Clear Sky Science · en
Convergent evolutionary shifts in AGT targeting between mitochondria and peroxisomes across mammal transitions to herbivory
How Plant-Eating Mammals Rewired a Detox Enzyme
Many mammals switched from eating mostly meat or insects to diets rich in leaves, fruit, or seeds. Plants are nutritious but chemically tricky: they generate toxic byproducts that animals must safely dispose of. This study asks how one key liver enzyme, AGT, has been repeatedly re-engineered during mammal evolution so that plant‑eaters can better detoxify compounds from their leafy meals.
A Cellular Traffic Problem Inside the Liver
AGT is a liver enzyme that prevents the build‑up of oxalate, a compound that can form damaging calcium oxalate crystals in organs such as the kidney. Where AGT sits inside cells matters. In meat‑eating mammals, a molecule called glyoxylate, which AGT converts to harmless glycine, is mostly produced in mitochondria, the cell’s power plants. In plant‑eating mammals, glyoxylate mainly arises in tiny sacs called peroxisomes, which handle many detox jobs. For AGT to work efficiently, it needs to be in the same compartment where glyoxylate appears. That means evolution has had to solve a cellular “traffic routing” task: should AGT be sent to mitochondria, peroxisomes, or both?
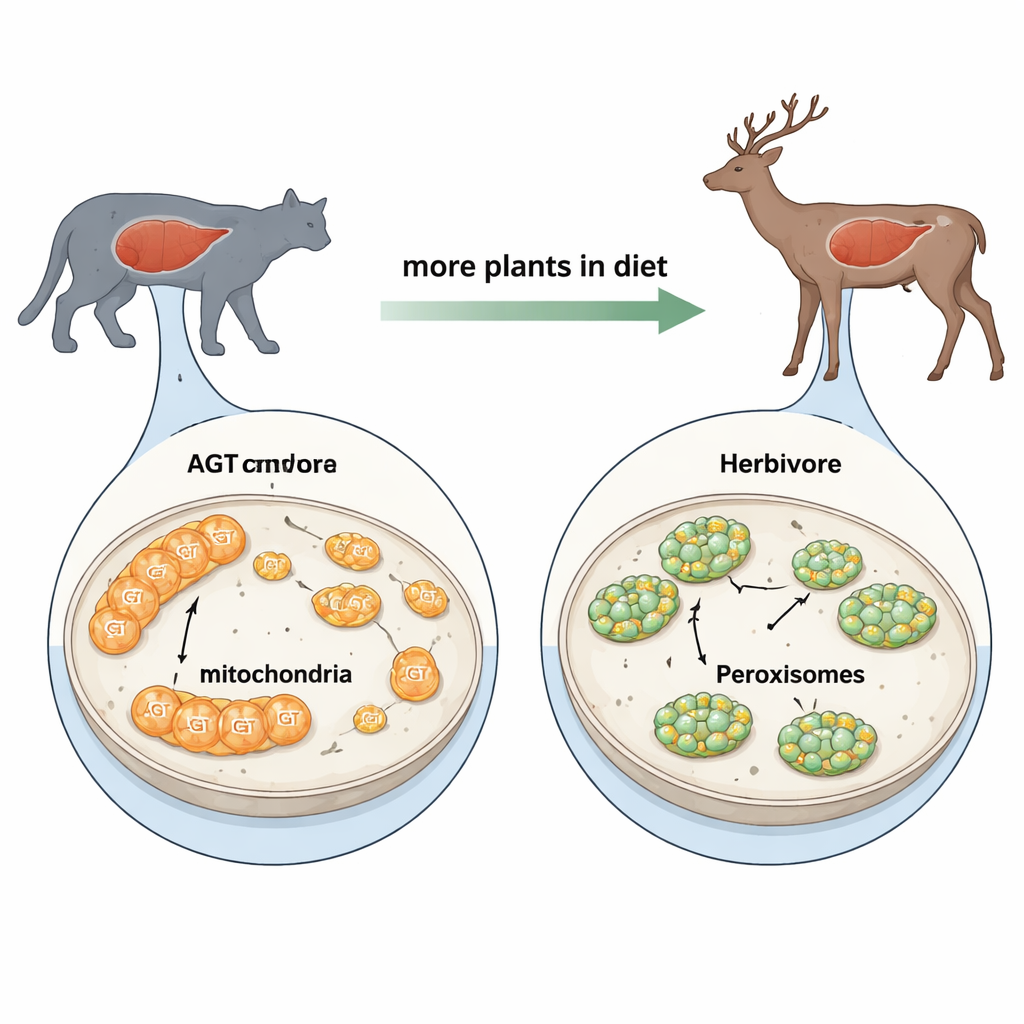
Two Address Labels That Compete
AGT carries two built‑in address labels. At its front end is a short stretch called the mitochondrial targeting sequence, which directs it to mitochondria. At its tail is a three‑letter code called PTS1, which sends it to peroxisomes. Earlier work mainly focused on the front label and treated the peroxisome code as a backup. By comparing AGT genes from almost 500 mammal species and running cell‑based experiments on dozens of them, the authors show this view is incomplete. Plant‑eating lineages often have damaged or truncated mitochondrial labels, but their PTS1 codes remain intact and are frequently upgraded to highly efficient versions. In contrast, meat‑eaters tend to keep strong mitochondrial labels and weaker peroxisome codes.
