Clear Sky Science · en
Label-free mass and size characterization of few-kDa biomolecules by hierarchical vision transformer augmented nanofluidic scattering microscopy
Seeing the Smallest Molecules
Many of the body’s most important messengers—such as hormones and immune signals—are so tiny that they have been nearly invisible to powerful microscopes unless they are tagged with fluorescent labels. Those labels can change how molecules behave, a serious drawback for drug development and basic biology. This study introduces a way to weigh and size individual molecules as small as insulin without any labels, using a combination of nanofluidic chips and advanced artificial intelligence. It opens a path to watching key biomolecules in their natural state, one by one.
Tiny Channels as Test Tubes
At the heart of the method is nanofluidic scattering microscopy, which uses a fingernail-sized chip etched with extremely narrow channels. These channels are only a few dozen nanometers across—thousands of times thinner than a human hair. Molecules dissolved in a buffer drift freely through these channels rather than being stuck to a surface. When the chip is illuminated with visible light, both the channel walls and any passing molecule scatter light. Because the channel is so much larger than the molecule, their scattered light interferes in a way that greatly amplifies the molecule’s optical signature. By subtracting images of an empty channel from images taken when a molecule is present, the researchers obtain a movie-like record (a kymograph) of its passage through the channel, containing information about both its mass and how quickly it diffuses.
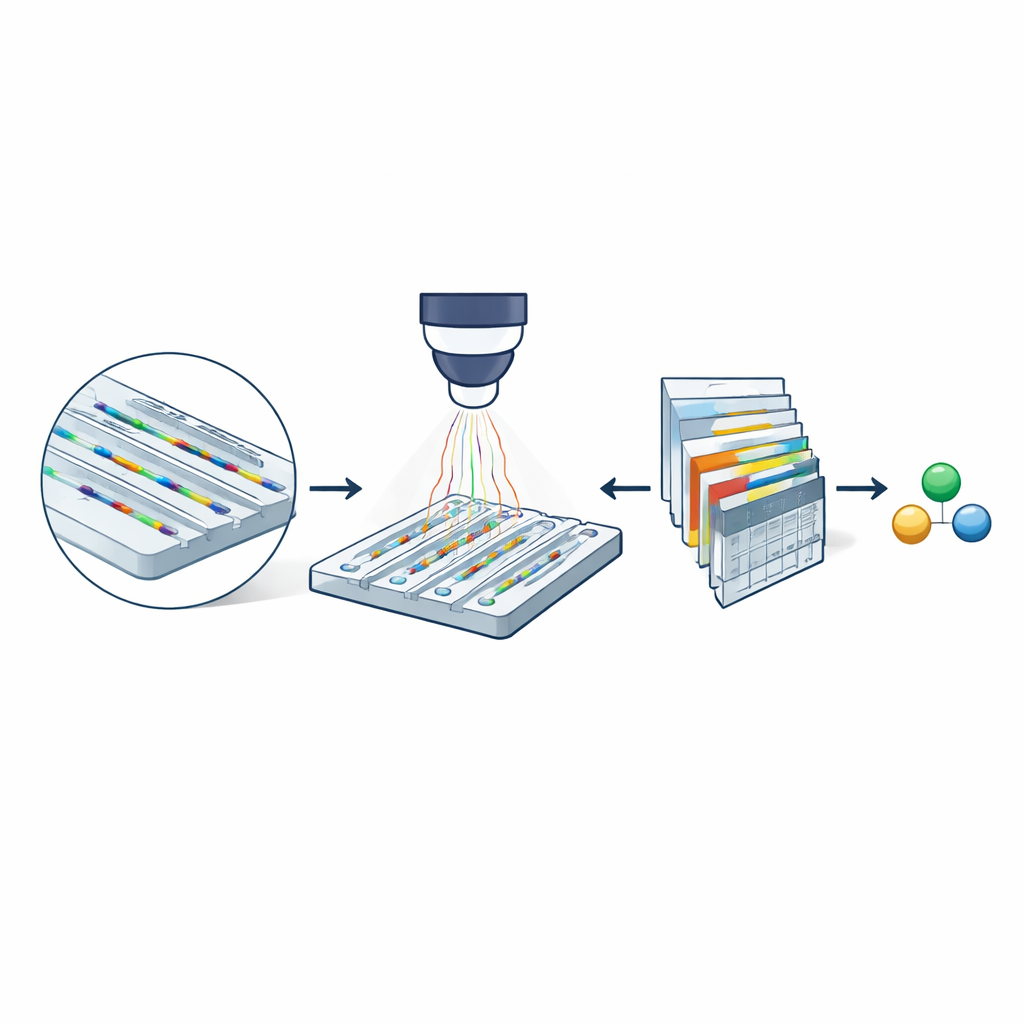
Why Smaller Channels Matter
Even with this optical boost, earlier versions of the technique could only characterize molecules heavier than about 60 kilodaltons—typical large proteins—because signals from smaller species drowned in noise. The team shows that shrinking the cross-section of the nanochannels dramatically improves sensitivity: the optical contrast from a molecule increases as the channel area decreases. Experiments comparing two channel sizes with the protein bovine serum albumin reveal that its path is clearly visible in the narrower channel but nearly lost in the wider one. This simple geometric change hints that, in principle, much smaller biomolecules might be detectable if their faint signals can be reliably extracted from noisy image sequences.
