Clear Sky Science · en
Identification of cis-regulatory elements provides insights into tissue-specific gene regulation in the sheep genome
Why the sheep genome matters to everyday life
Sheep feed millions of people and power rural economies, yet we still know surprisingly little about how their genes are switched on and off in different parts of the body. This study creates a detailed map of the control switches that tune gene activity in 24 tissues of a single sheep, from brain and lung to muscle and udder. By charting these hidden switches, the work lays the groundwork for healthier animals, better production traits like milk fat, and deeper understanding of how mammalian bodies, including our own, regulate genes.
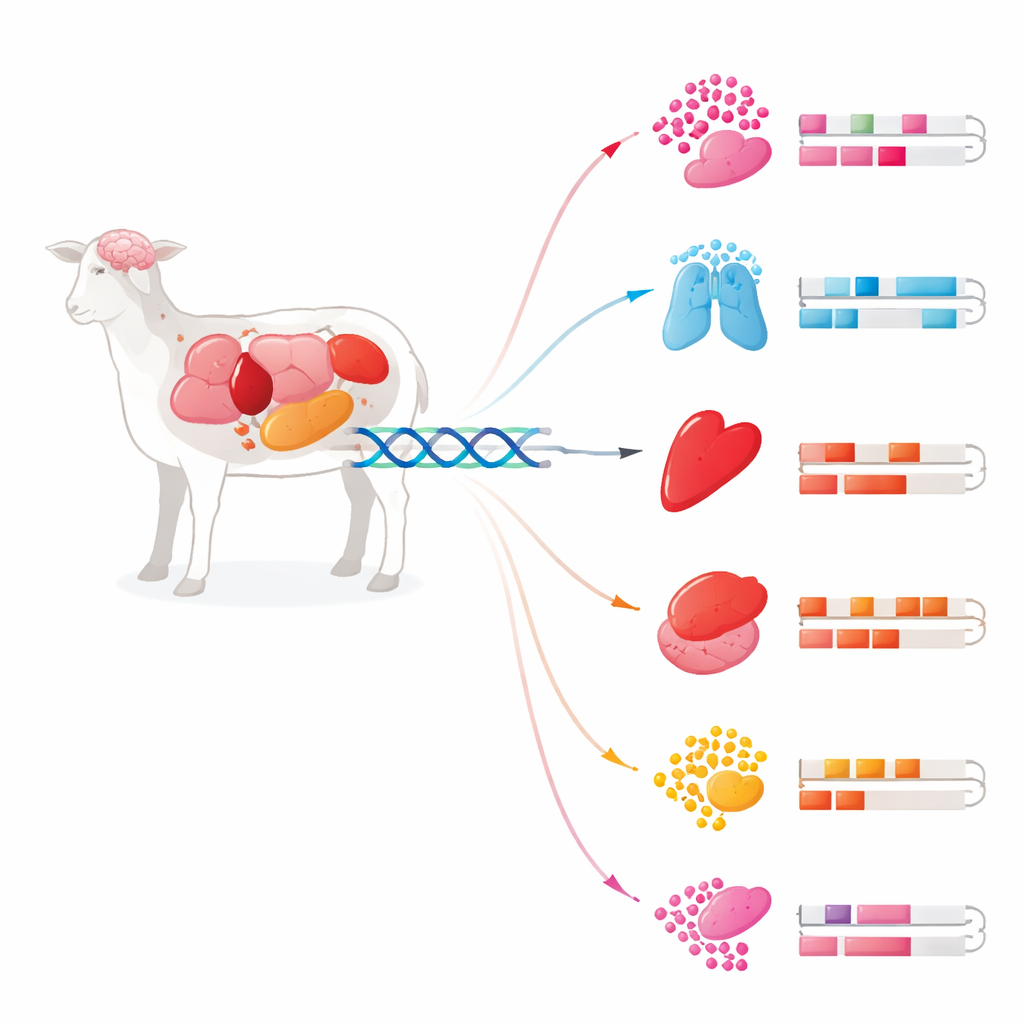
Hidden switches in DNA
Our DNA does not just contain genes; it also holds vast stretches of regulatory code that act like dimmer knobs, deciding when and where genes turn on. Two key types of elements are promoters, right next to genes, and enhancers, which can lie far away but still control them. The researchers combined six cutting-edge methods that read different aspects of genome activity: protein marks on DNA-packaging proteins, open chromatin, transcription start sites, DNA methylation, and RNA output. Using these overlapping clues, they pinpointed more than 270,000 enhancers and nearly 26,000 promoters across 24 tissues from the same Rambouillet ewe used to build the current sheep reference genome. This unified map shows where the genome is wired for regulation, not just which genes exist.
