Clear Sky Science · en
A genus-wide interaction atlas across NS4B orthologues identifies a conserved role for UFMylation in orthoflavivirus replication
Why this virus study matters
Viruses like dengue, Zika and West Nile are spreading into new regions and can cause anything from fever to brain infections and birth defects. Yet we still lack broad, reliable treatments. This study explores how a hidden partnership between these viruses and a little‑known cellular tagging system called UFMylation helps the pathogens multiply. By mapping this relationship across many related viruses, the authors uncover a shared weak spot that might be exploited to develop future antiviral drugs.
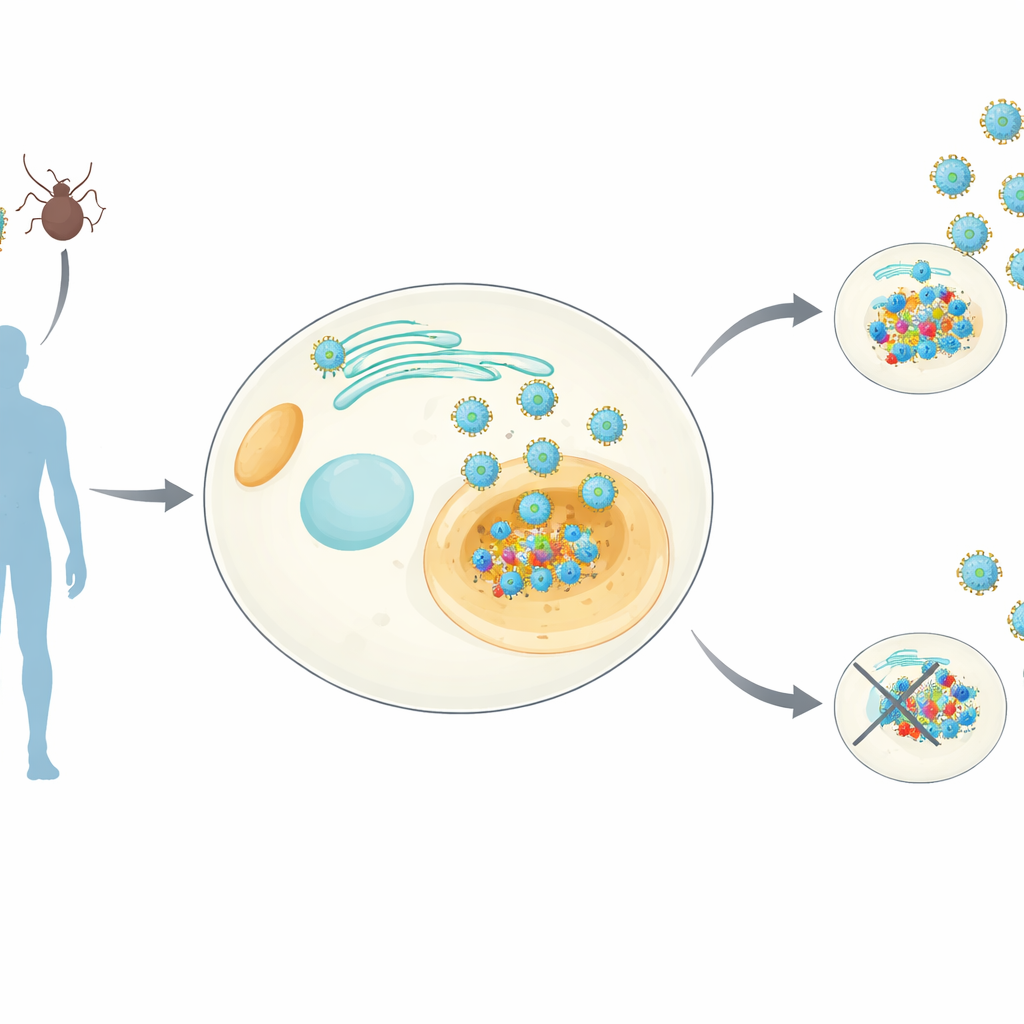
Following a key viral insider
Orthoflaviviruses – the group that includes dengue, Zika, yellow fever and others – all carry a tiny protein called NS4B that nestles into membranes inside our cells. NS4B is known to be essential for viral replication, but its exact jobs have remained murky. The researchers created human cell lines that stably produce NS4B from eight different orthoflaviviruses. They then used sensitive mass‑spectrometry techniques to pull down NS4B along with any human proteins stuck to it, and in parallel measured how NS4B expression alters the overall mix of proteins in the cell. This produced a genus‑wide “interaction atlas” – which they call the NS4Bome – showing both common and virus‑specific host targets.
