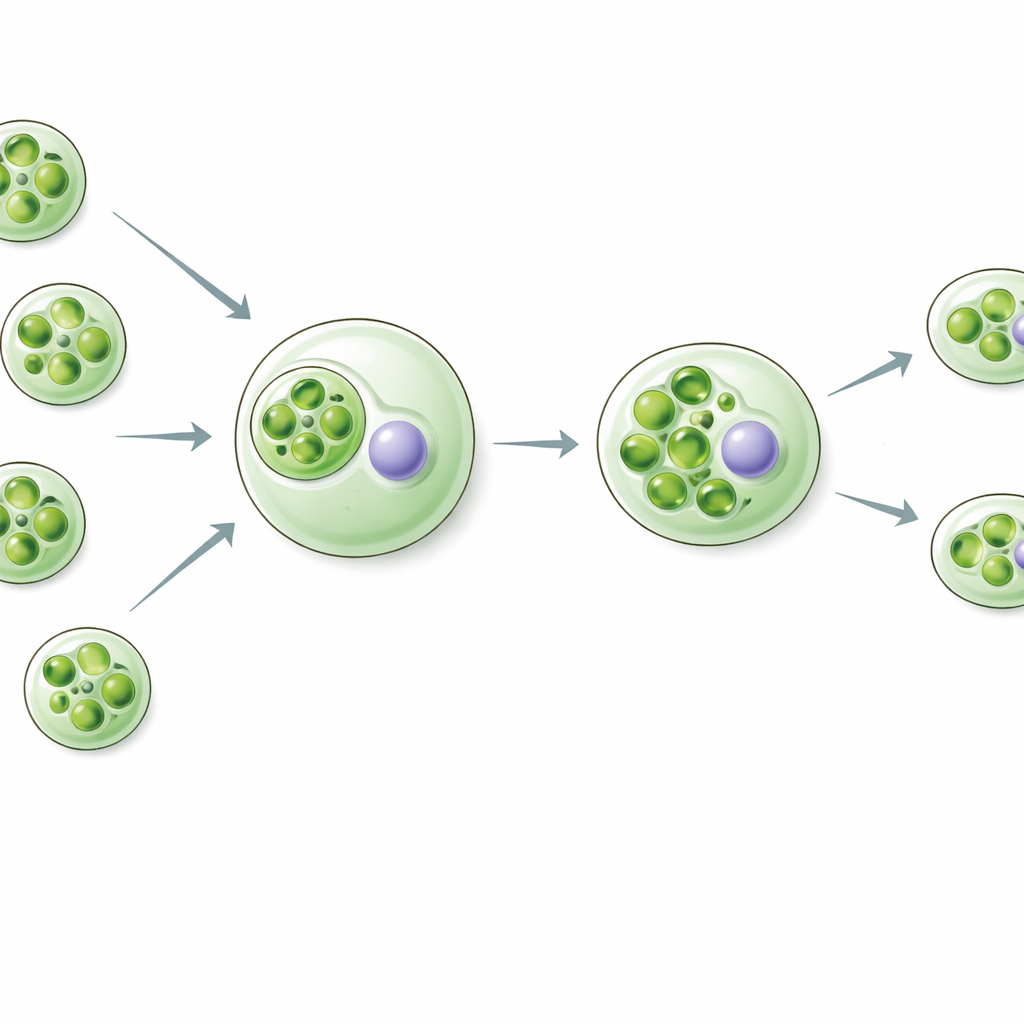
Host Genes Step In to Keep Stolen Parts Alive
The researchers asked a crucial question: if the algal nucleus is gone, who supplies the proteins needed to keep the kleptoplast running? By analyzing gene activity over time in Rapaza, they identified 37 host genes whose products look like they work inside chloroplasts. Many of these genes resemble components of the photosynthetic machinery—light-harvesting proteins, electron transport parts, and enzymes that fix carbon. Two stood out: a RuBisCO small-subunit–like protein (RvRbcS-like) and a RuBisCO activase–like protein (RvRca-like). Both are related to key helpers of RuBisCO, the central enzyme that captures carbon dioxide. These host genes switch on strongly after the prey is eaten, just when the kleptoplasts are being reshaped and prepared for long-term use.
Proving the Host Proteins Enter the Kleptoplasts
Finding suggestive gene sequences is not enough; the proteins must actually reach the kleptoplast interior. Using custom antibodies and fluorescent microscopy, the team tracked where RvRbcS-like and RvRca-like proteins accumulate. They engineered Rapaza strains with tiny detectable tags on RvRbcS-like and showed that the signal overlapped with the kleptoplasts and with RuBisCO itself. Similar imaging with an antibody against RvRca-like revealed that this protein also concentrates inside kleptoplasts. On protein gels, both proteins appeared shortened, consistent with the removal of a special front-end segment as they cross membranes—just like targeting signals that guide proteins into ordinary chloroplasts.
Why These Borrowed Helpers Matter
To test whether these host proteins are truly important, the authors used CRISPR-based genome editing to knock out each gene. Cells lacking RvRbcS-like grew poorly, lost photosynthetic strength, made far fewer energy-rich storage grains, and died much earlier than normal. Levels of the algal RuBisCO large subunit also fell, suggesting that without the host’s replacement small subunit, the enzyme complex falls apart. Removing RvRca-like had a milder but still clear effect: early growth remained close to normal, but later photosynthesis and carbon storage declined, revealing a gradual loss of efficiency. Together, these results show that Rapaza does not simply park stolen chloroplasts; it actively maintains and remodels them with its own protein toolkit.
