Clear Sky Science · en
Control of lysosome function by the GTPase-activating protein TBC1D9B and its binding partner TMEM55B
How Cell Recycling Centers Keep the Balance
Inside every one of our cells sit tiny sacs called lysosomes that act like recycling centers and control hubs for growth. When they work well, worn-out parts are broken down and their components are reused; when they fail, wastes pile up and diseases, including neurodegeneration and cancer, can emerge. This study reveals how two little‑known proteins help switch lysosomes between a roaming, growth‑friendly state and a clustered, waste‑clearing mode, shedding light on how cells adapt to feast and famine.
Moving Recycling Centers Around the Cell
Lysosomes are not fixed in place. They travel along internal tracks, pulled outward by one set of molecular motors and drawn inward by another. Where they sit in the cell matters: when nutrients are plentiful, lysosomes spread toward the cell’s outer regions and help support growth signals; when food is scarce, they retreat to the center, become more acidic, and ramp up the breakdown of cellular material. A small switch‑like protein called ARL8 is known to push lysosomes outward, but until now no one had identified a dedicated braking system to rein it in. The authors suspected that such a brake would be crucial for letting cells rapidly shift into a conservation mode during starvation.
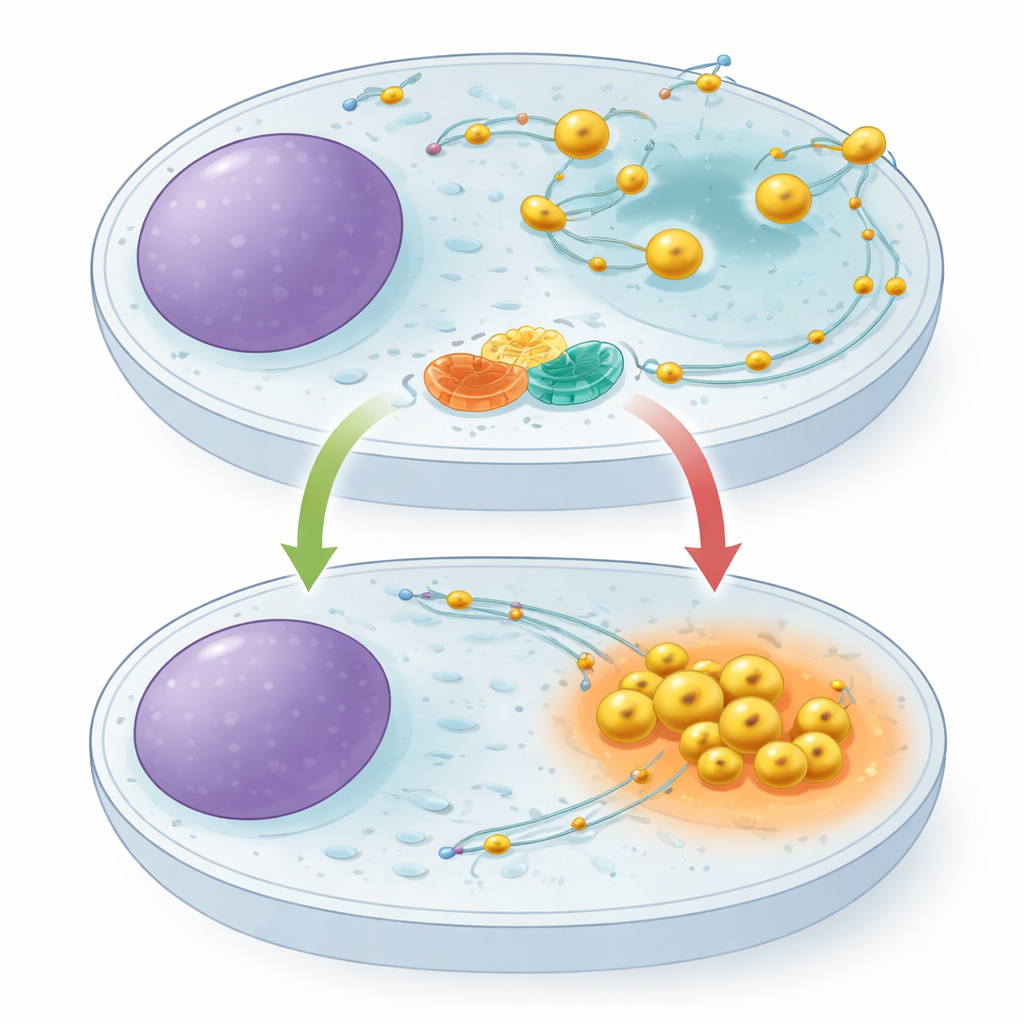
A Molecular Partnership at the Lysosome Surface
Using protein fishing techniques and mass spectrometry, the researchers discovered that a protein named TBC1D9B attaches to a lysosomal membrane protein called TMEM55B. TMEM55B sits across the lysosome’s outer shell, while TBC1D9B is largely soluble and can dock onto the