Different tissues, different control logic
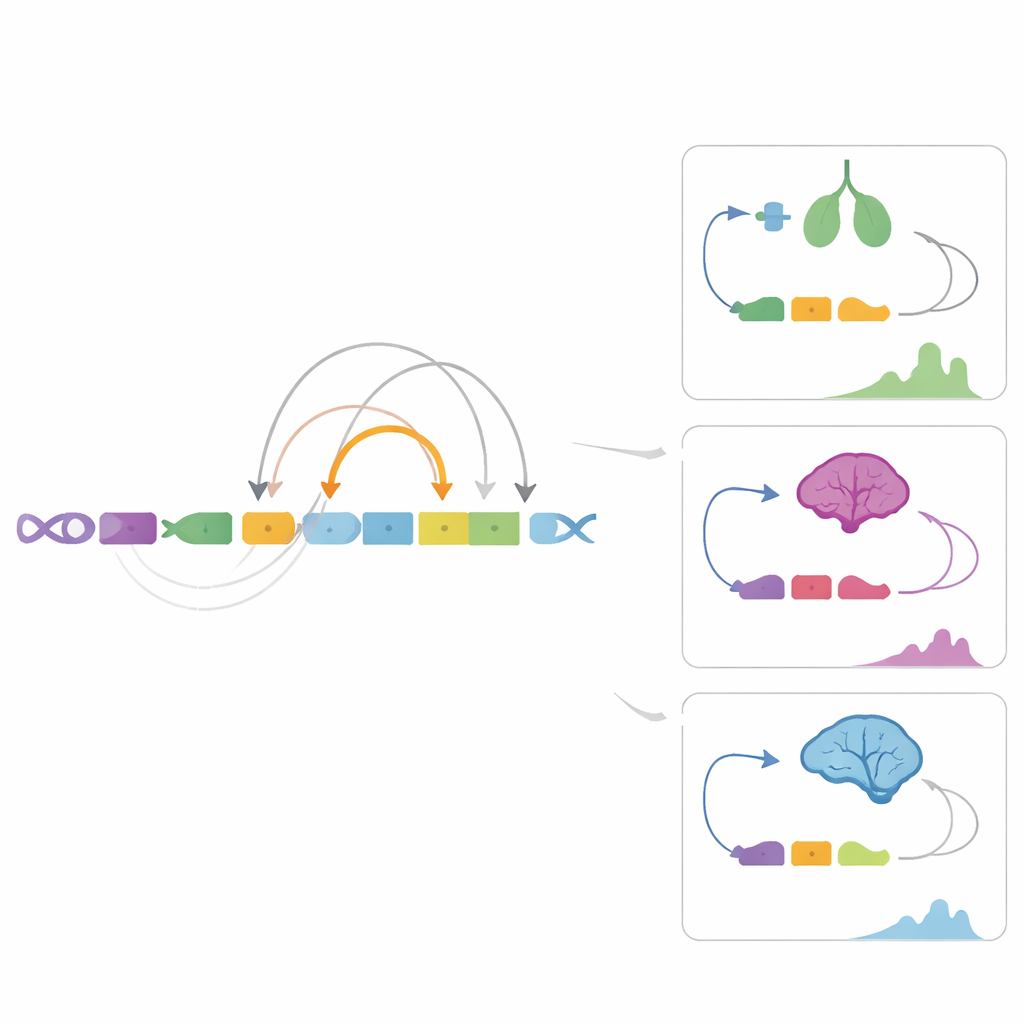
Although almost every cell carries the same DNA, tissues behave very differently because they use distinct sets of enhancers. The study found that promoters are relatively stable: many are shared across tissues, and their patterns are conserved across species. Enhancers, in contrast, are highly variable and far more tissue-specific. Brain tissues, such as the cerebellum and cerebral cortex, stood out with especially rich and diverse enhancer landscapes. Some neural genes had more than ten enhancers each, suggesting that crucial functions like nerve growth and communication are managed by densely layered regulatory control rather than a single on/off switch.
Zooming in on brain and organ specialisation
By correlating enhancer activity with gene expression levels, the authors linked nearly 9,000 enhancers to about 4,300 genes and uncovered many tissue-specific control pairs. In the cerebellum, for example, a unique enhancer region appears to drive a brain factor called BDNF differently from the nearby cortex, helping to explain subtle differences between parts of the brain that share the same gene. Similar patterns emerged in heart, gut, and adrenal tissues: organs can use overlapping gene sets, but distinct enhancers in each tissue fine-tune when and how strongly those genes are used. Analysis of DNA methylation showed that chemical marks on enhancers generally dampen gene activity, reinforcing their role as sensitive regulators rather than passive stretches of DNA.
What makes sheep and other mammals different
To see how sheep compare to other mammals, the team crossed their map with regulatory data from humans, mice, pigs, and cattle. Promoters proved highly conserved, but enhancers varied much more between species, echoing the idea that evolution rewires gene control more than it changes the genes themselves. The authors identified sets of enhancer–promoter pairs found only in ruminants like sheep and cattle, enriched for processes such as sugar breakdown in the rumen and long-chain fatty acid handling. This suggests that specialized digestive systems and metabolism in these animals are driven in part by unique regulatory wiring, rather than unique genes alone.
Links to traits farmers care about
Because many economically important traits depend on subtle shifts in gene regulation, the team overlaid millions of genetic variants and known trait-associated regions onto their regulatory map. They discovered variants inside enhancers that likely influence milk fat yield by altering control of a gene involved in copper and fat metabolism, COMMD1. They also found birth-weight–associated variants within a cerebellum-specific enhancer predicted to regulate the gene XKR4, providing a plausible route from DNA change to growth traits. These examples show how enhancer maps can turn anonymous genome-wide association signals into concrete biological hypotheses about how traits arise.
What this means for the future
For a layperson, the core message is that this study transforms the sheep genome from a static parts list into a wiring diagram, showing how and where genes are controlled in real tissues. By cataloging hundreds of thousands of regulatory switches, clarifying their activity in different organs, and linking them to traits and species differences, the work offers a powerful foundation for breeding healthier, more productive animals and for understanding how complex traits emerge from gene regulation. Similar maps in other species, including humans, will deepen our grasp of health, disease, and evolution by focusing on how genes are managed, not just what genes are present.
Citation: Xie, S., Davenport, K.M., Salavati, M. et al. Identification of cis-regulatory elements provides insights into tissue-specific gene regulation in the sheep genome. Nat Commun 17, 2413 (2026). https://doi.org/10.1038/s41467-026-70382-7
Keywords: tissue-specific gene regulation, sheep genomics, enhancers and promoters, livestock functional genomics, ruminant evolution