Custom Delivery Tags and Rebuilt Inner Structures
Most of the putative kleptoplast-targeted proteins in Rapaza share long, low-structure “heads” at their front end, often with predicted membrane-spanning segments. By fusing one of these heads to a luciferase reporter protein, the authors showed that this segment alone is enough to carry a cargo protein into kleptoplasts. Detailed sequence analysis revealed several classes of these targeting regions, which closely resemble those used by a related group of algae to import proteins into their permanent, triple-membrane chloroplasts. One especially striking host protein, RvRbcS-like, carries four RuBisCO-related domains and a floppy tail rich in repeating motifs thought to promote phase separation of proteins. The authors propose that this tail helps reorganize the pyrenoid—the dense carbon-fixing hub inside the kleptoplast—into multiple droplets that can be inherited by daughter cells.
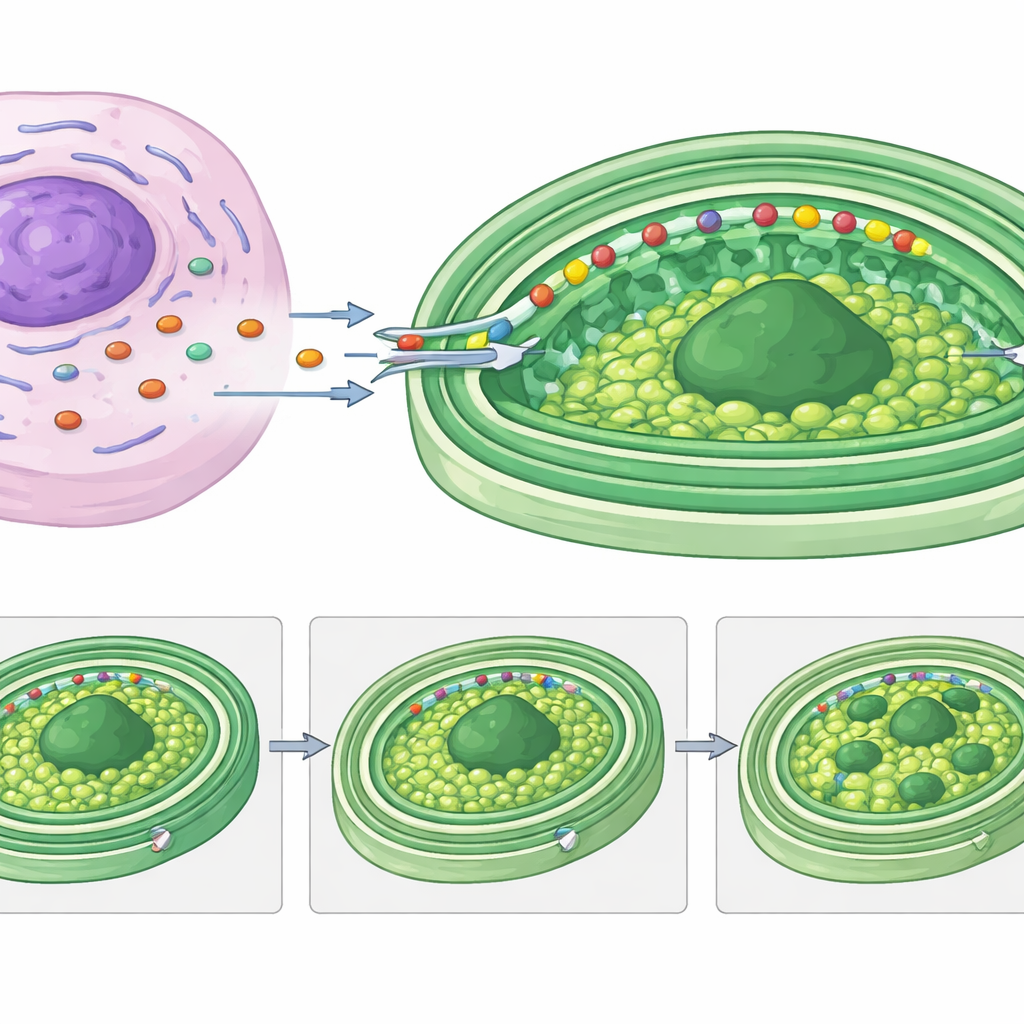
What This Means for the Story of Complex Cells
To a non-specialist, the key message is that Rapaza viridis shows a living, reversible version of what might have happened when ancient cells first turned free-living bacteria into permanent parts like chloroplasts. Here, the host rapidly builds a temporary molecular partnership: its own genes supply crucial replacement parts and even reshape the inner structure of the stolen chloroplasts, all guided by specialized delivery tags. This “transient molecular chimerism” demonstrates that even short-lived integrations between species can be sophisticated and finely tuned. Studying this system gives scientists a powerful model for uncovering how complex cells learned to control, feed, and safely use foreign energy factories—and how new organelles may still emerge over evolutionary time.
Citation: Kashiyama, Y., Maruyama, M., Nakazawa, M. et al. Transient molecular chimerism for exploiting xenogeneic organelles. Nat Commun 17, 2371 (2026). https://doi.org/10.1038/s41467-026-70516-x
Keywords: kleptoplasty, chloroplast evolution, Rapaza viridis, endosymbiosis, organelle biogenesis