membrane. The team showed that these two form a direct complex: purified TBC1D9B sticks to the exposed part of TMEM55B in test tubes, and tagged versions of the proteins pull each other down from cell extracts. Careful mapping of protein segments revealed that several regions of TBC1D9B contact TMEM55B, positioning TBC1D9B at the lysosome surface without blocking the part of the protein responsible for its catalytic activity.
When the Brake Fails, Lysosomes Run Wild
To find out what this partnership does, the authors used CRISPR editing to remove either TMEM55B or TBC1D9B from human cells. In both cases, lysosomes no longer clustered near the nucleus but instead scattered toward the cell edge and moved faster along their tracks, mimicking cells in a constantly fed state. Reintroducing normal TBC1D9B restored proper positioning, but a version lacking catalytic power did not, indicating that its enzymatic activity is essential. When the cells were starved, control cells pulled their lysosomes inward and boosted their digestive power, as shown by increased activity of key enzymes and enhanced breakdown of test cargo. Cells missing TMEM55B or TBC1D9B failed to do this: their lysosomes stayed peripheral and their degradative response to starvation was blunted. Autophagy, the process by which cells self‑eat damaged components, was also impaired, with less efficient processing of an autophagy reporter and accumulation of the adaptor protein p62.
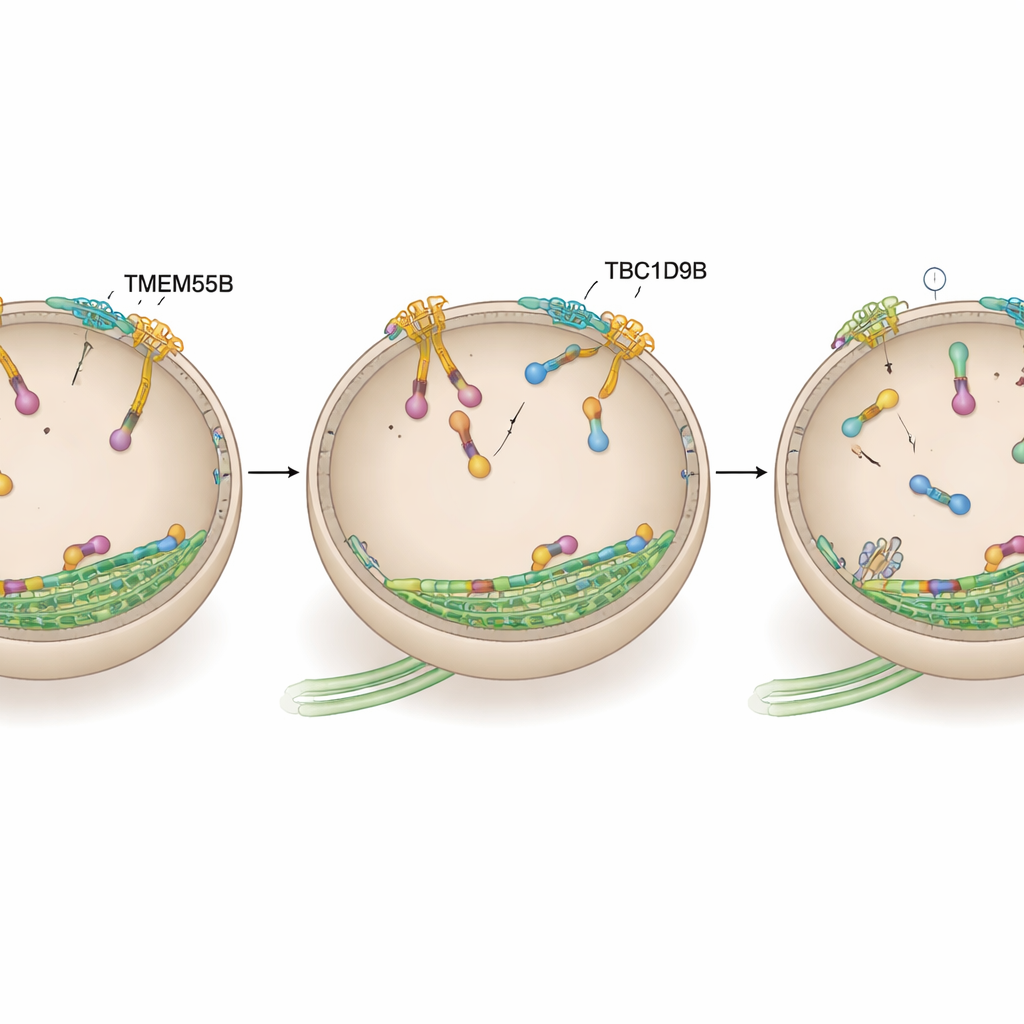
Turning Off the Lysosome Motor