Convergent Changes Linked to Plant-Rich Diets
Across the mammal family tree, the researchers found that efficient peroxisome codes—specific three‑letter endings such as SKL, SRL, or GKL—have evolved again and again in unrelated herbivores. In many of these species, lab imaging shows AGT clustering in peroxisomes, even when the mitochondrial label is still present. When the scientists experimentally removed the PTS1 code, plant‑eating species showed a sharp drop in peroxisomal targeting, whereas carnivores changed little. Genetic analyses further revealed that the PTS1 region has experienced stronger adaptive evolution than the rest of the enzyme, suggesting natural selection has repeatedly fine‑tuned this tiny address tag as diets shifted toward plants.
Changing Where the Enzyme Starts, Not Just Its Labels
AGT has another twist: the gene can be read from two different starting points. The longer version includes the mitochondrial label; the shorter one skips it and produces a form that relies mainly on the peroxisome code. Using RNA data from 172 mammal species, the team found that herbivores tend to favor the shorter, peroxisome‑bound version, while carnivores more often use the longer, mitochondria‑directed form. In species where epigenetic data were available, plant‑eaters showed weaker activity and lower DNA accessibility around the upstream start site, and stronger activity near the downstream one. This indicates that changes in gene regulation, not only in the protein sequence, help steer AGT toward the cellular compartment best matched to the animal’s diet.
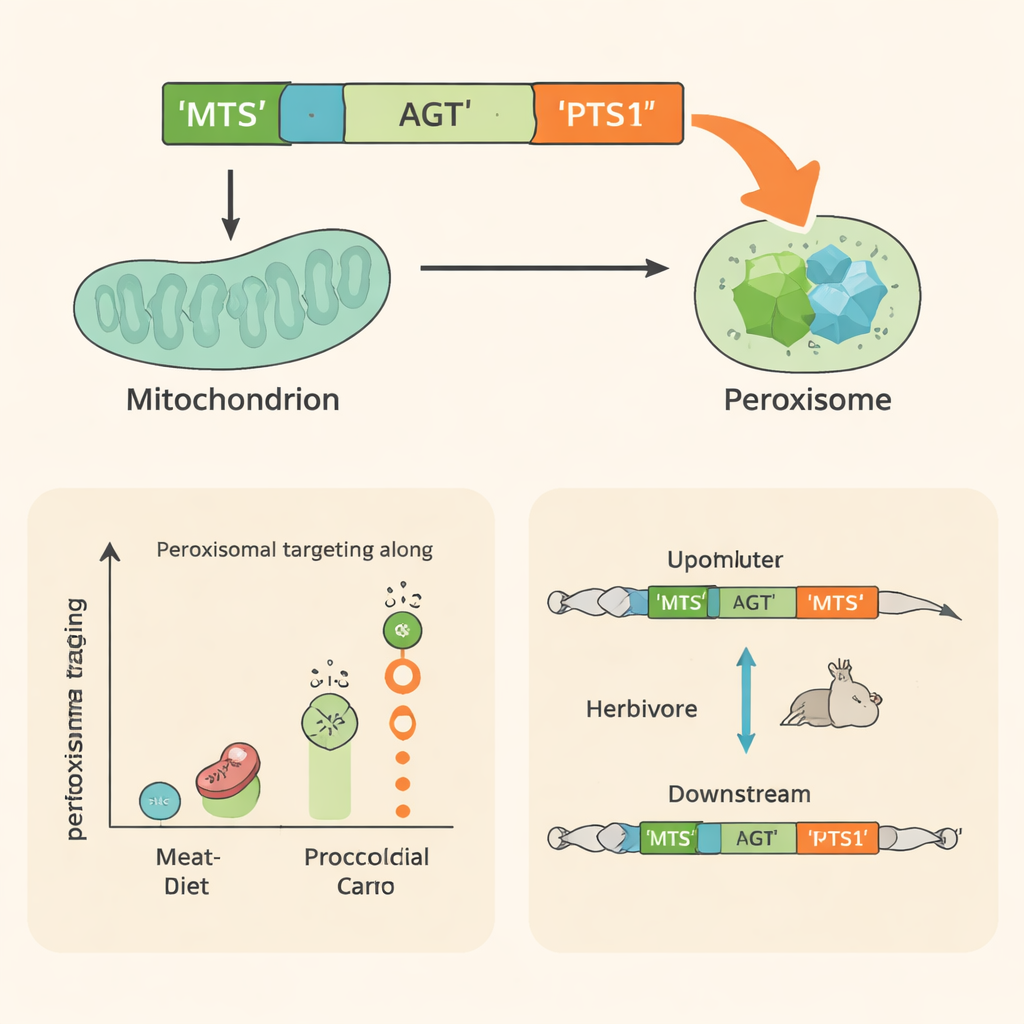
Multiple Paths to the Same Solution
By combining evolutionary comparisons, cell imaging, and gene‑expression analyses, this work reveals that mammals have repeatedly solved the same metabolic challenge—detoxifying glyoxylate from plant foods—through similar outcomes but diverse routes. Herbivores commonly boost AGT’s presence in peroxisomes by degrading the mitochondrial address, upgrading the peroxisome code, shifting transcription to avoid the mitochondrial label, or using combinations of these strategies. For non‑specialists, the message is that even tiny molecular details, such as a three‑letter tag on a protein or a change in where a gene is switched on, can be reshaped by natural selection to support major lifestyle transitions like the move from hunting prey to grazing on plants.
Citation: Huang, C., Wang, B., Yu, J. et al. Convergent evolutionary shifts in AGT targeting between mitochondria and peroxisomes across mammal transitions to herbivory. Nat Commun 17, 2161 (2026). https://doi.org/10.1038/s41467-026-70246-0
Keywords: herbivory, mammal evolution, cellular detoxification, protein targeting, glyoxylate metabolism