Letting an AI Read Noisy Movies
To reach this regime, the researchers developed a specialized deep-learning model called a hierarchical vision transformer. Instead of trying to reconstruct each molecule’s exact track pixel by pixel, the model scans the kymograph at multiple scales and produces two products: a probability map that marks where a molecule is likely to be at each point in space and time, and a property map that encodes estimates of molecular weight and hydrodynamic size. By weighting property estimates with the probability map, the model can ignore regions that are mostly noise. Training relies entirely on simulated trajectories laid over experimentally measured background noise, allowing the system to learn how real signals should look even when they are barely visible to the eye.
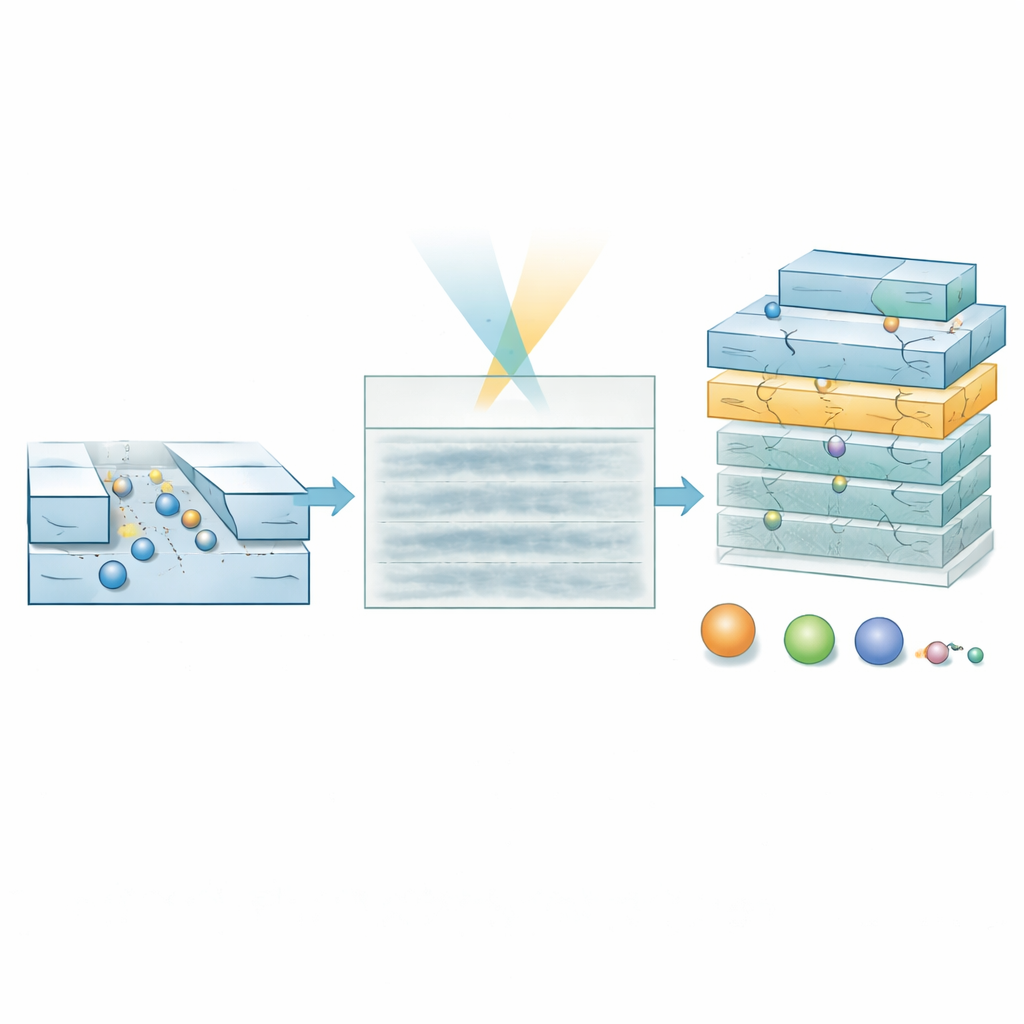
Testing on DNA and Hormone Molecules
The authors first validated the approach using a standard DNA “ladder,” a mix of double-stranded fragments of known lengths and masses, in relatively wide channels where all fragments are deliberately below the usual detection limit. In the raw movies, no clear DNA tracks can be seen. Yet the AI model produces probability maps that correctly highlight where molecules pass and predicts masses that match the expected values once low-confidence data are filtered out. The method also correctly reports smaller effective sizes for the stiff, rod-like DNA fragments than their full physical length, reflecting how their shape affects diffusion. The most demanding test used ultrasmall channels to study the peptide hormone insulin, which weighs only about 5.8 kilodaltons and is roughly 1.5 nanometers across. Here too, the kymographs look featureless, but the model extracts a tight cluster of mass and size values in excellent agreement with literature numbers, cleanly separated from buffer-only controls.
Pushing Against the Limits