Shared tricks and custom hacks
The atlas revealed 538 human proteins that interact with NS4B and over 500 whose abundance changes when NS4B is present. Many of these are concentrated in pathways that manage energy production in mitochondria, the shaping of internal membranes and the movement of newly made proteins and fats. Some interactions were shared by most viruses, hinting at evolutionarily conserved tricks that all orthoflaviviruses use. Others appeared only in certain species or strains, potentially explaining why some viruses prefer mosquitoes over ticks, or cause brain disease instead of hemorrhagic fever. This systematic view turns a tangled set of previous, piecemeal studies into a coherent map of how these viruses rewire the cell.
A surprising ally: the UFMylation system
Among the many partners that NS4B latched onto, one stood out: UBA5, the starter enzyme of the UFMylation pathway. UFMylation is a cellular system that attaches a small protein tag, UFM1, to selected targets, influencing processes such as protein quality control and stress responses. Using RNA interference and CRISPR‑based gene knockouts, the team showed that cells lacking UBA5 or otherwise unable to perform UFMylation had greatly reduced replication of Zika and several related viruses. Reintroducing normal UBA5 restored virus growth, whereas mutant forms that could no longer drive UFMylation did not, proving that the tagging activity itself is crucial for infection.
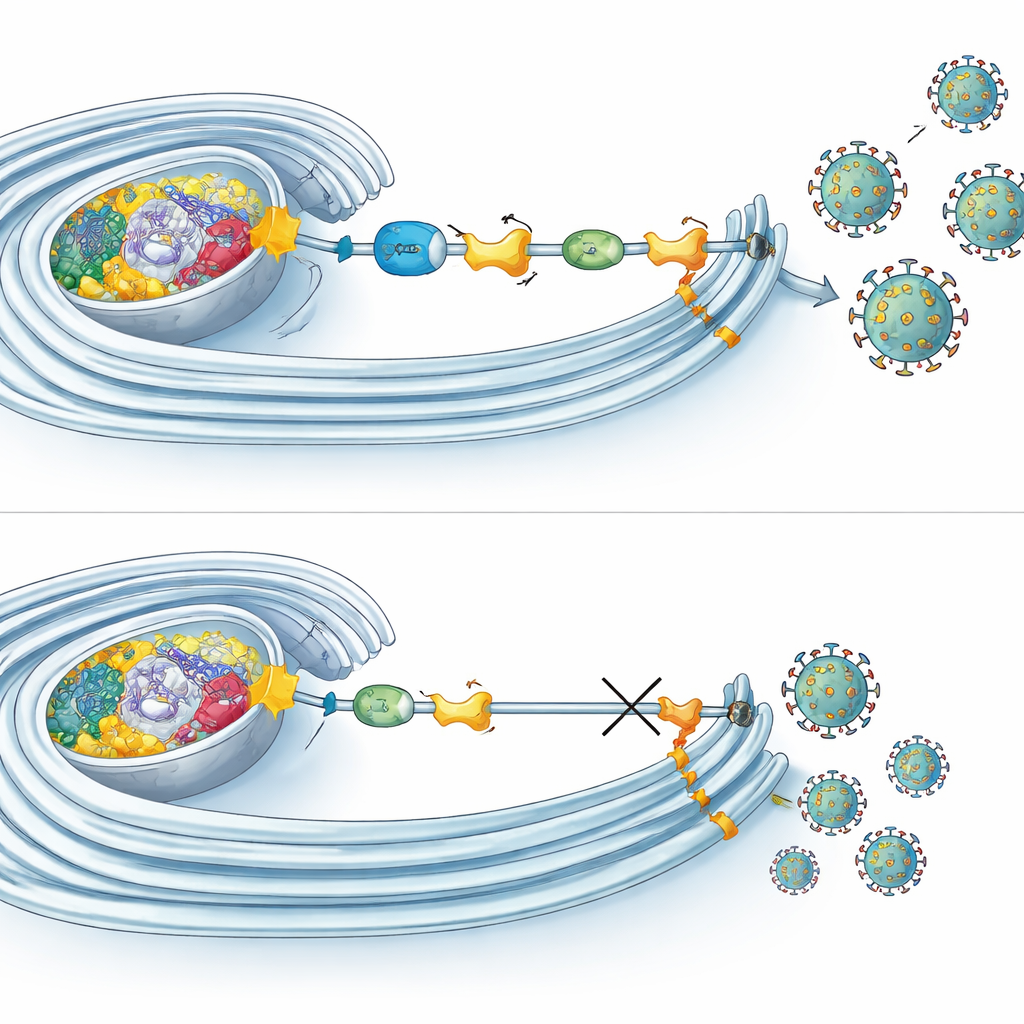
How tagging supports viral factories
Microscopy and biochemical experiments revealed that multiple UFMylation components gather at the very sites where viral RNA is copied inside remodeled membranes. There, they associate not only with NS4B but also with other viral proteins that build the replication machinery. Interestingly, blocking UFMylation did not stop viral entry, RNA production or protein synthesis. Instead, it mainly crippled the late stages of the cycle, such as assembly or release of infectious particles. At the same time, UFMylation turned out to influence how mitochondria breathe: when the pathway was disabled, cells showed reduced oxygen consumption and altered mitochondrial shapes. In contrast, early during Zika infection, mitochondrial respiration ramped up, suggesting that viruses may tap into UFMylation to boost the cell’s energy output right when they need it most.