The team then asked whether TBC1D9B exerts its effects by acting directly on ARL8, the outward‑driving motor switch. Using proximity labeling in human neurons and binding tests in cell lines and purified proteins, they showed that TBC1D9B selectively binds to the active, GTP‑loaded form of the ARL8B variant, but not to its inactive form or to the closely related ARL8A. Structural modeling predicted that key residues in TBC1D9B contact the GTP pocket of ARL8B. In biochemical assays, TBC1D9B sped up the hydrolysis of GTP bound to ARL8B, effectively converting the protein from its “on” to its “off” state; a mutant version of TBC1D9B lacking these residues could no longer do so. Consistent with this, cells lacking TMEM55B or TBC1D9B showed increased ARL8B on lysosomes, while overexpressing TBC1D9B pulled lysosomes back toward the center, similar to dialing down ARL8B activity.
A New Control Knob for Cellular Housekeeping
Finally, the authors tested whether this ARL8B brake explains the cellular changes seen when TBC1D9B is missing. When ARL8B was depleted, lysosomes stayed clustered near the nucleus regardless of whether TMEM55B or TBC1D9B were present, and the defects in autophagy caused by TBC1D9B loss were largely overridden. Together, the data support a model in which TMEM55B recruits TBC1D9B to lysosomes, where it inactivates ARL8B and allows lysosomes to shift from a dispersed, growth‑supporting state to a centralized, digestion‑focused one. For non‑specialists, this means the study has uncovered an important dial that cells use to decide when to recycle more aggressively, a process with implications for disorders of waste buildup in the brain, metabolism, and cancer.
Citation: Duhay, V., Tian, M., Kosieradzka, K. et al. Control of lysosome function by the GTPase-activating protein TBC1D9B and its binding partner TMEM55B. Nat Commun 17, 2487 (2026). https://doi.org/10.1038/s41467-026-70345-y
Keywords: lysosomes, autophagy, cell signaling, organelle transport, neurodegeneration