Clear Sky Science · en
CPF-CF-terminated snoRNAs shuttle through the cytoplasm via an mRNA guard protein-mediated surveillance mechanism
How tiny RNA guides take an unexpected detour
Inside every cell, the making of proteins depends on a precise molecular choreography. A key part of this dance is controlled by small nucleolar RNAs, or snoRNAs, which help sculpt the cell’s ribosomes—the machines that build proteins. This study reveals that some of these tiny guides unexpectedly leave the nucleus, briefly visit the cytoplasm, and then return, all because of how their production is stopped. Understanding this hidden traffic route sheds light on how cells safeguard RNA quality and protect their genetic information.
A closer look at the cell’s RNA helpers
SnoRNAs are short RNA molecules that act as guides, directing chemical tweaks on other RNAs, especially those that form ribosomes. In yeast cells, most snoRNAs are made in the nucleus and thought to stay there for life, forming stable partnerships with specific proteins to create snoRNPs, the working complexes that modify ribosomal RNA. Yet earlier work had detected snoRNAs contacting proteins normally involved in sending messenger RNAs out of the nucleus. This puzzling observation raised a question: do snoRNAs sometimes travel to the cytoplasm, and if so, why?
Discovering snoRNAs on the move
By re‑examining RNA sequencing data from carefully separated nuclear and cytoplasmic fractions, the authors found that many snoRNAs are indeed present in the cytoplasm of normal yeast cells, at levels similar to typical messenger RNAs. When they disabled key export factors, Mex67 and Xpo1, the cytoplasmic pool of snoRNAs shrank and immature, 3′‑extended snoRNA precursors built up in the nucleus. Microscopy using fluorescent probes confirmed this shift: signals that normally appeared both in the nucleolus and faintly in the cytoplasm became strongly nuclear when export was blocked. These findings show that a subset of snoRNAs, often still carrying extra sequence at their ends, is actively transported out of the nucleus and not just leaking or contaminating samples.