Testing a drug in cells and tiny fish
The authors then turned to a small‑molecule compound, DKM 2‑93, that blocks the active site of UBA5. In cultured human cells, this inhibitor cut Zika virus production roughly ten‑fold at doses that left cells largely healthy, and showed activity across several cell types. To see whether this strategy could work in a living organism, they used a zebrafish model of Zika infection, which mimics key features of disease in the developing brain. Treating infected embryos with DKM 2‑93 during early brain development significantly lowered viral RNA levels and reduced the proportion of fish showing severe malformations, indicating genuine antiviral benefit in vivo.
What this means for future treatments
By building a comprehensive map of how NS4B from many orthoflaviviruses interacts with human proteins, this study highlights UFMylation as a conserved helper pathway that these viruses depend on to produce infectious particles. Because the same host system appears to support dengue, Zika, West Nile and others, drugs that selectively dampen UFMylation might act as broad‑spectrum antivirals, rather than targeting one virus at a time. While more work is needed to understand exactly which proteins are tagged and to ensure safety, the findings identify a promising host‑based target that could one day help blunt outbreaks of multiple mosquito‑ and tick‑borne diseases with a single therapeutic approach.
Citation: Rajasekharan, S., Barragan Torres, V.A., Pinheiro Gomes, Y.C. et al. A genus-wide interaction atlas across NS4B orthologues identifies a conserved role for UFMylation in orthoflavivirus replication. Nat Commun 17, 2489 (2026). https://doi.org/10.1038/s41467-026-70437-9
Keywords: Zika virus, dengue, host–virus interactions, UFMylation, broad-spectrum antivirals