To understand how far this strategy can go, the team compared the model’s precision to the Cramér–Rao lower bound, a statistical limit that defines the best possible accuracy for any unbiased estimator given the noise in the data. Using large sets of simulations, they show that as the number of frames contributing to a molecule’s trajectory increases beyond about ten thousand, the model’s estimates of mass and size approach this theoretical optimum even for a six-kilodalton particle. Experiments on insulin at different concentrations reproduce the same trend: longer effective trajectories yield more precise measurements, limited mainly by how long tiny, fast-diffusing molecules remain in view. The authors propose future strategies—such as gently trapping molecules in the channels or reusing data with bootstrapping—to further increase usable observation time.
What This Means for Biology and Medicine
In everyday terms, this work shows that it is now possible to weigh and size some of the body’s smallest and most important molecules individually, without touching them with fluorescent tags or sticking them to a surface. By marrying nanofabricated channels with a purpose-built AI that reads noisy light-scattering movies, the method lowers the mass detection limit of this label-free microscopy approach by roughly a factor of ten. This opens the door to studying families of small signaling proteins and peptide hormones—such as cytokines, chemokines, and insulin—at the single-molecule level, and to characterizing tiny biological nanoparticles and drug carriers with unprecedented sensitivity. As the technology matures, it could become a powerful tool in drug discovery, diagnostics, and fundamental studies of how molecular interactions shape life at the smallest scales.
Citation: K. Moberg, H., Yeroshenko, B., Fritzsche, J. et al. Label-free mass and size characterization of few-kDa biomolecules by hierarchical vision transformer augmented nanofluidic scattering microscopy. Nat Commun 17, 2533 (2026). https://doi.org/10.1038/s41467-026-70514-z
Keywords: label-free single-molecule microscopy, nanofluidic scattering, vision transformer, small biomolecules, insulin detection