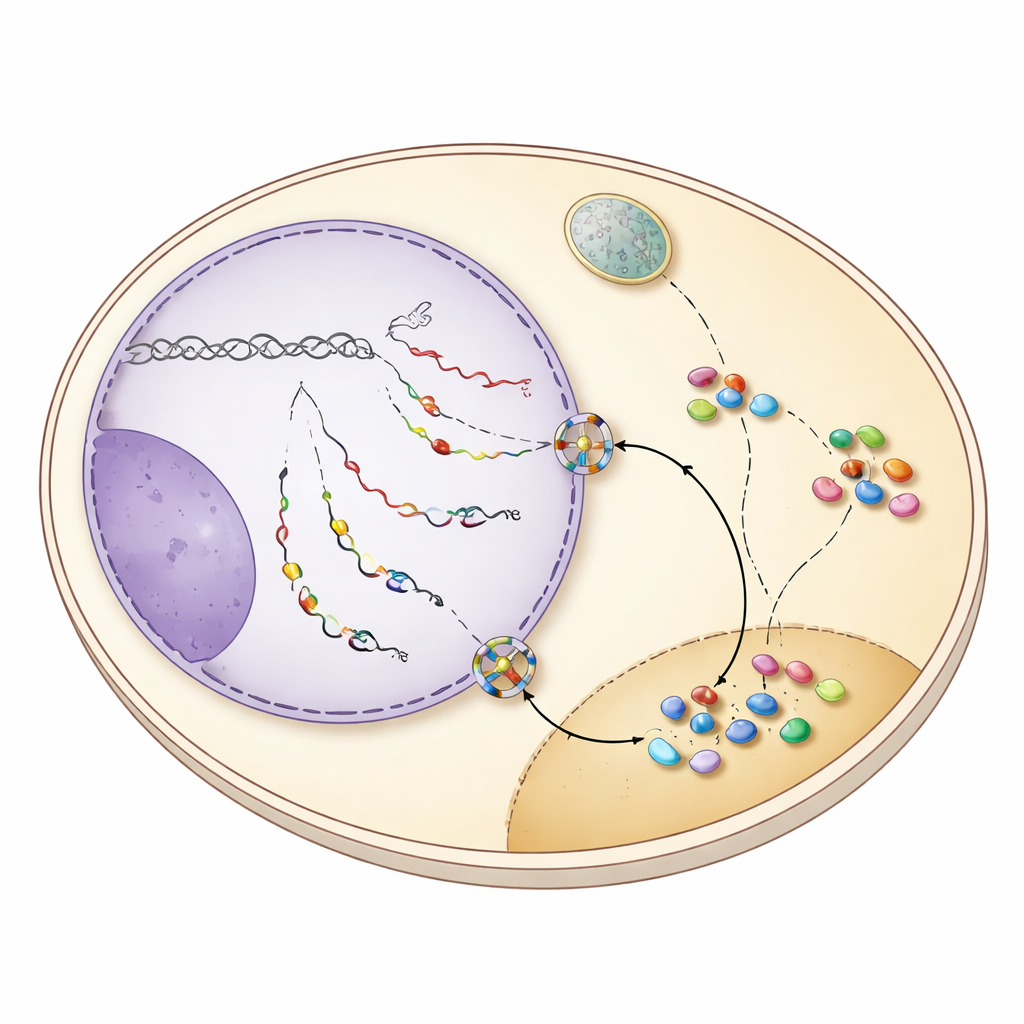
The guard‑protein switch that sends snoRNAs out
The key to this shuttling behavior lies in how snoRNA transcription is terminated. In yeast, most snoRNAs are normally cut loose from the transcription machinery by a system called NNS, which leaves them with a short tail that is quickly trimmed in the nucleus. Many snoRNA genes, however, also harbor backup stop signals recognized by a second system, CPF‑CF, best known for finishing messenger RNAs and adding long poly(A) tails. When NNS termination falters, snoRNA transcripts run on to these downstream CPF‑CF sites and receive a longer tail. This altered ending recruits a set of “guard” proteins—Hrp1 and Nab2 among them—that check correct processing and simultaneously attract the export factor Mex67. The study shows that when snoRNAs are finished by CPF‑CF, guard proteins and Mex67 work together to send these tailed snoRNAs through nuclear pores into the cytoplasm.
Round‑trip tickets and quality control
Once in the cytoplasm, snoRNAs are not left unattended. They remain bound by a protective ring of Lsm proteins and the factor Lhp1, which shield the vulnerable RNA end. These features allow two import receptors, Cse1 and Mtr10, to recognize the shuttling snoRNAs and bring them back into the nucleus. When the authors disabled these import factors or the Lsm ring, immature snoRNAs accumulated in the cytoplasm, confirming that this machinery mediates their return. Back in the nucleus, the long tails are trimmed by the nuclear exosome, core snoRNP proteins are fully assembled, and the mature complexes relocate to the nucleolus. Importantly, snoRNAs that had taken this detour remained fully functional: a model snoRNA, snR13, still carried out its specific chemical modifications on ribosomal RNA after being terminated by CPF‑CF and sent through the cytoplasm.
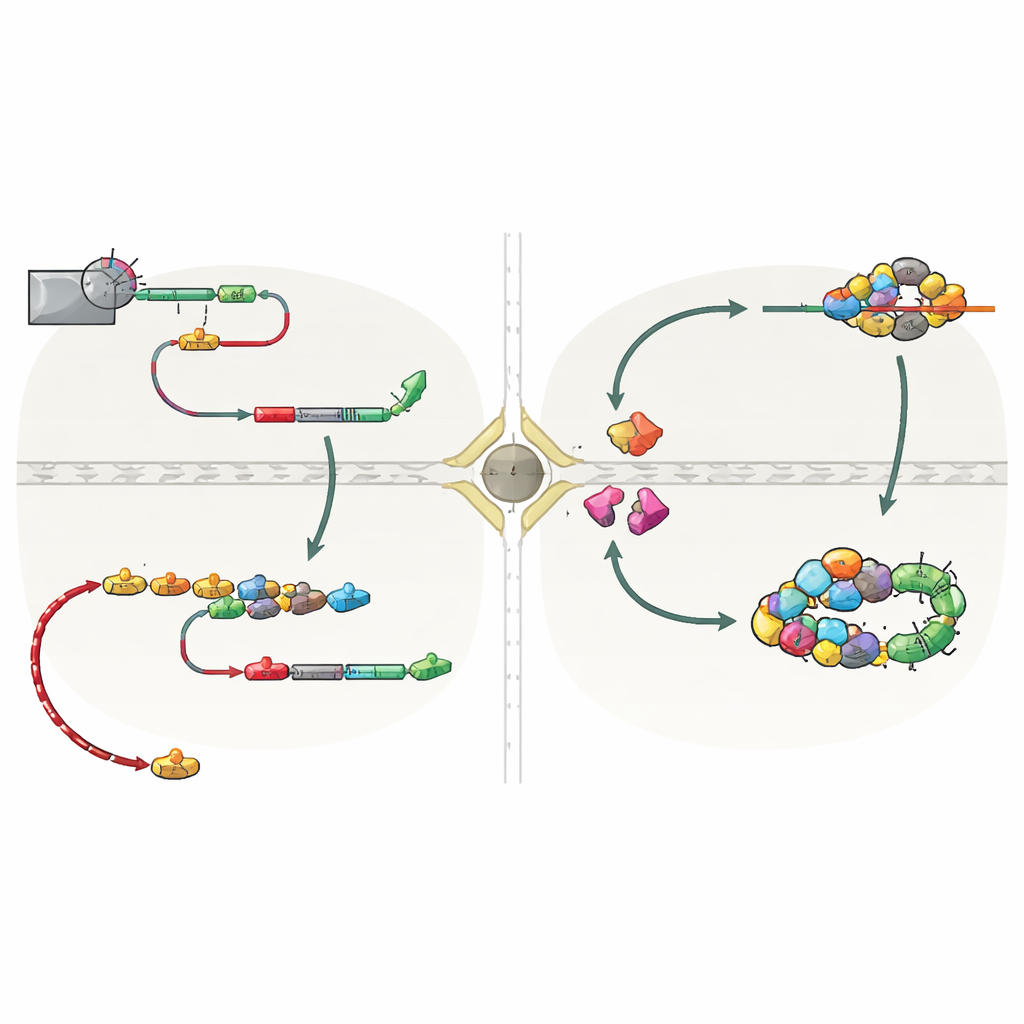
Why this hidden pathway matters
This work reveals that snoRNA travel is not a special case, but a built‑in backup route controlled by the way transcription ends. When the primary NNS system is weakened—for example, at particular cell‑cycle stages—downstream CPF‑CF signals rescue snoRNA production. This prevents waste of already synthesized RNA and avoids runaway transcription that could interfere with neighboring genes and damage DNA. The authors also find hints of a similar backup in human cells, where individually encoded snoRNAs can acquire poly(A) tails at related sites. In simple terms, the study shows that the “stop signal” chosen at the end of a snoRNA gene decides whether that RNA quietly matures in the nucleus or briefly leaves and returns under the watchful eye of guard proteins. This transcription‑termination‑driven surveillance mechanism helps keep the cell’s RNA landscape both flexible and safe.
Citation: Yu, F., Zaccagnini, G., Duan, Y. et al. CPF-CF-terminated snoRNAs shuttle through the cytoplasm via an mRNA guard protein-mediated surveillance mechanism. Nat Commun 17, 2328 (2026). https://doi.org/10.1038/s41467-026-70373-8
Keywords: snoRNA, RNA quality control, nuclear export, transcription termination, yeast genetics